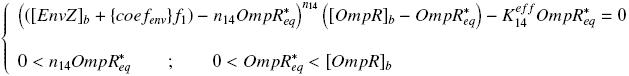Team:Paris/Modeling/f3bis
From 2008.igem.org
|
Method & Algorithm : ƒ3bis
In this experiment, we have [EnvZ]real = {coefenvZ} ƒ1([aTc]i) but we use [aTc]i = Inv_ƒ1( [EnvZ] ) so, at steady-states, phosphorylated OmpR verify : We can then solve it, and reintroduce the result in the previously characterized ƒ3( 0, [OmpR*] ) , to determine the parameters : ↓ Table of Values ↑
↓ Algorithm ↑
<Back - to "Implementation" | |
 "
"