Data Retrieval and Storage
From 2008.igem.org
(→Perl) |
|||
| (16 intermediate revisions not shown) | |||
| Line 8: | Line 8: | ||
!align="center"|[[Team:Calgary_Software/Team|The Team]] | !align="center"|[[Team:Calgary_Software/Team|The Team]] | ||
!align="center"|[[Team:Calgary_Software/Project|The Project]] | !align="center"|[[Team:Calgary_Software/Project|The Project]] | ||
| - | |||
!align="center"|[[Team:Calgary_Software/Notebook|Notebook]] | !align="center"|[[Team:Calgary_Software/Notebook|Notebook]] | ||
|} | |} | ||
| Line 23: | Line 22: | ||
[[Image:Perl_logo.PNG||thumb|right|160px]] | [[Image:Perl_logo.PNG||thumb|right|160px]] | ||
| - | <div align=justify> | + | <div align=justify>Our first challenge was finding a way to expand the database for EvoGEM. Last year, EvoGEM only had a small database of BioBrick parts, all of which were added manually. Since the iGEM registry consisted of hundreds of parts, manually adding parts was not practical. In addition, more parts were needed to make more sophisticated tests with EvoGEM. Also, we wanted to have some way of comparing the retrieved parts. We needed to answer the following questions about each part: |
| + | * If it is an enzyme, what reactions does it catalyze? | ||
| + | * If it is a molecule, what is its molecular structure, and what are the synonyms for the molecule name? | ||
| - | <div align=justify>Perl is a programming language that is powerful in text processing facilities. Since it uses string matching | + | The answers to these questions would allow EvoGEM to better distinguish between different compounds. How do we accomplish this, though? By creating a Perl script! </div> |
| + | |||
| + | <div align=justify>Perl is a programming language that is powerful in text processing facilities. Since it effectively uses string matching, it is an ideal language for searching text and manipulating text files, which is exactly what we needed for retrieving and expanding EvoGEM's local database.</div> | ||
<br style="clear:both"/> | <br style="clear:both"/> | ||
== UniProt == | == UniProt == | ||
| - | [[Image:UniProt.PNG|left|180px | + | [[Image:UniProt.PNG|thumb|left|180px]] |
<div align=justify> | <div align=justify> | ||
| - | If | + | If a protein makes up one of the parts retrieved from the iGEM database, the registry provides its amino acid sequence, which can be used to infer all other required information. Namely, the program sends this amino acid sequence to UniProt. UniProt is a large database of proteins and enzymes. This database can be queried by a Blast algorithm, which is a very powerful programming tool. When inputting the DNA or amino acid sequence, UniProt gives results that are closest to the initial search. Besides giving the name of the protein searched, UniProt will give the reagents from the reaction that the protein catalyzes. All this information is useful for EvoGEM and is stored in a local database. Visit [http://www.uniprot.com Uniprot] to see this database. </div> |
<br style="clear:both"/> | <br style="clear:both"/> | ||
== ChemSpider == | == ChemSpider == | ||
| - | [[Image:ChemSpider.PNG|right|180px | + | [[Image:ChemSpider.PNG|thumb|right|180px]] |
<div align=justify> | <div align=justify> | ||
| - | + | The reagents from the reaction that the protein catalyzes are put through ChemSpider. This large database is much like UniProt except that it is for chemistry. Searching and querying in ChemSpider is simple because molecules can be queried using synonyms. After a molecule is queried, ChemSpider produces information about the molecule such as its SMILES, which is a simplified molecular input line entry specification. As useful as this information can be, we needed something that is machine-readable and that could be used for comparisons of metabolic pathways. What is this machine readable format? It is known as the IUPAC International Chemical Identifier (InChI). This InChi is a unique "fingerprint" of the molecule that is not ambiguous like SMILES and is supplied only by IUPAC. An example of an InChi would look like this: | |
'''1/C6H8O6/c7-1-2(8)5-3(9)4(10)6(11)12-5/h2,5,7-8,10-11H,1H2/t2-,5+/m0/s1''' | '''1/C6H8O6/c7-1-2(8)5-3(9)4(10)6(11)12-5/h2,5,7-8,10-11H,1H2/t2-,5+/m0/s1''' | ||
| - | To see this database, | + | To see this database, visit: [http://www.chemspider.com ChemSpider] |
<br style="clear:both"/> | <br style="clear:both"/> | ||
| Line 53: | Line 56: | ||
<div align=justify> | <div align=justify> | ||
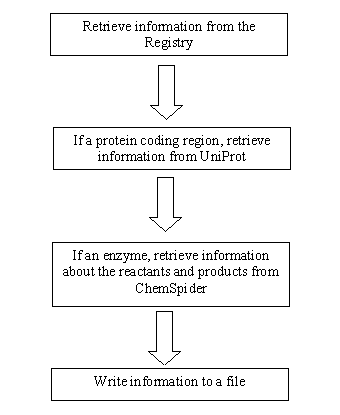
| - | The Perl script’s algorithm works in the following manner | + | The Perl script’s algorithm works in the following manner: First, the program goes to the iGEM registry and retrieves one of the parts, recording its name, type, and sequence. Then, if the part is a protein, it sends the information to Uniprot, where it undergoes the Blast algorithm. From there, it extracts the names of reactants and products and stores them in a file for EvoGEM's local database. Afterwards, if the protein catalyzes a reaction, the program searches for the catalzed compounds in ChemSpider. There, it retrieves more information, such as the InChi, in a local database. Consequently, we now have a large database ready for use for EvoGEM. </div> |
| - | [[Image:Retrieval flow chart.PNG|thumb|center|200px| | + | [[Image:Retrieval flow chart.PNG|thumb|center|200px| Data Retrieval Flow Chart]] |
== Navigation == | == Navigation == | ||
| Line 73: | Line 76: | ||
!align="center"|[[Team:Calgary_Software/Team|The Team]] | !align="center"|[[Team:Calgary_Software/Team|The Team]] | ||
!align="center"|[[Team:Calgary_Software/Project|The Project]] | !align="center"|[[Team:Calgary_Software/Project|The Project]] | ||
| - | |||
!align="center"|[[Team:Calgary_Software/Notebook|Notebook]] | !align="center"|[[Team:Calgary_Software/Notebook|Notebook]] | ||
|} | |} | ||
Latest revision as of 02:22, 30 October 2008

|
| Home | The Team | The Project | Notebook |
|---|
| Evolutionary Algorithm | Data Retrieval | Modeling | Graphical User Interface |
|---|
Contents |
Perl
- If it is an enzyme, what reactions does it catalyze?
- If it is a molecule, what is its molecular structure, and what are the synonyms for the molecule name?
UniProt
ChemSpider
The reagents from the reaction that the protein catalyzes are put through ChemSpider. This large database is much like UniProt except that it is for chemistry. Searching and querying in ChemSpider is simple because molecules can be queried using synonyms. After a molecule is queried, ChemSpider produces information about the molecule such as its SMILES, which is a simplified molecular input line entry specification. As useful as this information can be, we needed something that is machine-readable and that could be used for comparisons of metabolic pathways. What is this machine readable format? It is known as the IUPAC International Chemical Identifier (InChI). This InChi is a unique "fingerprint" of the molecule that is not ambiguous like SMILES and is supplied only by IUPAC. An example of an InChi would look like this:
1/C6H8O6/c7-1-2(8)5-3(9)4(10)6(11)12-5/h2,5,7-8,10-11H,1H2/t2-,5+/m0/s1
To see this database, visit: ChemSpider
The Algorithm
| Evolutionary Algorithm | Data Retrieval | Modeling | Graphical User Interface |
|---|
| Home | The Team | The Project | Notebook |
|---|
 "
"