Team:Caltech/Project/Vitamins
From 2008.igem.org
|
People
|
{{{Content}}} |
</div>
Vitamin productionBackground Information on FolateFolate, the generic term for the various forms of Vitamin B9, is an essential vitamin because it is heavily involved in amino acid synthesis as well as single-carbon transfer reactions. Folate deficiencies in women can result in birth defects such as neural tube defects and other spinal cord malformations. As important as folate is, humans are unable to produce folate, and so must obtain it from eating foods such as green leafy vegetables or folate-fortified cereals sybesma1. An engineered strain of bacteria that would constantly release folate into the gut would reduce the need to fortify breads and cereals with folate, as well as reduce folate-related birth defects in regions with little access to folate-containing foods. In addition to all the reasons stated above, folate is an ideal vitamin to be produced in the gut because, unlike many other vitamins, it has been shown to be absorbed in physiologically relevant quantities in the large intestineasrar. Structurally, folate consists of three main parts: pteridine, p-aminobenzoic acid (pABA), and L-glutamate.
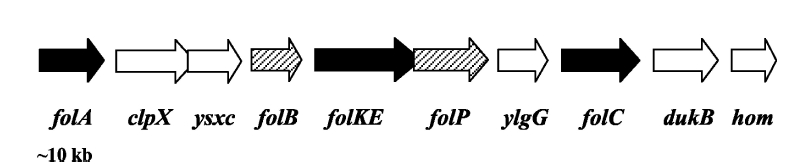
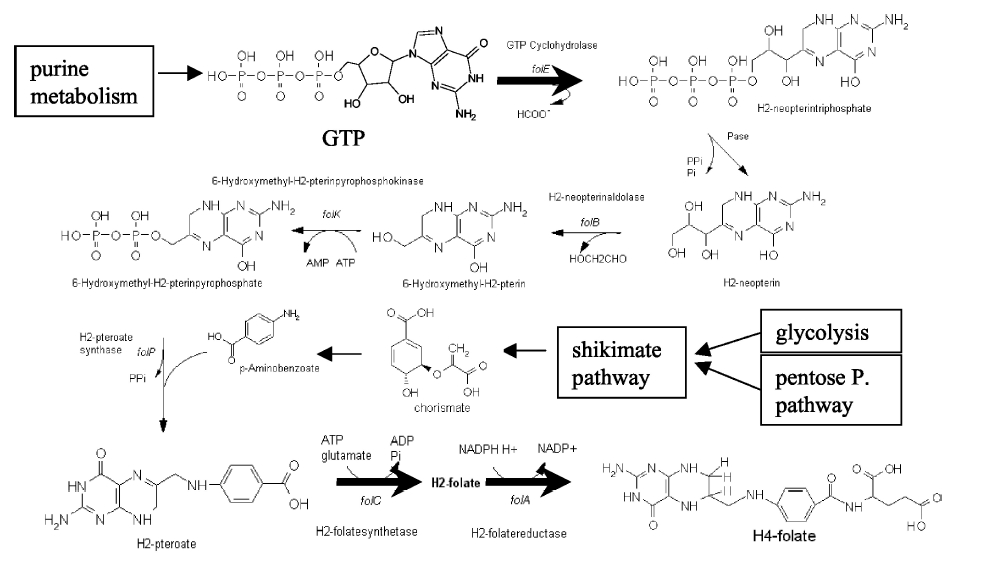
Folate Biosynthesis PathwayAlthough folate is naturally produced in E.coli, the folate biosynthesis pathway in the bacteria Lactococcus lactis has been more heavily characterized and studied. There are six major enzymes involved in folate synthesis, which, in L.lactis, are contained in five genes: folB, folKE, folP, folC, and folAsybesma1. The first four, which we have chosen to focus on, are located in a gene cluster approximately 4.4kb long. We’ve chosen not to focus on folA for the time being because folA encodes an enzyme which turns one form of folate (dihydrofolate) into another form of folate(tetrahydrofolate). Since our assay would detect both types of folate as part of the total folate production, folA was not a prime target for overexpression of folate. In previous studies, this folate gene cluster has been successfully transformed into the folate-consuming bacteria L.gasseri, turning the bacteria in to folate-producerswegkamp1. Therefore, we have chosen to also use the folate operon from L.lactis, which also offers the additional benefit of removing the operon from its natural regulatory context. Our strategy is to clone the entire folate operon from the L.lactis genome and to transform the entire operon into E.coli. However, because we are unsure of whether or not the ribosomal binding sites (RBS) within the L.lactis operon would work in E.coli, we are also going to unpack the operon by cloning each of the four genes individually, placing them behind E.coli RBSs, and then recombine them into a single empty BioBricks™ plasmid. In addition to the main folate operon, we will also be experimenting with overexpression of the para-aminobenzoic acid (pABA) synthesis pathway from chorismate. Wegkamp et al. have shown that in order to increase overall total levels of folate, both the pABA synthesis genes and certain folate production genes need to be overexpressedwegkamp2. The pABA pathway involves three genes, pabA, pabB, and pabC – though in L.lactis, pabB is actually a fusion gene encoding for both pabB and pabCwegkamp2. System DesignThe overall system design for testing folate production in E.coli is to construct two plasmids – one for the folate biosynthesis pathway, and one for the pABA synthesis pathway. In addition to ensuring that the plasmids are complementary, each plasmid would need to contain a different variable copy origin of replication, which would be low copy by default, but can be switched to high copy via the addition of Isopropyl β-D-1-thiogalactopyranoside (IPTG)to the media. This will allow us to test overexpression of each plasmid separately. In addition, each plasmid will contain a constitutive promoter, since we would want folate to be produced constantly. The purple dots represent ribosomal binding sites (RBS), followed by the gene (green arrow), and eventually terminating in a double stop (TAATAA) sequence, as regulated by the Registry of Standard Biological Parts. Folate Detection MethodsWe will be detecting folate production, and thus the relative success of our engineering efforts, via a microbiological assay involving the folate-dependent strain Enterococcus hiraehorne. This assay involves first the characterization of a standard growth curve of E. hirae given known quantities of folate present in the growth media. Once the standard curve has been established, then experimental growth levels, as quantified by spectrophotometry, can be interpolated. PABA concentrations will be measured via high performance liquid chromatography (HPLC) wegkamp2. Folate microbiological assay protocol para-aminobenzoic acid (pABA) HPLC protocol Relevant Registry PartsBasic Parts
Construction Intermediates
Composite Parts
References<biblio>
</biblio> |
 "
"