Team:Wisconsin/Project
From 2008.igem.org
| Home | The Team | The Project | Parts Submitted to the Registry | Modeling | Notebook |
|---|
|
| Triose Phosphate Isomerase Knockout |
|
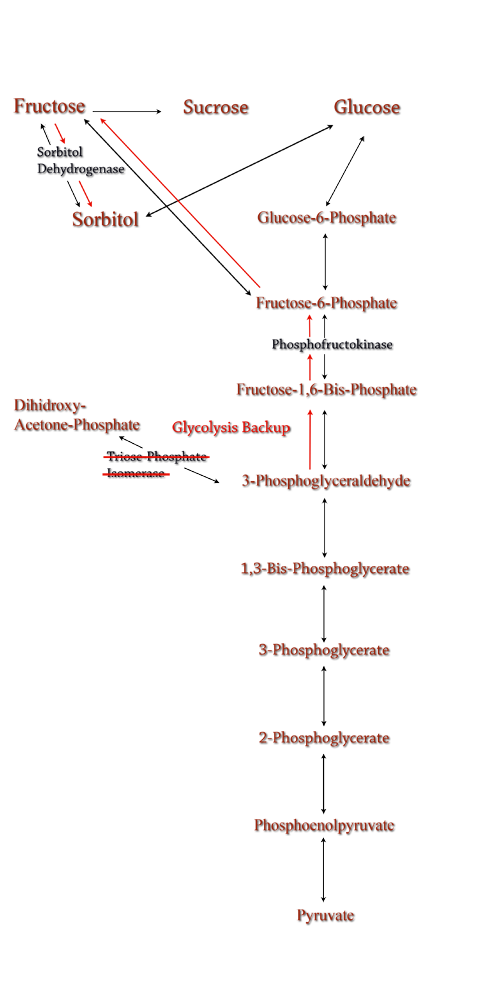
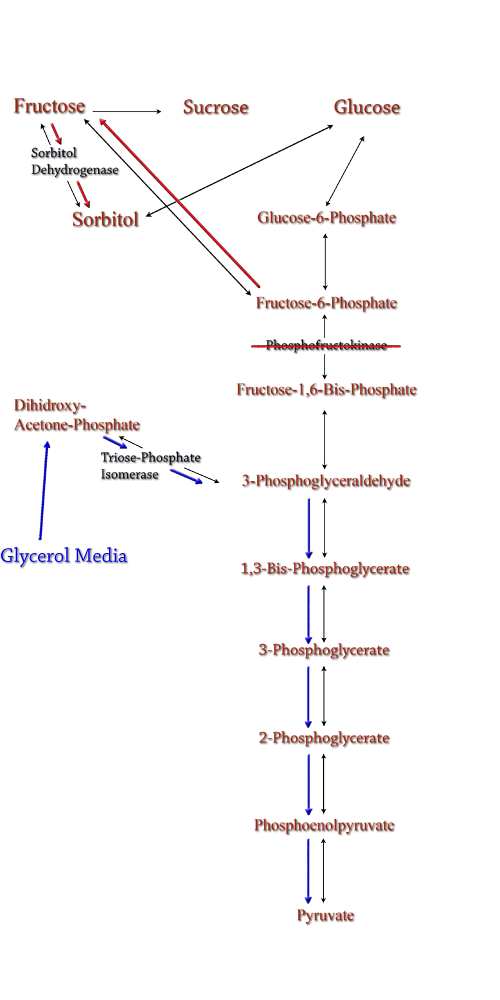
Computer modeling was done to determine gene knockouts that increased cellular flux towards sorbitol production. The model was given to us by Dr. Jenni Reed of University of Wisconsin-Madison. More extensive information about modeling can be found at our modeling page. Through the use of our model we determined that knocking out the gene encoding for triose phosphate isomerase would be the most effective at increasing sorbitol production when combined with srl gene up-regulation. Triose phosphate isomerase(TPI) is another enzyme in E.coli's glycolysis pathway. Its function is to catalyze the interconversion of 3-Phosphoglyceraldehyde to Dihidroxy-acetone-phosphate and back. Knocking out the gene which encodes TPI stops this interconversion, a process which removes products of glycolysis from the cellular reaction mixture shifting the overall reaction downwards towards pyruvate. In essence, knocking out the TPI gene backs up glycolysis and produces a shift towards sorbitol (Pictured right). TPI is a nonessential gene and as such we were able to obtain it from the Keio collection, a collection of non-essential single gene knockouts from the E.coli K-12 derivative strain BW25113. The increased cellular flux modeled in a TPI knockout combined with increased intracellular SDH production from a plasmid containing either the srlD gene or srl operon will hopefully result in a large scale production of cellular sorbitol. Even with the promising modeling results, our group worked on alternative routes for increased cellular sorbitol. One of these routes, knocking out phosphofructokinase, was an additional project that our group undertook.
|
| Phosphofructokinase Knockout |
|
The idea behind a phosphofructokinase (PFK) knockout is to literally stop the glycolysis pathway after fructose-6-phosphate. The backup of metabolic products caused by the knockout should pool and form a favorable pathway for increased sorbitol production. The obvious problem with this procedure is that knocking out a part of the glycolysis pathway under normal circumstances kills the cell. This is also why modeling didn't prove useful as the program showed the cell would be dead. We set out to determine how to bypass this normal cellular function in order to use it to increase sorbitol production. Experiments have been successfully done to create a viable PFK chromosomal knockouts that contain inducible plasmids encoded with the pfk gene. After obtaining this strain we have done countless growth curves with varying media to create ideal conditions for cell growth. After the optimal media is obtained, the addition of a plasmid containing srlD or the srl operon will be introduced to create what we hope will be this metabolic pathway(right). |
 "
"