Team:Cambridge/Bacillus/Lab Work
From 2008.igem.org
(→September 8th) |
(→September 10th) |
||
| Line 1,906: | Line 1,906: | ||
* Result : nothing on the gel!!! But we can not manage to PCR anything... there is a problem in our PCR protocol, or products! | * Result : nothing on the gel!!! But we can not manage to PCR anything... there is a problem in our PCR protocol, or products! | ||
| - | = | + | =August 29th= |
| - | == | + | ==Stock of ECE112== |
| - | - | + | - Bulk up ECE112 (Amp100) rfom 10mL LB (28/08), take 1mL and add to 100mL of LB in big flask |
| - | - | + | - Grow overnight in 37°C incubator with vigorous shaking |
| - | + | - Aliquot in 10mL tubes, pellet and freeze cell pellets | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ==Transformation of glycerol stock== | |
| - | + | * Strian : IA751 | |
| - | + | - Transfer into Eppendorf tubes and spin down for 30min | |
| - | + | -Remove glycerol and add medium B | |
| - | + | ||
| - | - | + | - Add ECE112 and ECE153, and one control tube |
| - | - | + | - incubate for 30min in 37°C |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| + | - Plate out ECE112 (Cm5), ECE153 (Spc50), DNA less (blank, Cm5, Spc50) | ||
=September 11th= | =September 11th= | ||
Revision as of 22:05, 28 October 2008
July 22nd
We have 4 tubes from last year. These strains are frozen in glycerol.
- PNZ8901 plasmid in E.coli MC1061 Strain
- Chloroamphenicol resistant
- "sure" vector
- does not integrate
- B. subtilis strain 1A1
- No resistance
- Same as strain 168 but tryptophan deficient
- E.coli strain MC1061, no plasmid
- PP182 plasmid in E.coli strain DH5alpha
- Ampicillin resistance
- integrates
Checking antibiotic resistance
Purpose : Grow each of them on a plate to test their antibiotic resistance
Protocol
- Warm up frozen tubes
- Take 15uL of each and put it on plates without antibiotic
- Incubate at 37°C
July 23rd
- Take one colony from each of the plates grown from the tubes yesterday and plated them again on LA without antibiotics
1/ Purpose Check antibiotic resistance of our different strains
| Strain | Plasmid | Extracted from | Antibiotic use | Concentration of antibiotic |
|---|---|---|---|---|
| DH5alpha | PP182 | tube | Amp | 100 |
| MC1061 | NONE | tube | Cm | 35 |
| MC1061 | PNZ8901 | tube | Cm | 35 |
| DH5alpha | PP182 | plate | Amp | 100 |
| MC1061 | NONE | plate | Cm | 35 |
| MC1061 | PNZ8901 | plate | Cm | 35 |
2/ Purpose : Find the right concentration of antibiotic so that B. subtilis survive
- Grow 15uL of B. 1A1 (frozen tube) in 5mL of LB
- Incubate at 37°C
3/ Purpose : Grow plasmids in TOP10, transformation
4 plasmids :
- I746000
- I746100
- I746101
- I746001
1 control : PUC19
- Add 20uL of TOP10 competent cells and 0.5uL of Plasmid in an eppendorf
- 30min on ice
- Heat shock : 60s at 42°C
- 2min on ice
- Add 89.5uL of SOC
- 60min at 37°C
- Put the mix on plate with ampicillin resistance
- Incubate at 37°C
July 24th
- Test of antibiotic resistances of strains of last year
| Strain | Plasmid | Extracted from | Antibiotic use | Observations | Conclusion |
|---|---|---|---|---|---|
| DH5alpha | PP182 | tube | Amp | Many colonies | Amp Resistance OK |
| DH5alpha | PP182 | plate | Amp | Many colonies | Idem |
| MC1061 | NONE | tube | Cm | Nothing | As expecting, no Cm Resist. |
| MC1061 | NONE | plate | Cm | Nothing | Idem |
| MC1061 | PNZ8901 | plate | Cm | Maybe a few colonies | Contamination?? |
| MC1061 | PNZ8901 | tube | Cm | No colonies | no Cm Resist. |
- Transformation of E.coli with different plasmids from last year
| Strain | Inserted plasmid | Antibiotic | Observation | Conclusion |
|---|---|---|---|---|
| E.coli | I746000 | Amp | No colonies | Antibiotic resistance unknown, no Amp Resist., Pb : no terminator |
| E.coli | I746100 | Amp | No colonies | Antibiotic resistance unknown, no Amp Resist., Pb : no terminator |
| E.coli | I746001 | Amp | Many colonies | Transformation OK |
| E.coli | I746101 | Amp | Many colonies | Transformation OK |
Antibiotic test for Bacillus resistance
We want to find the lowest concentration of antibiotic which kills Bacillus.
Dilution of Amp. [ 1:1000 means 1 part stock to 1000 part Sterile Distilled Water ]
| Concentration (μg/mL) | 100 | 75 | 50 | 25 | 10 |
|---|---|---|---|---|---|
| 100 mg/mL Stock | 1:1000 | 3:4000 | 1:2000 | 1:4000 | 1:10000 |
Dilution of Cm. [ 1:1000 means 1 part stock to 1000 part Sterile Distilled Water ]
| Concentration (μg/mL) | 35 | 25 | 15 | 10 | 5 |
|---|---|---|---|---|---|
| 100 mg/mL Stock | 1:1000 | 1:1400 | 3:7000 | 1:3500 | 1:7000 |
- Add Disks in antibiotic solution for 5 mins [ Protocol to sterilize tweezers : Wipe with Kimwipes, Ethanol, Flame ]
- Melt Soft Agar
- 3mL SA + 10μL Cells 1A1 from LB prepared on 23/7/08
- Pour SA+Cells over blank hard agar plates
- put disks with different concentration of Amp/Cm above SA
- Incubate at 37°C
Results [Plates were prepared before lunch, at 5pm there were visible growth]
-Amp Plates: Huge rings of no growth around 100, 75, 50, 25, and 10.
-Cm PLates: Tiny rings of no growth around 5, 10, 15 and 25. Small ring of no growth around 35 but not well defined. (No clear zones)
Salts for making Bacillus Competent
10x Medium A Base
- Yeast Extract 10g
- Casamino Acid 2g
- add Distilled water to 900mL
-Aliquot into 5 different bottles. [180mL each]
10x Bacillus Salts
- NH4)2SO4 20g
- K2PO4 anhydrous 139.66g
- KH2PO4 60g
- Tri-Sodium Citrate 10g
- MgSO4.7H2O 2g
- Add Distilled water to 1000mL
-Aliguot into 5 different bottles. [200mL each]
Medium B
- Preparing 50mM CaCl2.2H2O
- CaCl2.2H2O 1.470g
- Add Distilled water to 20mL [Final conc. 500mM]
- Take 10mL of 500mM
- Add 90mL of distilled water [Final conc. 50mM]
- Preparing 50mM CaCl2.2H2O
- Preparing 250nM MgCl2.6H2O
- MgCl2.6H2O 0.508g
- Add Distilled water to 10mL [Final conc. 250mM]
- Take 1mL of 250mM
- Add 99mL of Distilled water [Final conc. 250μM]
- Take 1mL of 250μM
- Add 99mL of Distilled water [Final conc. 250nM]
- Preparing 250nM MgCl2.6H2O
- Add 100mL of 50mM CaCl2.2H2O
- Add 100mL of 250nM MgCl2.6H2O
-Total volume 200mL
NOTE: To complete Medium B, Take 0.2mL of this solution
July 25th
Results from Yesterday
- Antibiotic test
Results for Amp
Even with a concentration of 10μg/mL, Amp kills Bacillus. Since we want to know which is the lowest concentration of Amp which kills B.S, we are going to test with some lower concentrations.
Results for Cm
Even with a concentration of 35μg/mL, ther is not a clear area around the antibiotic disk. So we have to test some higher concentration of Cm.
Wet Work
- New antibiotic tests
Dilution of Amp
| Concentration (μg/mL) | 10 | 7.5 | 5 | 2.5 | 1 |
|---|---|---|---|---|---|
| 10 μg/mL Stock (from yesterday) | 1:1 | 3:4 | 1:2 | 1:4 | 1:10 |
Dilution of Cm
| Concentration (μg/mL) | 35 | 37.5 | 40 | 42.5 | 45 |
|---|---|---|---|---|---|
| 35 μg/mL Stock (from yesterday) | 1:1 | 5:6 (from 45μg/mL) | 1:875 | 17:18 (from 45μg/mL) | 9:7000 |
- Melt Soft Agar - 3mL SA + 10μL Cells 1A1 from LB prepared on 23/7/08 - Pour SA+Cells over blank hard agar plates - Put 5 blank disks on each agar plate - Add 40μL of antibiotic on each disk - Incubate at 37°C
Results
Nothing! 1A1 cells were kept in the freedge! B.S. can not be kept in the fridge, low temperatures kill them!
July 28th
- Results of antibiotic plates from yesterday
For Cm, 35μg/mL should be enough to kill B.S.
For Amp, nothing can be concluded!
Wet Work
- Check plasmid ppL82
We had 2 samples of ppL82 in DH5α in LB solution (~4mL), one from the frozen glycerol tube and one from a colony picked on a plate. Normally plasmid and cells should not be kept in freezer. So, we want to extract this plasmid, check its size and then keep it in the fridge. We do that for the 2 different samples.
- Plasmid Miniprep (standard protocol)
- Test the concentration of DNA in each tube
- ppL82 plate : 89.8ng/mL
- ppL82 tube : 71ng/mL
- Preparation of different stocks of strains and plasmids
PNZ8901
- Plate a single colony (from 23/07/08) on a Cm plate
- Incubate at 37°C
- Grow the entire frozen glycerol tube in 20mL of LB without antibiotic to check if this stock is still good (we will check the plasmid on a gel tomorrow)
- Incubate at 37°C
MC1061
-Pick e single colony of MC1061 (from 22/07/08) and put it in 10mL of LB (to make glycerol stocks)
-Incubate at 37°C
1A1
- Add 100μL of LB+1A (mix of friday) and 10mL of LB
- Incubate at 37°C
- Make B.S. 1A1 competent and transform them
- Prepare Medium A
Add 81mL of SDW, 10mL of 10X Medium A base and 9mL of 10X Bacillus salts
- Make B.S. 1A1 competent
- In 10mL of medium A and add about 10 colonies of B.S.
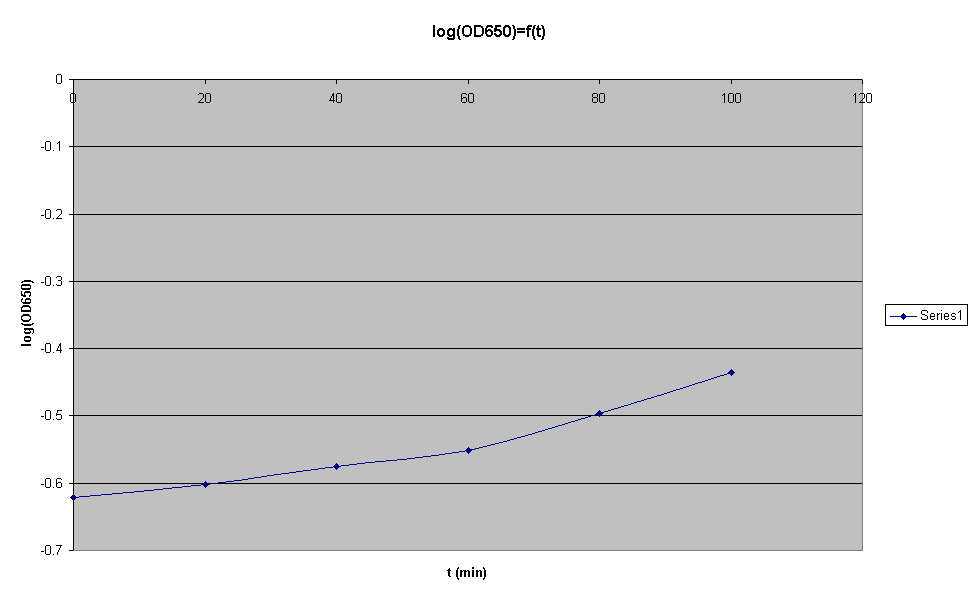
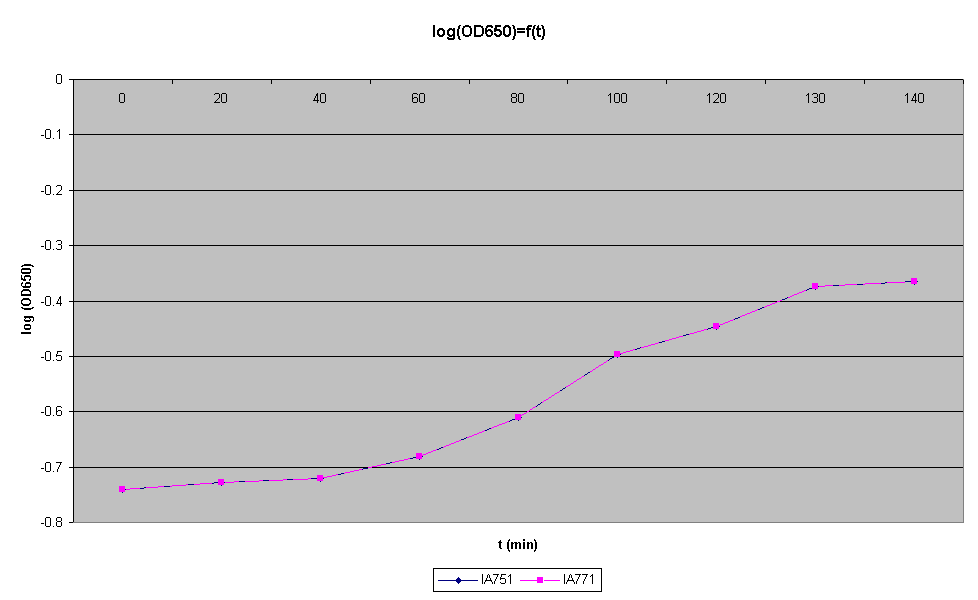
- Check the OD650, you should have an OD between 0.1 and 0.2. t0 for OD = 0.1876
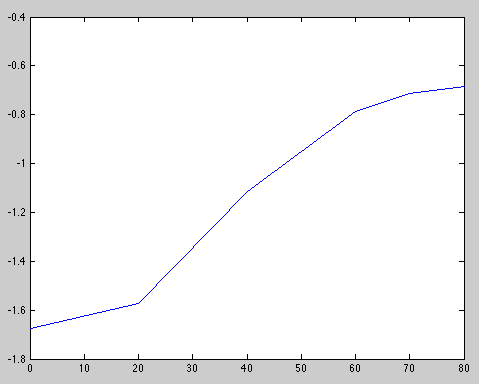
- Check the OD650 every 20min and plot OD against time on semi-log paper. When the point at the culture leaves log growth, it is ok
| Time (min) | 0 | 20 | 40 | 60 | 70 | 80 |
|---|---|---|---|---|---|---|
| OD650 | 0.1876 | 0.2074 | 0.3282 | 0.4545 | 0.4895 | 0.5040 |
At, t0 = 70min, log growth seems to stop.
- Incubate at 37°C 90min after t0
- Warn 10 Ependorf tubes with 0.45mL of Medium B
- Add 50μL of culture in each tube of Medium B at t90
- Incubate the diluted culture at 37°C with vigorous aeration for 90min
- Storage : we want to store 7 tubes
6 tubes in the freezer (with 60μL of glycerol)
1 tube on the bench
- Transformation
- We make ppL82 from plate, ppL82 from tube, and a control
- Add 0.5μg of DNA (15μL)
- Incubate at 37°C for 30min
- Plate on Cm plates
July 29th
- Result from the gel (29/07/2008)
- Lane8 : ppL82 (plate)
- Lane9 : ppL82 (tube)
- Lane 10 : I746001
- Lane 11 : I746101
- Lane 12 : hyperladderI
The ladder seems to be wrong. So it was really difficult to ckeck the size of our plasmid ppL82. To make, that, we assume that the size of our biobricks was ok, and we estimate the size of our plasmid. The plasmid ppL82 seems to have the right size, but we will have to check again to be able to use a ladder.
- Competence of B.S. kept on the bench
We observed B.S. with a microscope. Problem, all B.S. seems to be dead!
Possible reasons of this problem :
- Problem with the vector (it could be a wrong vector, we are not really sure)
- Quantity of DNA, we added 0.5μg. The protocol was 1μg, but normally it should be ok.
- B.S. may not be competent (we have to test motility with microscope)
- Cells could be dead : replate them
- Try to add tryptophan in the medium
Probable reason : we forgot to dilute the medium B, that's why it did not work
- Results from transformation plates (B.S.)
There are 2 colonies on a plate, but it does not look like Bacillus. Moreover, there is also a colony on our negative control plate. SO it must be contamination (resistant to CM!). We will do the transformation again.
Wet Work
- Prepare medium for transformation of B.S.
- Preparation of medium B: 10mL of medium A and 0.2mL of 50mMCaCl22(H2O) + 250mMgCl26(H2O)
- Preparation of medium A with tryptophan : 81mL of SDW, 9mL of 10X Bacillus salts, 10mL of 10X Medium A base and 0.1mL of Tryptophan (11mg/mL)
- Check plasmid PNZ8901
- Plasmid miniprep (same protocol with 60μL of elution buffer)
- Digest : with PstI and SalI from Biolabs, and Buffer 3 (add 15μL of DNA)
- Gel (17μL of sample and 3μL of dye)
Results from the gel
- Lane9 : PNZ8901
- Lane10 : HyperladderI
The sizes we expected were about 1100kb and 2100kb. The sizes should be ok!
July 30th
- Transforming Bacillus Subtilis with medium A and medium A + tryptophan
Medium A
| Time (min) | 0 | 20 | 40 | 60 | 73 | 85 | 95 | 105 |
|---|---|---|---|---|---|---|---|---|
| OD650 | 0.1394 | 0.1370 | 0.1660 | 0.2304 | 0.2778 | 0.3138 | 0.3373 | 0.3583 |
For this medium, the curb stop to increase logarithmically at 100min.
Medium A + tryptophan
| Time (min) | 0 | 20 | 40 | 60 | 73 | 85 | 95 |
|---|---|---|---|---|---|---|---|
| OD650 | 0.1493 | 0.1636 | 0.2080 | 0.2909 | 0.3512 | 0.4002 | 0.4165 |
For this medium, the curb stop to increase logarithmically at 90min.
- Incubate 90min at 37°C by shaking
- Prepare 7 tubes for medium A and 7 tubes for medium A + tryptophan : add 0.45mL of prewarmed medium B and 0.05mL of culture and incubate 90min at 37°C by shaking.
- Transform B.S. by adding DNA and incubate 30min at 37°C
We have one transformation with ppL82 (add 7.5μL of DNA), one with PNZ8901 (add 10μL of DNA) and one negative control without DNA.
- Plate 500μL on Cm plate (35μg/mL)
- For the remaining tubes, in one, add 60μL of glycerol and keep it in the freezer and keep the other one on the bench (for each medium)
- Checking our big stock of biobricks and PNZ plasmid
We did big culture of biobricks and PNZ8901 plasmid to be able to make stocks. We want to check them before making stocks.
- Plamsid miniprep (for I746001, I746101 and PNZ8901 from big flasks
- Measure DNA concentration in our samples to decide the volume of DNA we have to add in our preparation
| 260/280 | DNA concentration (ng/nl) | |
|---|---|---|
| I746001 | 1.79 | 50.5 |
| I746101 | 1.87 | 54.3 |
| PNZ8901 | 1.84 | 87 |
- Digest
| For I746001 and I746101 | For PNZ8901 plasmid |
|---|---|
| 14μL of SDW | 14μL of SDW |
| 2μL of 10X Fast Digest Buffer | 2μL of Buffer 3 (Biolabs) |
| 2μL of DNA of biobricks | 2μL of DNA of PNZ8901 |
| 1μL of EcoRI | 1μL of PstI |
| 1μL of SpeI | 1μL of SalI |
- Gel
- Results from this gel
- Lane 9 : I746001
- Lane 10 : I746101
- Lane 11 : PNZ8901
- Lane 12 : HyperladderI
Everthing is really too big! There is a problem, either dimerization, either contamination, either a problem in our work. So we are going to run a new gel.
- New gel to check
- Miniprep plasmid from growth bottles (I746001, I746101 and PNZ8901); from plates (I746001, I746101 and PNZ8901).
- Do single digest for PNZ8901 (one with PstI, one with SalI)
- Run a gel with : PNZ8901 from friday, PNZ8901 digest with PstI, PNZ8901 digest with SalI, 3 samples from growth bottles, 3 samples from plates (to check the size of the uncut vectors)
- Result from this second gel
The ladders are really bad! But the size of our biobricks and plasmid are too big! So we can not trust these big cultures. They is a problem. With tests from the first week, we know that biobricks are right, so we are going to grow new cultures from these first cultures, to make stocks.
Concerning the vectors, they are from last year, so we are not sure of what they are. Since we ordered new well defined vectors (we should receive them on friday), we will use them in the next steps to be sure of our work.
August 1st
- Grow received vectors
| Vectors | AmpR (μg/μL) | CmR (μg/μL) | KanR (μg/μL) |
|---|---|---|---|
| ECE112 | 100 | 0 | 0 |
| ECE147 | 50 | 0 | 0 |
| ECE149 | 50 | 0 | 0 |
| ECE150 | 50 | 0 | 0 |
| ECE151 | 50 | 0 | 0 |
| ECE153 | 100 | 0 | 0 |
| ECE162 | 100 | 0 | 0 |
| ECE165 | 100 | 10 | 0 |
| ECE166 | 100 | 10 | 0 |
| ECE171 | 50 | 0 | 10 |
| ECE172 | 100 | 10 | 0 |
| ECE176 | 100 | 5 | 0 |
- Preparation of Agar plates
We have bottles of 200mL.
| Type | Amp (μL) | Cm (μL) | Kan (μL) |
|---|---|---|---|
| Amp100 | 200 | 0 | 0 |
| Amp50 | 100 | 0 | 0 |
| Amp100 + Cm10 | 200 | 57.1 | 0 |
| Amp100 + Cm5 | 200 | 28.6 | 0 |
| Amp50 + Kan10 | 50 | 0 | 80 |
- Preparation of LB tubes
We prepare 10mL of LB in which we add a single colony.
| Type | Amp (μL) | Cm (μL) | Kan (μL) |
|---|---|---|---|
| Amp100 | 10 | 0 | 0 |
| Amp50 | 5 | 0 | 0 |
| Amp100 + Cm10 | 10 | 2.9 | 0 |
| Amp100 + Cm5 | 10 | 1.4 | 0 |
| Amp50 + Kan10 | 5 | 0 | 4 |
August 2nd
Yesterday's Results
- All Plates grew!
- ECE 172 seemed to have trouble growing on the plate (no single colonies, just one big lump at the start of the streak)
- ECE 150 grew too well! (possibly hard to pick out single colony)
- Tubes:
- ECE 176, 172, 153 seem to be quite clear!!
- ECE 150, 151 had strange floating 'colonies' in the tube
- All other tubes were quite cloudy
Lab Work
Fridged all plates except ECE 172 ==> Benched.
All tubes palleted and placed in freezer although ECE 176, 172, 153 doesn't seem to have any pallets
All red plates does not seem to have any extra growth despite the air conditioning had been turned off when I entered the lab this afternoon.
August 4th
Wet Work
- Check vectors
- for each vector, plasmid miniprep from frozen pellets (step 7 only once, with 60μL of Elution Buffer for ECE112, ECE147, ECE149, ECE 150 and only 30μL for the others)
- Single digest (15μL of SDW, 2μL of Buffer, 2μL of DNA, 1μL of enzyme, then incubate 10min at 37°C and 30 min for the one using HindIII with EcoRI buffer)
- Double digest (mix 7.5μL of each single digest, and then incubate 10min (or 30min for EcoRI + HindIII and EcoRI buffer) at 37°C, then heat shock 5min at 80°C
| Vectors | Enzyme 1 | Enzyme 2 | Buffer | Entire size | Size of part1 | Size of part2 |
|---|---|---|---|---|---|---|
| ECE112 | XbaI | EcoRI | Fast Digest | 10156 | 6900 | 3295 |
| ECE147 | EcoRI | HindIII | EcoRI | 5482 | 4913 | 600 |
| ECE149 | SpeI | PstI | Fast Digest | 6192 | 4919 | 1300 |
| ECE150 | SpeI | HimdIII | Buffer 2 | 6166 | 5300 | 778 |
| ECE151 | SpeI | EcoRI | Fast Digest | 6166 | 4825 | 1300 |
| ECE153 | SalI | Buffer 3 | ||||
| ECE162 | SalI | Buffer 3 | 7600 | 6000 | 1600 | |
| ECE165 | EcoRI | HindIII | EcoRI | 5952 | 5200 | 793 |
| ECE166 | EcoRI | HindIII | EcoRI | 7262 | 5100 | 2103 |
| ECE171 | EcoRI | PstI | Fast Digest | 5444 | 3600 | 1826 |
| ECE172 | HindIII | Buffer 2 | 6462 | 3900 | 2540 | |
| ECE176 | EcoRI | XbaI | Fast Digest | 866 | 4250 | 5128 |
- Results
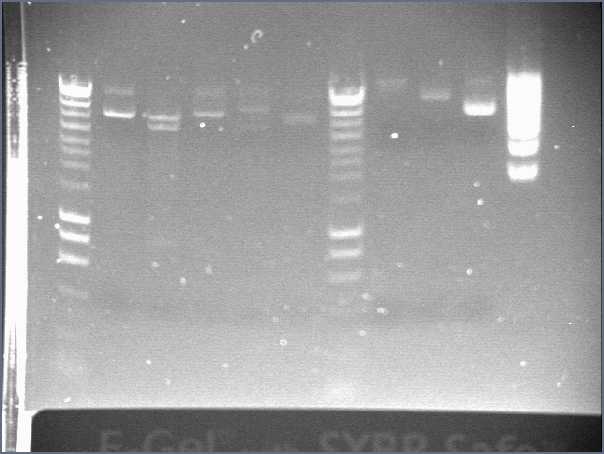
- Lane1 : HyperladderI
- Lane2 : ECE112
- Lane3 : ECE147
- Lane4 : ECE149
- Lane5 : ECE150
- Lane6 : ECE151
- Lane7 : ECE162
- Lane8 : ECE165
- Lane9 : ECE166
- Lane10 : ECE171
- Lane11 : ECE172
- Lane12 : HyperladderI
- ECE112, ECE165, ECE166, ECE171 : right bands, ok
- ECE172 : no band, not enough DNA
- ECE147, ECE150, ECE162 : only one band, maybe not enough DNA
- ECE149 : wrong size!
- ECE151 : not sure, check again
- Plates
- Streak new plates with different strains of Bacillus : 1A1, IA751 and IA771
August 5th
Checking vectors
- Plasmid miniprep for ECE166 (for stock), ECE172 and ECE153
- Digest of ECE153, ECE172 and ECE162 (with plasmid miniprep from yesterday)
- Run on a gel single digest for ECE147 (from yesterday with more DNA), ECE149 (from yesterday with more DNA), ECE150 (from yesterday with more DNA) and ECE 153, ECE162, ECE172
- Results
- Lane3 : HyperladderI
- Lane4 : ECE147
- Lane5 : ECE149
- Lane6 : ECE150
- Lane7 : ECE151
- Lane8 : ECE1162
- Lane9 : ECE172
- Lane10 : hyperladderI
Very low bands : not enough DNA!!!
Transformation of Bacillus
- Result of Nanodrop
| Vector | 260/280 | ng/μL |
|---|---|---|
| ECE112 | 1.75 | 64.6 |
| ECE166 | 172 | 138.6 |
- Prepare medium A with tryptophan, and medium B
- add 5mL of medium A in 3 different tubes, inoculate each tube with colonies (1A1, IA751 ans IA771)
- Check OD every 20min
- Incubate 90min
- Add 0.45mL of medium B and 0.05mL of culture in Ependorf tubes
- Incubate 90min
- Transform : add 1μg of DNA (some with ECE112, some with ECE166) (so you need to nanodrop samples before!)
- Incubate 30min
- Pipette 200μL of solution, spread it on aech plate, wait 10min, and do it again
- Incubate 24hours
- A few glycerol tubes to stock cells : add 2/7 glycerol to cell tubes
- Transformation from glycerol stock from 30/07/2008
- Spin glycerol stocks, pipette out glycerol
- Add 0.5mL of medium B, incubate for 1 hour
- Add 10μL of ECE112 (640ng)
- Incubate 2hours
- Plate 200μL, and 10min later, still 200μL
New stocks
- Do glycerol stock of I746001 and I746101 (no sterile glycerol)
- Put IA751, IA771 in 10mL LB
- Put ECE 176 in 10mL LB + antibiotic
- Reinoculate the tube of LB from yesterday with ECE166 plate
- ECE176 replated onto Amp100 + Cm5
August 6th
Result from transformation
PLATES HAD BEEN BINNED!! OOPS
Check Vectors
- Plasmid miniprep for ECE176 (from 05/08 LB stock and from 04/08 LB stock with a second inoculation)
We want to check uncut plasmid, single and double digest for ECE147, ECE149, ECE150, ECE153, ECE162, ECE172, ECE176. In order to have enough DNA (to see the bands), we will add 1μg of DNA to make single digest, and wr will incubate during 1h (instead of only 10min).
- Nanodrop
| Vetor | 260/280 | μg/mL |
|---|---|---|
| ECE147 | 1.67 | 50.8 |
| ECE149 | 1.66 | 51.9 |
| ECE150 | 1.78 | 107.0 |
| ECE153 | 1.68 | 31.2 |
| ECE162 | 1.56 | 33.1 |
| ECE172 (04/08) | 1.64 | 22.8 |
| ECE172 (05/08) | 1.89 | 213.1 |
| ECE176 (04/08) | 1.89 | 111.1 |
| ECE176 (05/08) | 1.83 | 96.2 |
We made singe digest for the samples from 05/08 only.
- Double digest (1 hour of incubation in water bath at 37°C)
- Gel
Gel1
- Lane1 : HyperladderI
- Lane2 : ECE147 with EcoRI
- Lane3 : ECE147 with HindIII
- Lane4 : ECE149 with SpeI
- Lane5 : ECE149 with PstI
- Lane6 : ECE150 with SpeI
- Lane7 : ECE150 with HindIII
- Lane8 : ECE153 with SalI
- Lane9 : ECE162 with SalI
- Lane10 : ECE172 with HindIII
- Lane11 : ECE176 with EcoRI
- Lane12 : HyperladderI
Gel2
- Lane1 : HyperladderI
- Lane2 : ECE176 with XbaI
- Lane3 : ECE176 double digest
- Lane4 : ECE150 double digest
- Lane5 : ECE149 double digest
- Lane6 : ECE147 double digest
- Lane7 : HyperladderI
- Lane8 : ECE149 uncut
- Lane9 : ECE153 uncut
- Lane10 : ECE172 uncut
- Lane11 : supercoiled ladder
- Results
Our supercoiled ladder was very bad, so it was impossible to conclude for uncut vectors.
- ECE147, ECE 150, ECE153, ECE 176 single digest : ok
- ECE149 : problem, 2 cutting sites for PstI (only one in the sequence)
- ECE162 : only one band whereas SalI should have 2 cutting sites
- ECE172 : wrong sizes!!!
- Double digest for ECE176 : ok!
- Double digest for ECE 150, 149 and 147 : only one band!!!
There may be a problem when we do the double digest by adding two single digest. We will try again with a direct double digest.
August 7th
Transformation of B.S. IA771
New transformation with the same protocol than 2 days ago. We will try to add less liquid on plates (to avoid growing colonies in liquid)
| Tube | Plate 1 | Plate 2 | Plate 3 | Plate 4 |
|---|---|---|---|---|
| ECE166 | Cm5 + 200μL of cells | Cm5 + 100μL of cells | Cm5 + 50μL of cells | Cm10 + 100μL of cells |
| ECE153 | Spc50 + 200μL of cells | Spc50 + 100μL of cells | Spc50 + 50μL of cells | |
| ECE166 | Cm5 + 100μL of cells | Cm10 + 100μL of cells | ||
| ECE153 | Spc50 + 100μL of cells | |||
| no DNA | Cm5 + 100μL of cells | Cm10 + 100μL of cells | Spc50 + 100μL of cells | |
| ECE166 (glycerol stock) | Cm5 + 100μL of cells | Cm10 + 100μL of cells |
Check Vectors
We want to make a final check for ECE147, 149, 150, 153 and 162.
- Double digest + gel
- Results
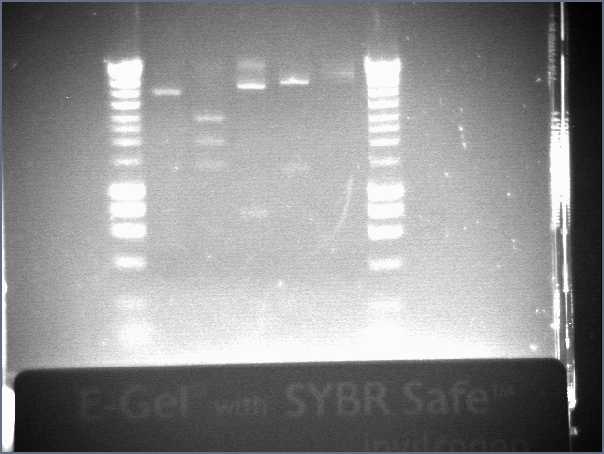
- Lane3 : HyperladderI
- Lane4 : ECE147
- Lane5 : ECE149
- Lane6 : ECE150
- Lane7 : ECE153
- Lane8 : ECE162
- Lane9 : HyperladderI
- ECE 147, ECE150, ECE153 :ok!
- ECE 149 : 3 bands! not the right vector
- ECE162 : ony one band! It has been uncut... maybe the good vector (according to precedent gel), but not sure
Test ECE112 transformation
- for the three different strain (1A1, IA771, IA751), make 12 spots
- take one colony with a loop, and put it on a Cm5 plate and on a Cm5 + Spc100 plate (do that for the 12 spots)
- incubate
New stock
- Put ECE171 in 10mL of LB
- Incubate
August 8th
Result from yesterday transformation
| Vector | Antibiotic | Quantity of cells added | number of colonies |
|---|---|---|---|
| ECE166 | Cm5 | 200μL | 0 : problem!!! |
| ECE166 | Cm5 | 100μL | 2 |
| ECE166 | Cm5 | 50μL | 1 |
| ECE166 | Cm10 | 100μL | 0 : no resistance to Cm10! |
| ECE166 (spin) | Cm5 | 100μL | a lot, confluent |
| ECE166 (spin) | Cm10 | 100μL | 0 |
| ECE166 (glycerol + spin) | Cm5 | 100μL | about 150, very small |
| ECE166 (spin) | Cm10 | 100μL | 0 |
| ECE153 | Spc50 | 200μL | 21 + a lot of confluent colonies |
| ECE153 | Spc50 | 100μL | 7 + confluent (a few) |
| ECE153 | Spc50 | 50μL | 4 |
| ECE153 (spin) | Spc50 | 100μL | 25 + about 200 (small and almost confluent) |
| No DNA | Cm5 | 100μL | 0 |
| No DNA | Cm10 | 100μL | 0 |
| No DNA | Spc50 | 100μL | about 12, maybe more : problem!!!!! |
Result for test of ECE112 transformation
On Cm5 plates, we have some colonies for each strains, on Cm5 + Spc 100 plates, no colonies on each plates. This result seems good, however, we forgot to make a control. Since, we had some colonies on control plates, it is possible that our bacillus are resistant to Cm5, even if they are not transformed. We addded some control colonies.
Control with erythromycin
- Prepare erythromycin, stock 5mg/mL : 0.05g in 10mL of SDW
- Prepare plates : add 20μL of Ery (5mg/mL) into 200mL of agar (final concentration : 0.5μg/mL)
- Do several spots on each plate
| Plate | Colonies added from |
|---|---|
| 1 | ECE166 + 100μL of cells (2 different plates) |
| 2 | ECE153 + 100μL of cells (2 different plates) |
| 3 | ECE166 (from glycerol stock) |
| 4 | Control (from a plate of IA771 without antibiotic) |
| 5 | IA771 + ECE112 (05/08) |
| 6 | IA771 + ECE166 (05/08) |
Amylase production screening
- Prepare agar plate with starch
- Starch : 1g of starch in 10mL of SDW
- Add 2mL of Starch solution into 200mL of agar, mix
-Inoculate on different plates : 1A1 +ECE112 (plate from 05/08), IA751 + ECE112 (plate from 05/08) and IA771 + ECE153
New stocks
- Take 1mL from LB stock oh ECE171 (from 07/08) and put it into 99mL of LB
August 11th
Erytromycin Experiment
- New preparation of Ery : 0.05g into 10mL of ETHANOL
- We plated again our plates (same than 08/08/08)
Amylase screening experiment
- New method : add 2g of starch powder into 200mL of Agar, shake, pipette to plate (and avoid bubbles)
- Same plates than 08/08/08
Test our stock of ECE171
-Plasmid miniprep 9from the samples of 10mL and the one of 100mL)
-Nanodrop both DNA
| Vetor | 260/280 | μg/mL |
|---|---|---|
| ECE171 (10mL) | 1.73 | 136.3 |
| ECE171(100mL) | 1.82 | 99.5 |
- Double digest (EcoRI and PstI) with 10μL of DNA and 1h of incubation
- Samples have been frozen, they should be run onto a gel
Control of Spc resistance of Bacillus
- Spc50 plate with transformed IA771 (ECE153, which is Spc resistant) and with IA771 9which should not be Spc resistant)
August 12th
Results from yesterday
- Ery plates
- IA751 (control) : most of colonies did not grow, which is good, but 2 or 3 grew...
- IA7771 + ECE166 : all colonies grew : ok 9non integration vector)
- IA771 + ECE153 : growth of all colonies... problem!!! if IA771 is transformed with this integration vector, it should lose its EryR
- IA771 + ECE112 : growth of all colonies... problem!!! if IA771 is transformed with this integration vector, it should lose its EryR
- Spc resistance
- IA771 : no growth on Spc50 plate, so no natural resistance
- transformed cells grew
- Amylase plates
- Add iodine solution for 1min
- Problem : no aparent blue, no diffusion of iodine into LA agar + a lot of contamination
New plates and LB stocks
- In order to try to make plasmid miniprep at the end of the day, grow IA771 transformed with ECE166 in 10mL LB + Cm5
7hours is not enough to reach exponential phase!!! We will try again tomorrow by incubating longer
- Grow IA771 and IA751 ( first plate from sterile disks of strain) imto 10mL of LB
- Grow ECE166, 171, 153 in E.coli with approriate antibiotics 9to transform tomorrow)
- Grow IA771 transformed with ECE153 into LB+Spc50 (to check fluorescence with xylose induction)
New test for amylase
We tried 2 different methods.
- Dilute 1g of starch into 100mL of agar and try to dilute it and boil it to sterilize. The problem is that it is very difficult to dilute...
- The second method seem to be better. We diluted 1g of starch into 100mL of Soft Agar (it dilutes very well). Then plate blank agar plate (LA) and then add a thin layer of SA ith 1% of starch. Poke the plates.
- We plated IAI+ECE112, IA751+ECE112 and also IA751 and 1A1 for control.
August 13th
Results for Starch plates
- Add 5mL of iodine
- 1A1 or IA751 : big zones of clearance
- IA751 + ECE112 : no zones of clearance (photos), just small white points on colonies : the gene AmyE seems to be knocked out, transformation ok!
- 1A1 + ECE112 : problem, soft agar melted... impossible to observe!
Transformation of IA751
- We used IA751 plate from 12/08/08, vectors ECE 153, 166, 171
- Spectrophotometer : blank made with medium A
| Time (min) | 0 | 20 | 40 | 60 | 80 | 100 | 120 |
|---|---|---|---|---|---|---|---|
| OD650 | 0.1487 | 0.1541 | 0.1642 | 0.1980 | 0.2470 | 0.3205 | 0.3643 |
- t0 = 120min
- Follow the protocol of transformation
- Plasmid miniprep of ECE153, 166, 171 (to make the transformation)
- Nanodrop ( to add 0.5μg of DNA for transformation)
| Vetor | 260/280 | μg/mL | quantity of DNA to add for transformation (μL) |
|---|---|---|---|
| ECE153 | 1.5 | 18.4 | 27.2 |
| ECE166 | 1.83 | 39.5 | 12.7 |
| ECE171 | 1.83 | 115.8 | 4.4 |
- Plate (with appropriate antibiotics)
- ECE166 : Cm5
- ECE153 : Spc50
- ECE171 : Kan5 (to prepare agar plate, 40μL of Kan 25mg/mL into 200mL of agar)
Check our stock of ECE171
- We made big stocks of ECE171, before keeping it, we wanted to check it, so we run double digest on a gel!!
- Results : problem!!
- 2 possible causes : Double digest was made " days before and kept in the freezer (possible degradation) + too much DNA on the gel
- New double digest (from pellets in the freezer) + new gel
- Results : ok!
Glycerol stocks
- for IA751 and IA771, add 100μL of culture and 500μL of glycerol (60%)
Xylose experiment
- To test the transformation of ECE153, we want to induce the promoter Pxyl, and we will have green fluorescence
- Add 1mL of culture IA751 + ECE153 (from yesterday), 8mL of LB and 45μL of Spc50, 1mL of xylose (1g into 10mL)
Plasmid miniprep for B.S.
- We first tried to use the same protocol than for E.coli (Zyppy kit) for ECE166 transformed in IA751
- Nanodrop to see the result
| Vetor | 260/280 | μg/mL |
|---|---|---|
| ECE166, colony 1 | 1.69 | 11.2 |
| 1.41 | 14.5 | |
| ECE166, colony 2 | 1.34 | 7.0 |
- We have very low concentration of DNA, we will digest that plasmid tomorrow morning to see if it is ECE166, and if it does work, we will try to modify our protocole tomorrow
Transformation of ECE188, 189, 190 in E.coli
- We received this vector in DNA stocks, so we have to transform them to test them 9because we do not have enough DNA)
- Use TOP10 competent cells
- Pellet, add 100μL of CaCl2 solution, for each transformation, use 50μL of cells
- Add 2μL of DNA, and 1μL of PUC19 for control
- 30min on ice
- 2min at 42°C, 2min on ice
- 2h at 37°C
- Plate on Amp100 plates
In Preparation of Beta-galactosidase Assay (Promoter Assay PA)
Biobrick Extraction:
E0040
- GFP (Amp resistance)
- Extracted from 2007 Plate 1 Well 5H
I13522
- E.coli constitutive promoter with RBS and GFP (Amp resistance)
- Extracted from 2007 Plate 3 Well 13C
B0034
- E. coli RBS (Amp resistance)
- Extracted from 2008 Plate 1000 Well 2E
R0040
- E. coli promoter (Amp resistance)
- Extracted from 2008 Plate 1000 Well 4C
Transformation of competent TOP10 with the 4 biobricks above along with pUC19 control
- 20μL TOP10 used for each transformation with 2μL DNA
- Followed standard protocol
- Plated out neat and 1/10 on Amp100 plates after 2 hours of incubation, then plates are kept at 37°C overnight
August 14th
Results of transformation
| Vector | Antibiotic | number of colonies | Transformation efficiency |
|---|---|---|---|
| ECE166 | Cm5 | 31 | 62 |
| Control | Cm5 | 0 | |
| ECE171 | Kan5 | 5 + a lot of small colonies | ? |
| Control | Kan5 | 1 + 5 very small | |
| ECE153 | Spc50 | about 25 (on a side) | 50 |
| Control | Spc50 | 6 + 2 very small |
- Problem with our control, maybe we should check the resistance of competent cells (before transformation)
BioBrick extraction for testing promoters and RBS in B.subtillis
The primers for inducible B.subtillis promoters have been ordered. Meanwhile, we would like to be able to compare the RBS and promoter strengths in E.coli and B.subtillis, using GFP fluorescence to quantify gene expression.
We attempted to isolate 4 BioBricks: I13522 (GFP under constitutive promoter), E0040 (GFP only), R0040 (a promoter), & B0034 (an E.coli RBS). So far, only the former two, extracted from 2007 wells, grew after transformation. Single-colony PCR was used to test the transformants.
Expected VF2-VR fragment sizes:
I13522 - ~2.4 kb E0040 - 958 bp R0040 - 292 bp B0034 - 250 bp
Transformation of vectors 188, 189, 190
- Transformation of yesterday did not work! there was nothing on plates, even on the control plate
- Try again, same protocol, 2μL of DNA, control : PUC9
- Plate on Amp100 for control and Amp100 + Cm5 for vectors
Double digest of ECE166 (extracted from transformed Bacillus yesterday)
- Transformation from 07/08
- 16μL of DNA, 2μL of Buffer EcoRI, 1μL of EcoRI (Biolab) and 1μL of HindIII
- Incubator at 37°C for 35min, heat shock at 80°C for 5min
- Run on a gel with 4μL of DNA
- Result : no bands! not enough DNA!
- Run on a new gel with 16μL of DNA
- Result : " bands, one of about 2000b, and one of about 5000b> The big one seem to be too small...
We are going to check this transformation with fluorescence too!
Erythromycin plate
- Erythromycin plate for the transformation 13/08/08
- IA771 (control, they should survive)
- IA771 + ECE166 (non integration vector, so colonies should survive)
- IA771 + ECE171 (integration vector but in another locus, should survive)
- IA771 + 153 9integration vector, they should die if they are transformed)
New stocks
- LB stocks with antibiotic of ECE151, 153, 166
- LB stock of ECE153 with Spc50 (one from colonies from LB stock from 11/08 and one from 13/08 transformation plates)
Check fluorescence with microscope
- ECE166 from the plate from 13/08
We diluted one colony from this plate into SDW and observed with microscope.
Result : Bacillus is fluorescent! There are some bacteria which are brighter than others, but it is because it is not an integration vector. This transformation worked!!!
- ECE153
We added xylose, but we did not incubate our sample after that, so it did not work> We will try again tomorrow with a period of incubation>
We will have to
Plates from yesterday for Biobrick Extraction (PA)
- Growth observed on Amp100 plates for I13522 and E0040 only but not for R0040, B0034 and pUC9 control
- Re-plated overnight culture neat on Amp 75 plates and incubate at 37°C overnight
Single Colony PCR of I13522 and E0040
- 5 colonies picked from each neat agar plate onto Amp75 plates
- PCR following standard protocols
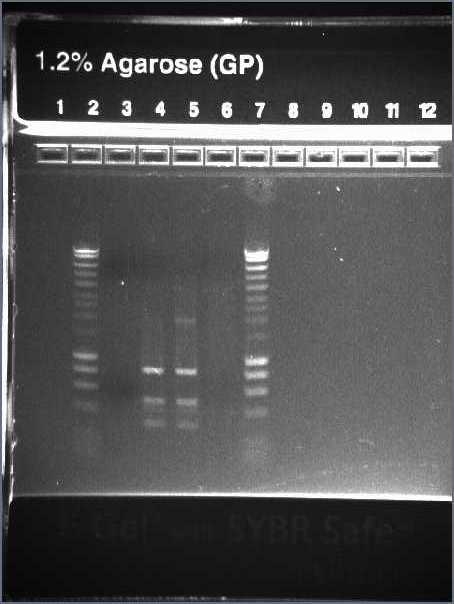
- Run PCR product on 1.2% agarose gel at 70V which is later soaked in EtBr
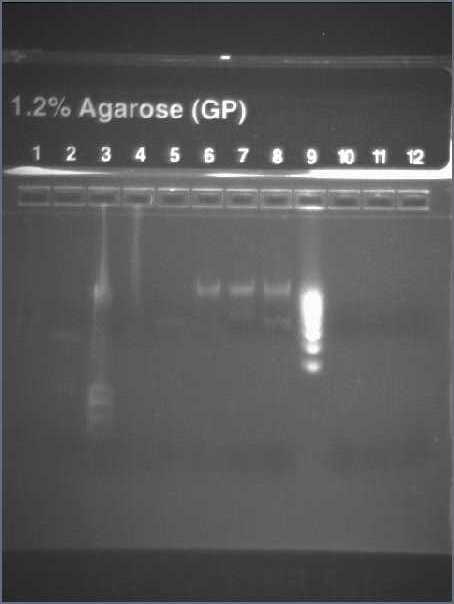
First Row:
- Lane 2 - Hyperladder IV
- Lane 3-7 - I13522 Colonies 1-5 PCR
- Lane 9-13 - E0040 Colonies 1-5 PCR
Second Row:
- Lane 2 - Hyperladder IV
- Lane 3-6, 9 - R0010 Colonies 1-5 PCR
Result:
- Expected size of band: I13522 - 2375bp; E0040 - 958bp; R0010 - 292bp
- Picked colonies 4 and 5 for E0040 into 10ml LB with Amp100 and incubate overnight at 37°C as the band corresponds to about 958bp
- Band of R0010 colonies 3 and 5 correspond to 292bp and will incubate in LB tomorrow when colonies have grown on agar plates
August 15th
Transformation ECE188, 189, 190 into E.coli
- Results : only our control plate grew!!
- Maybe we did not add enough DNA, or there is another problem. We will e mail the guy from Bacillus center before trying again, because we don't have enough DNA.
Plasmid miniprep ECE 151 (x2), ECE 153, ECE 166
- Nanodrop results :
| 260/280 | ng/μL | |
|---|---|---|
| ECE 151 (3) | 1.79 | 86.8 |
| ECE 151 | 1.88 | 106.3 |
| ECE 153 | 1.80 | 32.2 |
| ECE 166 | 1.87 | 36.2 |
Xylose Induction with ECE 153
- 4 Tubes
- No fluorescence! We have to think about the transformation of integration vectors into Bacillus.
Glycerol Stock Transformation
- Spin down cells (competent) from 5/8 and 13/8 stocks
- Remove liquid, resuspend in Medium B
- Incubate for 60 mins at 37°C
- Add ECE 166 into tubes
- Incubate for 30 mins at 37°C
- Plate out on CM 5 plates
- 4 plates
- 771 (5/8) + ECE 166
- 771 (5/8) Control
- 771 (13/8) + ECE 166
- 771 (13/8) Control
Transforming ECE 188, 189, 190 (3rd try)
- 2 tubes of chemically competent top 10
- Spin down, remove liquid
- Resuspend cells in 100μL of CaCl2 50mM
- 4 tubes with 50μL of cells
- ECE 190 (all)
- ECE 189 (all)
- ECE 188 (all)
- PUC 9 (5μL) [ Control ]
- Ice for 30 mins
- 42°C for 2 mins
- Ice for 2 mins
- Incubate at 37°C for 2 hours
- Plate out on Amp 100 (Neat)
Biobrick E0040 and I13522 (PA)
- PCR E0040 colonies 4 and 5 after miniprep of plasmid to ensure plasmid identity again
- Protocol:
- SDW 7.5μL
- MasterMix 10μL
- VR and VF2 1x2μL
- DNA sample 0.5μL
- Run on 1.2% SyBr E-gel
- Lane 2 - 100bp ladder
- Lane 3 - E0040 colony 4
- Lane 4 - E0040 colony 5
- Result: Bands at around 1000bp on both lanes 3 and 4 with a brighter band for lane 3. Correct band as E0040 VF-VR is 958bp
- Minprep of plasmid E0040 from colony 4 is kept
- Transform chemically compotent TOP10 with I13522 biobrick extract again
- Transformation following standard protocols
- TOP10 with I13522 plated neat and 1/10 on Amp. 100 plates and incubate at 37°C overnight
- Will pick single colonies tomorrow for PCR analysis and into LB with Amp100
August 16th
Result from the days before
- For Ery plates
- Everything grew, we will try again to see if there is a problem with the antibiotic (and we will do a negative control with the strain IA751)
- Transformation of E.coli with ECE188, 189, 190
- We have a result from our 3 transformation in E.coli. To transform E.coli with these vectors we have to wait for 2 days before seeing cells!! We will check the vectors next week.
- Transformation of B.S. IA771 (glycerol stocks)
- Nothing! Problem with the storage of competent IA771???
August 18th
Test Ery efficiency
Since all Ery plates grew, we want to check the antibiotic stock.
- New Ery plate of IA751 (they should die)
- New LB stock (Ery5) with IA751
Single colony plates
- New plates (Cm5 + Amp100) with single colony of ECE188 (from 14/08), ECE189 (from 13/08) and ECE190 (from 14/08)
- Grow the same single colony into LB with antibiotic
Transformation of glycerol stocks of B.S.
The result from the last transformation of glycerol stocks was negative> We tried again with two different protocols.
- First protocol
- Spin the glycerol stock, throw out the supernatant (just keep about 100μL), add 0.5μL of Medium B
- Incubate about 1h15
- Add 0.6μg of ECE166 (because we managed to transform BS with this vector)
- Incubate 1h30
- Plate on Cm5 plates (DNA less control and transformed cells)
- Second protocol
- Spin the glycerol stock, throw out the supernatant (just keep about 100μL), add 0.5μL of Medium B
- Add 0.6μg of ECE166 (because we managed to transform BS with this vector)
- Incubate 30min
- Plate on Cm5 plates (transformed cells)
- Plate cells after 10min of incubation (without DNA) on blank plates (to see if they are alive)
PA with I13522, E0040 and B0034 Preparation
- R0040 TOP10 miniprep from LB culture done by Kevin
- 5 single colonies of I13522 picked onto Amp100 plate and LB with Amp100 done on Sunday
- Miniprep of the 5 colonies with sinigle colony PCR
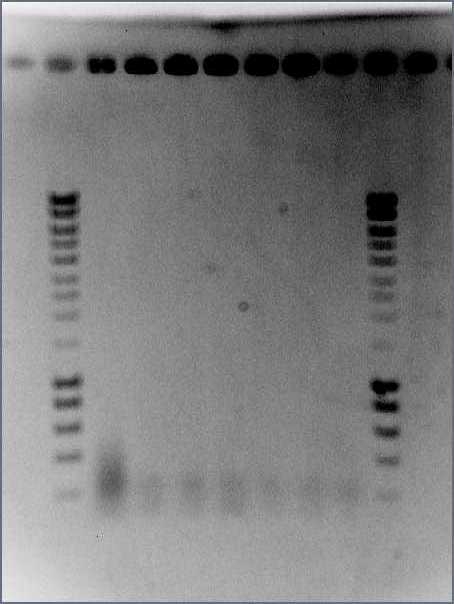
- PCR followed standard protocol and product is run on 0.8% EtBr E-gel
- iGEM34 porgramme is used for PCR
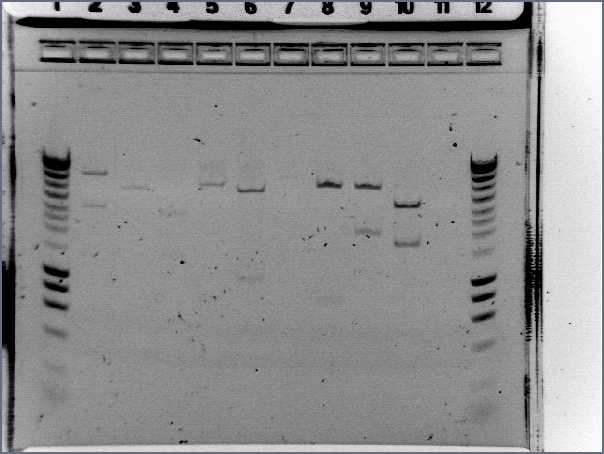
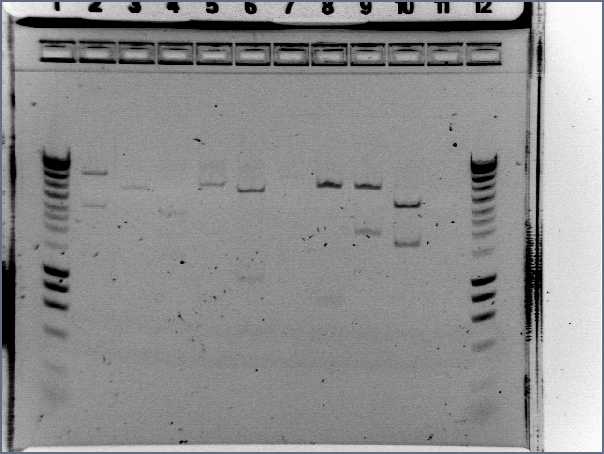
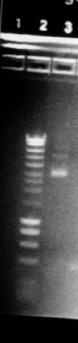
Gel:
- 4μL dye with 20μL PCR product
- 5μL Hyperladder I with 15μL SDW into lane 2
- Lanes 3-7: I13522 Colonies 1-5
- Lane 8: R0040
- Lane 9: Hyperladder I
Result:
- Size of I13522 is not right as the bands are all around 4500bp but it should be 2375bp
- R0040 is of the correct size
- Transformed chemically compotent TOP10 with B0034 biobrick extract and pUC9 control
- Plate on Amp100 plates after standard transformation
August 19th
Results from the transformation with glycerol stocks
We used two different glycerol stocks, one from 07/08 and one from 13/08. We also had 2 different protocols (cf 18/08 page).
We tested our stocks on blank plates, cells are alive, the problem is to know if they are still competent or not.
- Protocol 1 (with incubation time before adding DNA)
- Stock from 07/08 : nothing on the control and nothing on the transformation plate.
- Stock from 13/08 : nothing on the control and nothing on the transformation plate.
- Protocol 2 (without incubation time before adding DNA)
- Stock from 07/08 : nothing on the control and nothing on the transformation plate.
- Stock from 13/08 : nothing on the control, and 1 colony on the transformed plate. We checked the fluorescence of bacteria into this colony (vector ECE166 with promoter and GFP). We have fluorescence, so our transformation is ok.
The result of this confirm that our stock of competent cells in glycerol is still competnet since we managed to transform one colony. The problem is certainly not the storage, but our protocol when we use frozen competent cells> We need to improve the efficiency of this protocol.
Test our Erythromycin stock
- We put some colonies on a Ery 0.5 plate. They grow, but in a very small amount, so our antibiotic seems to be fine.
- Cells in LB with antibiotic : no growth!
So our antibiotic is ok, either we did not transform our cells and that is why everything grew on Ery plates, either there is a problem in our protocol for Ery test.
Plasmid miniprep of ECE153, ECE166 and ECE171
- Plasmid miniprep LB stocks from yesterday
- Nanodrop
| 260/280 | ng/μL | |
|---|---|---|
| ECE 153 | 1.63 | 32.6 |
| ECE 166 (1) | 1.73 | 56.2 |
| ECE 166 (2) | 1.67 | 57.7 |
| ECE 171 | 1.81 | 185.5 |
Transformation of IA771
We used IA751 plate from 12/08/08, vectors ECE 153, 166, 171.
Spectrophotometer : blank made with medium A
| Time (min) | 0 | 20 | 40 | 80 |
|---|---|---|---|---|
| OD650 | 0.1512 | 0.1498 | 0.1476 | 0.1535 |
There was a problem with our colonies. At t=80min, we inoculated again our tube of Medium A.
- We used IA751 plate from 12/08/08, vectors ECE 153, 166, 171
- Spectrophotometer : blank made with medium A
| Time (min) | 0 | 20 | 40 | 60 | 80 | 100 |
|---|---|---|---|---|---|---|
| OD650 | 0.2393 | 0.2504 | 0.2663 | 0.2807 | 0.3184 | 0.3672 |
- Continue the protocol, add DNA (ECE153, 166, 171)
- Plate on antibiotic plates
- New glycerol stocks
- 8 tubes : spin cells, throw away the supernatant, add 500μL of glycerol 60%
- 8 tubes : spin cells, keep only 100μL of medium B, resuspend cells and add 500μL of glycerol 60%
- Put them in the freezer at -80°C
Dealing with R0040, R0062, S0168, I13522 for PA
- Plated R0040, R0062, S0168 received from MIT onto Amp100 plates and incubate at 37°C
- Pciked 4 single colonies for each after colonies have grown and incubate overnight
Check I13522 biobrick DNA
- PCR biobrick DNA from 2007 wells
- Run 0.8% EtBr E-gel
- Lane 5: I13522
- Lane 7: Hyperladder I
Result:
- Size of band is too small under 1000bp should be 2375bp instead
- Transform TOP10 again but with I13522 from 2008 wells
August 20th
Check vector ECE190
- Plasmid miniprep
- Nanodrop
- Double digest : 6μL of SDW, 8μL of DNA, 1μL of buffer, 1μL of XbaI and 1μL of PstI
- Run on a gel
- Result : 2 bands (lane3, a little bit more than 3000b and a little bit more than 5000b) :ok!
August 21st
Check ECE188
- Plasmid miniprep
- Double digest with XbaI and PstI
- Gel (lane6)
- Result : nothing! not enough DNA
August 22nd
Stock ECE112, 166, 153
- Plasmid miniprep (2 tubes of ECE112, 2 tubes of ECE166 and 1 tube of ECE153)
- Nanodrop
Transformation of Bacillus
- For strain IA751 : transform with ECE166, 171, 153 and 112
- For strain IA771 : transform with ECE166, 171
- for glycerol stocks (one with 100μL of medium B and glycerol, and one with only glycerol) : transform with ECE166
- We used the same protocol than before with a few changes :
- add sterile 50% glucose in medium A
- don't keep samples after using them for spectrophotometer
- spin each tubes and keep only 100μL of medium A (to concentrate cells) before adding DNA
| Time (min) | 0 | 20 | 40 | 60 | 80 | 100 | 120 | 130 | 140 |
|---|---|---|---|---|---|---|---|---|---|
| OD650 for IA751 | 0.1817 | 0.1870 | 0.1929 | 0.2115 | 0.2543 | 0.3291 | 0.3695 | 0.4188 | 0.4224 |
| OD650 for IA751 | 0.1818 | 0.1874 | 0.1907 | 0.2088 | 0.2452 | 0.3182 | 0.3575 | 0.4228 | 0.4314 |
- For glycerol stocks, resuspend cells into 100μL of medium B, add DNA and wait for 30min, end plate them
August 23rd
Results of transformation from 23/08/2008
| Vector | Strain | Antibiotic | number of colonies | Conclusion |
|---|---|---|---|---|
| ECE166 | IA771, glycerol stock with medium B | Cm5 | one large colony and a lot of very small colonies | Contamination? |
| Control | IA771, glycerol stock with medium B | Cm5 | a lot of very small colonies | Contamination? |
| ECE166 | IA771, glycerol stock without medium B | Cm5 | many very small colonies | Contamination? |
| Control | IA771, glycerol stock without medium B | Cm5 | a few tiny colonies | Contamination? |
| ECE166 | IA751, fresh | Cm5 | 9 large colonies, no small ones | to test |
| Control | IA751, fresh | Cm5 | no growth, scratches | ok |
| ECE166 | IA771, fresh | Cm5 | 1 large colony, many small worm-like growth, scratches? | to test |
| Control | IA771, fresh | Cm5 | no colonies at all | ok |
| ECE171 | IA771, fresh | Kan5 | no colonies | ?? |
| Control | IA771, fresh | Cm5 | no colonies | ok |
| ECE171 | IA751, fresh | Kan5 | about 30 large colonies and 100 small colonies | to test? |
| Control | IA751, fresh | Cm5 | 3 large colonies, about 20 small to medium colonies | contamination? |
| ECE112 | IA751, fresh | Cm5 + Spc100 | no growth | ?? |
| Control | IA751, fresh | Cm5 + Spc100 | no growth | |
| ECE153 | IA751, fresh | Spc50 | about 500 medium to large colonies | to test |
| Control | IA751, fresh | Spc50 | no growth | ok |
August 26th
Xylose test
- for ECE153
- Prepare 4 tubes with 10mL LB and Spc50, inoculate each tube with a colony from IA751 + ECE153
Starch plate
- prepare some blank agar plates
- add 1g of starch into 100mL of soft agar, mix, boil
- put a thin layer of soft agar with starch
- plate with IA751, IA771, IA751+ECE153 and IA751 (control plate, to see if we mixed some plates)
- Incubate at 37°C
Ery plate
- Plate IA751, IA771, IA751 (control plate) and IA751+ECE153
August 27th
Xylose test
- Preparation of xylose tube
- Add 1mL of culture from yesterday, 1mL of xylose (0.1g/mL) and 8mL of LB and 45μL of Spc (do that for the 4 different colonies)
- Incubate 5hours
- Observation with microscope : some green task, but they do not really match with bacillus cells, not conclusive!
Starch plate
- Add 5mL of iodine solution on starch plate, wait for 1min, throw the liquid
- Result
| Strain | added vector | Observation | Conclusion |
|---|---|---|---|
| IA771 | none | no clearance zones | ok |
| IA751 | none | big clearance zones | ok |
| IA751 | ECE153 | no clearance zones | trnsformation worked! |
| IA751 or IA771 | none or ECE171 | big clearance zones | we have some colonues on control plate with IA751, contamination? or antibiotic resistance? |
Ery test
- IA771 alive and IA751 dead, confirmation that we did not mix plates (same conclusion than with starch plate, last line)
Test glycerol stocks of competent bacillus
- Pellet bacillus cells, add 100μL of prewarmed mediumB, add 0.6μg of DNA, incubate 30min at 37°C by shaking, plate
| Strain | added vector | Antibiotic |
|---|---|---|
| IA771 | none | blank plate (to check they are alive) |
| IA771 | none | Cm5 (negative control) |
| IA771 | ECE166 | Cm5 |
| IA751 | none | blank plate (to check they are alive) |
| IA751 | none | Cm5 (negative control) |
| IA751 | none | Spc50 (negative control) |
| IA751 | ECE112 | Cm5 |
| IA751 | ECE153 | Spc50 |
August 28th
Results from Bacillus transformation with glycerol stocks
| Strain | added vector | Antibiotic | number of colonies | Conclusion |
|---|---|---|---|---|
| IA771 | none | blank plate | confluent colonies | cells are alive |
| IA771 | none | Cm5 | 0 | no contamination |
| IA771 | ECE166 | Cm5 | 2 | transformation ok, very low efficiency |
| IA751 | none | blank plate | confluent colonies | cells are alive |
| IA751 | none | Cm5 | 0 | no contamination |
| IA751 | none | Spc50 | 0 | no contamination |
| IA751 | ECE112 | Cm5 | 0 | problem, we can not manage to transform this vector |
| IA751 | ECE153 | Spc50 | 156 | transformation ok, good efficiency |
Test AmyE insertion
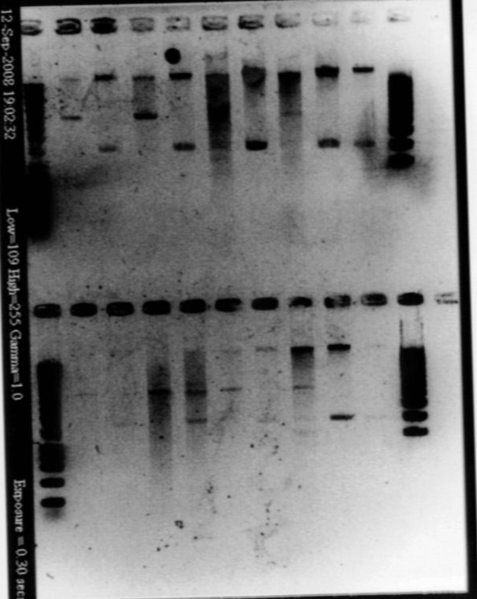
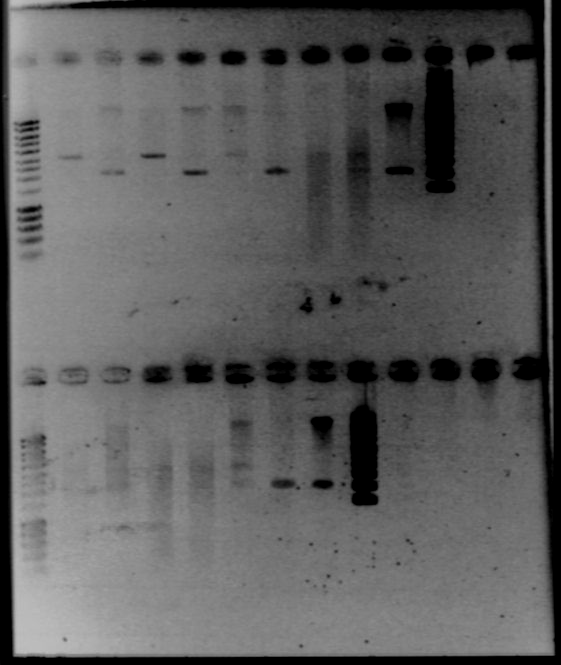
- Colonies PCR for Bacillus (IA751 transformed with ECE153, IA751, and IA771)
- 20μL of lysozyme, a few colonies for each
- Cycle : 15min qt 37°C, 15min at 99°C, 1min at 4°C, 1min at 99°C and 1min at 4°C
- for each PCR : 12μL of cells, 1μL of cells, 1μL of primer1, 1μL of primer2, 5μL of master mix
- PCR (iGEM program)
- run a gel
- Result : nothing on the gel!!! But we can not manage to PCR anything... there is a problem in our PCR protocol, or products!
August 29th
Stock of ECE112
- Bulk up ECE112 (Amp100) rfom 10mL LB (28/08), take 1mL and add to 100mL of LB in big flask
- Grow overnight in 37°C incubator with vigorous shaking
- Aliquot in 10mL tubes, pellet and freeze cell pellets
Transformation of glycerol stock
- Strian : IA751
- Transfer into Eppendorf tubes and spin down for 30min
-Remove glycerol and add medium B
- Add ECE112 and ECE153, and one control tube
- incubate for 30min in 37°C
- Plate out ECE112 (Cm5), ECE153 (Spc50), DNA less (blank, Cm5, Spc50)
September 11th
RBS screening
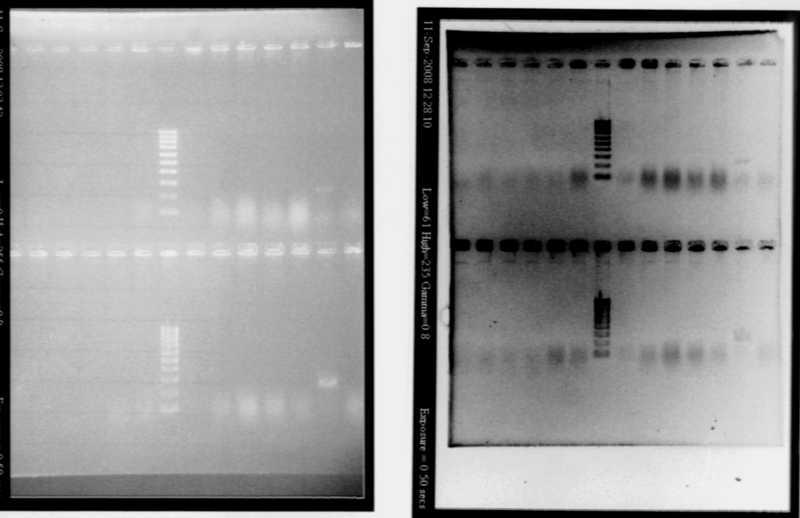
- We run our PCR products from yesterday on a new 1.8% agarose gel
- Gel1
- Lane2 : RBSS1
- Lane3 : RBS S2
- Lane4 : RBS S3
- Lane5 : RBS S4
- Lane6 : RBS S5
- Lane7 : RBS S6
- Lane8 : HyperladderIV
- Lane9 : RBS S7
- Lane10 : RBS S8
- Lane11 : RBS S9
- Lane12 : RBS S10
- Lane13 : RBS S6 with VF and VR primers
- Lane14 : RBS S11
- Gel2
- Lane2 : RBSW1
- Lane3 : RBS W2
- Lane4 : RBS W3
- Lane5 : RBS W4
- Lane6 : RBS W5
- Lane7 : RBS W6
- Lane8 : HyperladderIV
- Lane9 : RBS W7
- Lane10 : RBS W8
- Lane11 : RBS W9
- Lane12 : RBS W10
- Lane13 : RBS W6 with VF and VR primers
- Lane14 : RBS W11
- Result
- Nothing, except for the PCR with VF and VR primers, either our primers are not right, either we have no insert.
RBS screning but single digest
- Grow some colonies of Top10 transformed with RBS into 10mL of LB with antibiotic
We want to check our transformation with single digest. If we have self ligation in our transformation, we will loose the XbaI cutting site.
Backbone for ligation
- New pellets in 600μL of SDW
- Plasmid miniprep
September 12th
RBS screening (single digest)
- Plasmid miniprep 6 colonies of RBS S and 6 colonies of RBS W (without endo wash buffer)
- Nanodrop
| product | concentration (ng/μL) | 260/280 | quantity of DNA to add to have about 300ng (μL) | added SDW for single digest (μL) |
|---|---|---|---|---|
| RBS W1 | 57.1 | 1.57 | 6 | 11 |
| RBS W3 | 54.1 | 1.53 | 6 | 11 |
| RBS W5 | 15.1 | 1.85 | 17 | 0 |
| RBS W 7 | 21.8 | 1.66 | 15 | 2 |
| RBS W9 | 21.8 | 1.78 | 15 | 2 |
| RBS W11 | 13.8 | 1.73 | 17 | 0 |
| RBS S2 | 19.5 | 1.7 | 15 | 2 |
| RBS S4 | 82.13 | 2.57 | 4 | 13 |
| RBS S6 | 44.7 | 1.64 | 7 | 10 |
| RBS S8 | 104.0 | 1.48 | 3 | 14 |
| RBS S10 | 13.1 | 1.82 | 17 | 0 |
| RBS S11 | 20.1 | 1.73 | 15 | 2 |
- Single digest : with EcoRI and XbaI, add SDW and DNA (according to the previous table, 300mg of DNA), 2μL of fast digest buffer, and 1μL of enzyme
- Gel1
- Lane2 : HyperladderI
- Lane3 : RBS W1 cut with EcoRI
- Lane4 : RBS W1 cut with XbaI
- Lane5 : RBS W3 cut with EcoRI
- Lane6 : RBS W3 cut with XbaI
- Lane7 : RBS W7 cut with EcoRI
- Lane8 : RBS W7 cut with XbaI
- Lane9 : RBS W9 cut with EcoRI
- Lane10 : RBS W9 cut with XbaI
- Lane11 : RBS W9 uncut
- Lane12 : supercoiled ladder
- Gel2
- Lane2 : HyperladderI
- Lane3 : RBS S4 cut with EcoRI
- Lane4 : RBS S4 cut with XbaI
- Lane5 : RBS S6 cut with EcoRI
- Lane6 : RBS S6 cut with XbaI
- Lane7 : RBS S8 cut with EcoRI
- Lane8 : RBS S8 cut with XbaI
- Lane9 : RBS S11 cut with EcoRI
- Lane10 : RBS S11 cut with XbaI
- Lane11 : RBS S4 uncut
- Lane12 : supercoiled ladder
- Result
The size of our plasmid is about 3000bp (ok on the gel)
- RBS W
- W1 with EcoRI : cut
- W1 with XbaI : uncut, self ligation
- W3 with EcoRI : cut
- W3 with XbaI : uncut, self ligation
- W7 with EcoRI : impossible to see
- W7 with XbaI : uncut, self ligation
- W9 with EcoRI : cut
- W9 with XbaI : uncut, self ligation
- RBS S
- S4 with EcoRI : cut
- S4 with XbaI : uncut, self ligation
- S6 with EcoRI : cut
- S6 with XbaI : 2 band, this plasmid is partially cut!!! Transformation with RBS S
- S8 with EcoRI : cut
- S8 with XbaI : uncut, self ligation
- S11 with EcoRI : cut
- S11 with XbaI : uncut, self ligation
Double check of RBS S6 and stock
- Grow RBS S6 in LB with Cm35
Check more RBS W
- Grow RBS W2, 4, 6, 8, 10 in 10mL of LB with antibiotic
Check PCR products
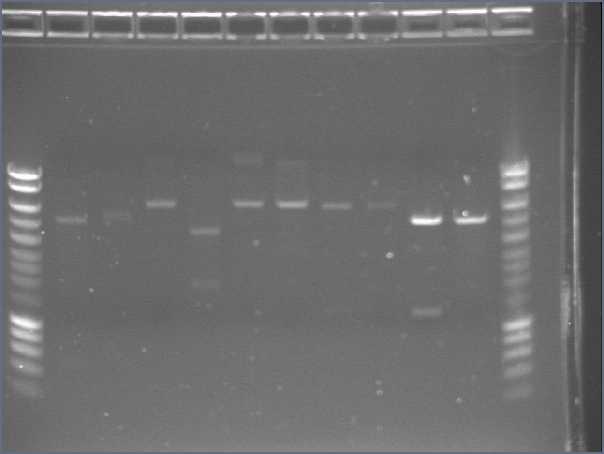
- Gel (2μL of each product)
- Lane1 : λ ladder
- Lane2 : agrA
- Lane3 : agrB
- Lane4 : agrC
- Lane5 : rep
- Lane6 : pSB4C5 1
- Lane7 : pSB4C5 2
- Lane8 : Pxyl
- Lane9 : Ppac
- Lane10 : RBS S
- Lane11 : RBS W
- Lane12 : HyperladderI
- Results
- agr B : 2 bands?
- agrC : ok
- rep : small band but good size
- backbones : ok
- promoters : ok
- RBS : nothing
September 13th
Grow RBS S6
- Put the 10mL of LB with RBS S6 from yesterday into 200mL of LB with antibiotic
- Incubate at 37°C with shaking
RBS W screening
- Plasmid miniprep of RBS W2, 4, 6, 8, 10 9witouh endo wash buffer)
- Nanodrop
| product | concentration (ng/μL) | 260/280 | quantity of DNA to add to have about 300ng (μL) | added SDW for single digest (μL) |
|---|---|---|---|---|
| RBS W2 | 42.4 | 1.62 | 8 | 9 |
| RBS W4 | 35.1 | 1.61 | 10 | 7 |
| RBS W6 | 41.0 | 1.67 | 8 | 9 |
| RBS W8 | 72.8 | 1.61 | 5 | 12 |
| RBS W10 | 51.0 | 1.60 | 6 | 11 |
- Single digest with EcoRI and XbaI
- Gel1
- Lane2 : HyperladderI
- Lane3 : RBS W2 cut with EcoRI
- Lane4 : RBS W2 cut with XbaI
- Lane5 : RBS W4 cut with EcoRI
- Lane6 : RBS W4 cut with XbaI
- Lane7 : RBS W5 cut with EcoRI
- Lane8 : RBS W5 cut with XbaI
- Lane9 : RBS W6 cut with EcoRI
- Lane10 : RBS W6 cut with XbaI
- Lane11 : RBS W8 uncut
- Lane12 : supercoiled ladder
- Gel2
- Lane2 : HyperladderI
- Lane3 : RBS W8 cut with EcoRI
- Lane4 : RBS W8 cut with XbaI
- Lane5 : RBS W10 cut with EcoRI
- Lane6 : RBS W10 cut with XbaI
- Lane7 : RBS W11 cut with EcoRI
- Lane8 : RBS W11 cut with XbaI
- Lane9 : RBS W8 uncut
- Lane10 : supercoiled ladder
- Result
- W2 with EcoRI : cut
- W2 with XbaI : uncut, self ligation
- W4 with EcoRI : cut
- W4 with XbaI : uncut, self ligation
- W5 with EcoRI : cut
- W5 with XbaI : uncut, self ligation
- W6 with EcoRI : cut
- W6 with XbaI : cut (2 bands), transformation ok for RBS W6
- W8 with EcoRI : impossible to see
- W8 with XbaI : impossible to see
- W10 with EcoRI : cut
- W10 with XbaI : maybe cut, it seems to be 2 bands, but quite difficult to see
- W11 with EcoRI : cut
- W11 with XbaI : uncut, self ligation
Double check of RBS W6 and stock
- Grow RBS W6 in 10mL of LB with antibiotic
 "
"