Team:University of Chicago/Project
From 2008.igem.org
| Home | Team | Project | Parts | Modeling | Notebook | Meetings | Papers |
|---|
Contents |
Synthetic Biology: Engineering E. coli to express Mefp-3,5, and Mgfp-5 proteins
Email questions, comments, etc to parijata@uchicago.edu
Synthetic biology is generally defined as the engineering of biology: the synthesis of complex, biologically based (or inspired) systems, which display functions that do not exist in nature. This engineering perspective may be applied at all levels of the hierarchy of biological structures —from individual molecules to whole cells, tissues and organisms. It is the fusion of genetic engineering and bioinformatics, in an effort to enable the design of biological systems in a rational and systematic way . We are interested in designing and synthesizing artificial biological systems in a systematic, hierarchical manner. This is about developing a biological programming language. We want to move from an ad-hoc genetic engineering approach to a scalable engineering framework.
Our research group is interested in furthering this effort by engineering E. coli to express Mefp-3,5, and Mgfp-5 proteins, which are found naturally in mussels (Mytilus edulis and Mytilus galloprovincialis). These mussel foot proteins are strong bioadhesives and powerful anti-biofouling agents, with applications for biomaterials and biomedical research . We aim to produce results that will achieve the initial goals of genetic engineering, as well as further the conceptual goals of synthetic biology. The final goal of this research is to produce four biological Systems meeting the specifications of the Standard Registry of Biological Parts: 1) Expression of Mefp-3 in E. coli, 2) expression of Mefp-5 in E. coli, 3) expression of Mgfp-5 in E. coli, and 4) concomitant expression of Mefp-3,5 and Mgfp-5 in E. coli.
The basic ideas underlying synthetic biology are, in short: 1) automated DNA construction and 2) standards of abstraction. It is the goal of synthetic biologists to produce a standardized registry of “biological parts”, which enables the development of an open-source biological programming language . Automated DNA construction is becoming easier and cheaper, enabling us to synthesize and express it on increasingly larger scales. Abstraction is the process of generalization by reducing the information content of a concept or an observable phenomenon, typically in order to retain only information that is relevant for a particular purpose. It is a mechanism and practice to reduce and factor out details so that one may focus on a few concepts at a time. For example, abstracting “a well-worn, bouncy, orange basketball” to simply “a ball” retains only the information on the attributes and behavior of a general ball. This will, however, allow us to calculate the behavior of the ball, using generalized equations for similar objects. In the same way, such abstractions allow us to hide the complexity of the genetic code, and utilize a greater amount of parts in a simpler design.
With this in mind, special attention will be paid to organization of the sequence data, with the intent of contributing to the abstraction hierarchy used in the Standard Registry of Biological Parts (an open-source effort located at partsregistry.org). Within synthetic biology, the genetic code is abstracted into chunks, known primarily as biological "parts." These parts allow us to build increasingly complex systems; putting several parts together creates a "device," which is regulated by start codons, stop codons, restriction sites, and similar coding regions known as "features." The hierarchy is characterized primarily thus far by levels of biological parts, devices, and systems. Parts are meant to be interchangeable, and typically consist of sequence designed to perform a single function (e.g., protein coding regions, terminators, conjugation, regulation, ribosome binding sites, etc). Devices are built up of parts (e.g., reporters, inverters, signalers, protein generators, composite devices, etc). Systems are composed of several devices, designed to perform a more complex series of tasks within the cell. It is our aim to organize the genetic material necessary to express the selected mussel foot proteins into standard biological parts, devices, and systems, and add them to the Registry, making them available for widespread use.
In mussels, these protein adhesives are secreted as fluids that undergo an in-situ cross-linking reaction, leading to the formation of a solid adhesive plaque, which allows the mussel to attach itself to various substrates. The most notable of the structural features of mussel adhesive proteins (MAPs) is the presence of L-3,4-dihydroxyphenylalanine (DOPA), an amino acid responsible for both adhesive and cross-linking characteristics of MAPs. DOPA is formed in these proteins by post-translational hydroxylation of tyrosine residues. Although the exact role of DOPA in these proteins is unknown, evidence suggests that bulk oxidation of DOPA residues leads to intermolecular cross-linking of the plaque proteins, giving rise to solidification of the adhesive. Similarly, interfacial adhesion to substrates is generally believed to be caused by chemical interactions between the unoxidized catechol form of DOPA, and functional groups at the surface of the solid substrate. Our group is interested in the engineered expression of three of these proteins in particular: Mefp-3,5 and Mgfp-5. These three proteins contain the highest DOPA concentrations of the mussel foot protein family, and so would provide us with the highest predicted yield when expressed in E. coli.
Methods
A tentative outline of the methods to be used, based upon recent research in the field, is described here. Both methods for conducting the experiment and determining success (through SDS-PAGE and Western blot analysis, MALDI-TOF mass spectroscopy analysis modification of tyrosine residues, investigation of coating of various substances, measurement of protein on a gold surface, measurement of adhesion force) are tentative, and will rely heavily on revision/input from Phillip Messersmith:
Vector Design. Using sequence data obtained from GenBank, Waite et al , Cha et al , and others, we will organize the information for Mefp-3 Mefp-5 and Mgfp-5 into the standardized hierarchy of biological parts, devices, and systems. The aim is to produce four final standardized systems: one for the expression of each protein, and one larger system that provides the necessary information for the production of all three. It is the goal to create blocks of genome that may be inserted into the E. coli much like an object or command may be entered into a block of code. All regulatory information necessary for expression will be included, and separated into devices and parts, in accordance with the standards outlined in the Registry . Finished systems, devices, and parts will be uploaded to the Registry for public use.
Vector Construction. Our group has already gathered the sequence data necessary for the expression of Mefp-3,5 and Mgfp-5. Construction of the expression vector will require the mature cDNA PCR fragments of Mefp-3,5 and Mgfp-5, as described in the previous section, inserted into the PstI and EcoRI sites of the pTrcHisA vector (Invitrogen) containing a hexahistidine (His6) tag at the N-terminus, to simplify protein purification, and a trc promoter that is inducible by isopropyl-β-D-thiogalactopyranoside (IPTG). The constructed vectors will be classified in accordance with the Standard Registry of Parts.
Cell culture and analysis. E. coli will be grown and maintained in Luria-Bertani (LB) medium. Cell growth will be monitored by measuring optical density using a UV-visible spectrophotometer. When cultures reach an OD600 of 0.7 to 0.8, 1mM (final concentration), IPTG will be added to induce expression of recombinant Mefp-3,5 and Mgfp-5. Samples will then be centrifuged and cell pellets will be stored at -80°C for further analysis. Total protein concentration will be determined using the Bio-Rad Bradford assay with bovine serum albumin (BSA) as a protein standard.
Purification of recombinant Mefp-3,5 and Mgfp-5. Immobilized-metal affinity chromatography purification will be performed using the Acta Prime purification system (Amersham Biosciences). Affinity purification will be performed under denaturing conditions. Harvested cell pellets will be resuspended in buffer B (8 M urea, 10 mM Tris-HCl, 100 mM sodium phosphate [pH 8.0]) per g (wet weight). Samples will be lysed by gentle shaking for 1 h at room temperature, lysates centrifuged at 14,000 rpm for 20 min at room temperature, and the supernatant collected for purification. The column will be equilibrated with 5 resin volumes of buffer B, after which 10 ml of denatured cell lysate was loaded, and then the column washed with buffer C (8 M urea, 10 mM Tris-HCl, 100 mM sodium phosphate, [pH 6.3]) and buffer D (8 M urea, 10 mM Tris-HCl, 100 mM sodium phosphate [pH 5.9]). Target recombinant Mefp-3.5 and Mgfp-5 shall be eluted with buffer E (8 M urea, 10 mM Tris-HCl, 100 mM sodium phosphate [pH 4.5]). Eluted recombinant Mefp-3.5 and Mgfp-5 samples will be dialyzed in 5% acetic acid overnight at 4°C, using Spectra/Por molecular porous membrane tubing (Spectrum Laboratories).
SDS-PAGE and Western blot analysis. Samples will be resuspended in protein sample buffer (0.5 M Tris-HCl [pH 6.8], 10% glycerol, 5% sodium dodecyl sulfate [SDS], 5% ß-mercaptoethanol, 0.25 % bromophenol blue) and heated to 100°C for 5 min. After centrifugation for 1 min, proteins will be separated by SDS-polyacrylamide gel electrophoresis (PAGE) (15% [wt/vol] polyacrylamide) and then detected using Coomassie blue staining (Bio-Rad) or silver staining (Bio-Rad), or Western blotted. The membrane will be scanned, and its image analyzed using gel-analyzing software.
MALDI-TOF mass spectroscopy analysis. Matrix-assisted laser desorption ionization (MALDI) time-of-flight (TOF) mass spectrometry analysis will be performed on a Sinapinic acid in 30% acetonitrile-0.1% trifluoroacetic acid matrix solution. Samples will be diluted 1:25 with matrix solution, and 1 µl per plate will be spotted onto gold-plated sample plates and evaporated using a vacuum pump. Mass spectra will be acquired in positive-ion mode
Modification of tyrosine residues. To investigate adhesion, purified recombinant Mefp-3,5 and Mgfp-5 that was resolved in 5% acetic acid buffer to prevent auto-oxidation of DOPA residues to o-quinone under basic pH conditions will be modified with tyrosinase to convert tyrosine residues into DOPA. As a positive control, Cell-Tak is a commercially available naturally extracted M. edulis mussel adhesive protein mixture of Mefp-1 and Mefp-2, that already contains DOPA residues in 5% acetic acid buffer.
Investigation of coating of various substances. The ability of recombinant Mefp-3,5 and Mgfp-5 to coat the following surfaces will be investigated: glass slide, poly(methyl methacrylate) plate, polystyrene plate, commercial silicone-based antifouling agent (SigmaGlide; Sigma Coatings)-coated slide, and aluminum plate. Each material surface is to be cleaned by washing with distilled water several times and drying with nitrogen gas. A 10-µl drop of 1.44-mg/ml protein solution will be added to each material surface and incubated in a humid environment for 12 h at 25°C. After being dried, each surface shall be washed thoroughly with deionized water for 2 h with shaking. The coating by each protein will be visualized using Coomassie blue staining.
Measurement of protein on a gold surface. The quartz crystal (Seiko EG & G) used for the quartz crystal microbalance (QCM) experiment is to be an AT-cut piece of quartz 5 mm in diameter with a basic resonant frequency of 9 MHz. A 5-µl drop of 1.44-mg/ml protein sample will be placed onto the gold surface of the quartz crystal and incubated in a humid environment for 12 h at 25°C. After being dried, the gold surface will be rinsed thoroughly in deionized water for 2 h with shaking, and the remaining deionized water will be evaporated using a vacuum chamber. Dried quartz crystal will be connected to a quartz crystal analyzer (QCA917; Seiko EG & G), and variations in resonance frequency measured and converted to mass using the following equation, on the basis that resonant frequency decreases as a function of increasing mass: 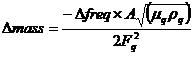
where ∆mass is mass change, ∆freq is resonant-frequency change, µq is the AT-cut quartz crystal constant (2.947 x 1.011 g/cm/s2), ρq is the quartz crystal density (2.648 g/cm2), Fq is the reference frequency (9.00 MHz), and A is the quartz crystal surface area (0.196 cm2).
Measurement of adhesion force. The force-distance curve will be obtained using atomic force microscopy (AFM) (SPA400; Seiko Instruments). All force data will collected by the methods of Messersmith et al.
We hope to accomplish these goals through a comprehensive and collaborative study of genetics, systems biology, computer science, and trial-and-error.
One-Paragraph Lay Summary
Synthetic biology is a novel approach to engineering biology, focusing on organizing genetic information into “standard parts,” comparable to interchangeable parts in manufacturing industries such as automobiles or electronics. The fundamental goal of synthetic biology is the organization of genetic information into standardized regions of code, that will create the functional equivalent of an open-source biological programming language. As opposed to ad hoc genetic engineering methods, that do not necessarily emphasize the organization of genetic data, synthetic biology aims to move towards a scalable engineering framework. This framework is built upon the principles of standards of abstraction. Abstraction, the process of generalization to reduce information content, allows us to hide the complexity of the genetic code, and utilize a greater amount of information in a simpler design. There is a centralized database to catalog this effort, undertaken by a community of scientists, and manifested in the Standard Registry of Biological Parts (partsregistry.org). This is the primary library of standardized biological parts to which researchers around the globe contribute. It is our aim to add to the Registry, by organizing the genetic material necessary to express mussel foot proteins (Mefp-3, Mefp-5, and Mgfp-5) in E. coli. These proteins have both bio-adhesive and anti-biofouling properties. Bio-adhesion gives mussels the ability to attach to various substrates, while anti-biofouling prevents the accumulation of unwanted biological material on selected surfaces. This makes these proteins a unique and promising target of study for biomedical research (e.g. synthetic implants, pacemakers, artificial organs, and internal prosthetics).
 "
"
