Team:Cambridge/Bacillus
From 2008.igem.org
| Line 20: | Line 20: | ||
=Creating new Biobrick-compatible ''B.subtilis'' vectors= | =Creating new Biobrick-compatible ''B.subtilis'' vectors= | ||
| - | + | [http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2008&group=Cambridge Vectors submitted:] | |
We are using the In-Fusion™ PCR method from ClonTech to create new Biobrick-compatible integrative and episomal vectors. These vectors will allow us to insert Biobricks into the ''Bacillus subtilis'' genome at the AmyE locus and as a 3-5 copy plasmid that will replicate autonomously in ''Bacillus''. | We are using the In-Fusion™ PCR method from ClonTech to create new Biobrick-compatible integrative and episomal vectors. These vectors will allow us to insert Biobricks into the ''Bacillus subtilis'' genome at the AmyE locus and as a 3-5 copy plasmid that will replicate autonomously in ''Bacillus''. | ||
| Line 45: | Line 45: | ||
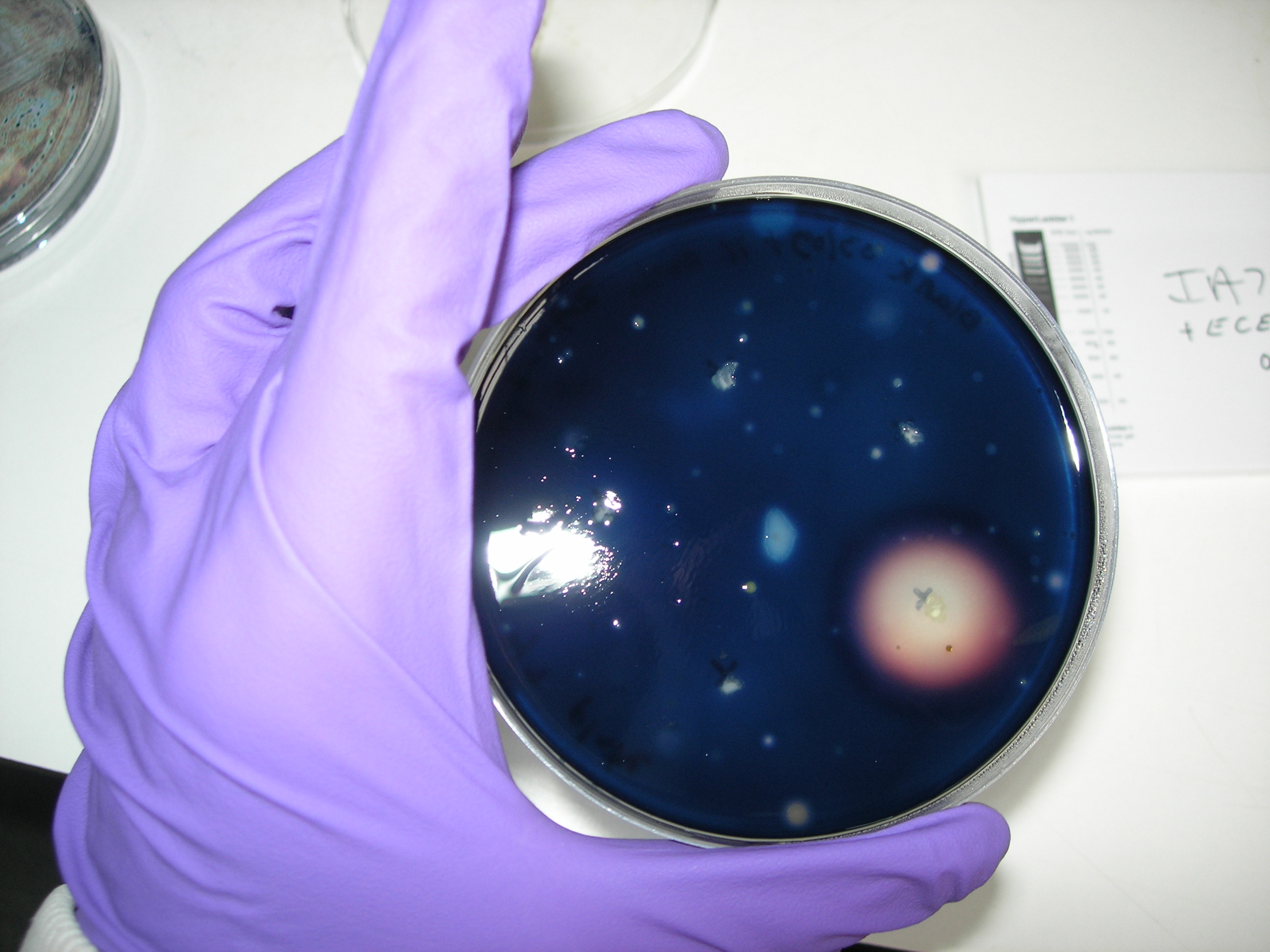
[[Image: 112Amylase.jpg | thumb | 200px | center | IA751 transformed with ECE112 : SUCCESFUL TRANSFORMATION for 4 colonies out of 5, no zone of clearing ]] | [[Image: 112Amylase.jpg | thumb | 200px | center | IA751 transformed with ECE112 : SUCCESFUL TRANSFORMATION for 4 colonies out of 5, no zone of clearing ]] | ||
| - | + | B. subtilis Promoters Designed: | |
| - | + | [http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2008&group=Cambridge Biobricks submitted:] | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
* Pupp - a strong constitutive ''Bacillus cereus'' promoter present on ''B. subtilis'' vector ECE166 | * Pupp - a strong constitutive ''Bacillus cereus'' promoter present on ''B. subtilis'' vector ECE166 | ||
* Ppac - constitutive, present on ''B. subtilis'' vector ECE151 | * Ppac - constitutive, present on ''B. subtilis'' vector ECE151 | ||
| Line 76: | Line 69: | ||
# Transform ''B. subtilis'' with the plasmid which contains amyE for integration and lacZ for beta-galactosidase assay | # Transform ''B. subtilis'' with the plasmid which contains amyE for integration and lacZ for beta-galactosidase assay | ||
# Perform beta- galatosidase assay! | # Perform beta- galatosidase assay! | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
<html> | <html> | ||
Revision as of 03:33, 30 October 2008
|
x
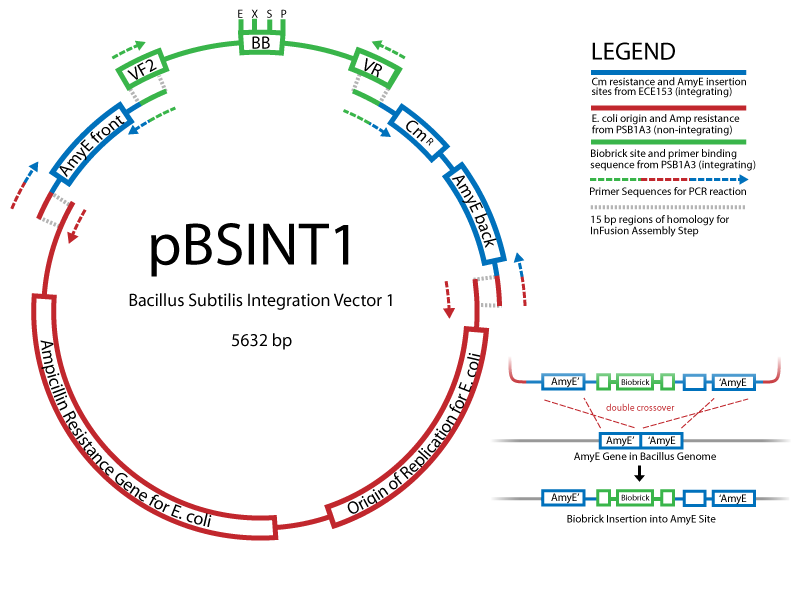
Working with B.subtilisBuilding on the work of the last year's Cambridge iGEM team, we are exploring applications of the gram-positive chassis B.subtillis. Easy to handle and transform, this bacterium is much better to use than E.coli wherever protein import/export is concerned, e.g. in our signalling project. An important part of our effort is to establish standard protocols and parts to work in B.subtillis, characterise control elements, and develop new vectors. You can find our Bacillus protocols here. Creating new Biobrick-compatible B.subtilis vectorsVectors submitted: We are using the In-Fusion™ PCR method from ClonTech to create new Biobrick-compatible integrative and episomal vectors. These vectors will allow us to insert Biobricks into the Bacillus subtilis genome at the AmyE locus and as a 3-5 copy plasmid that will replicate autonomously in Bacillus.
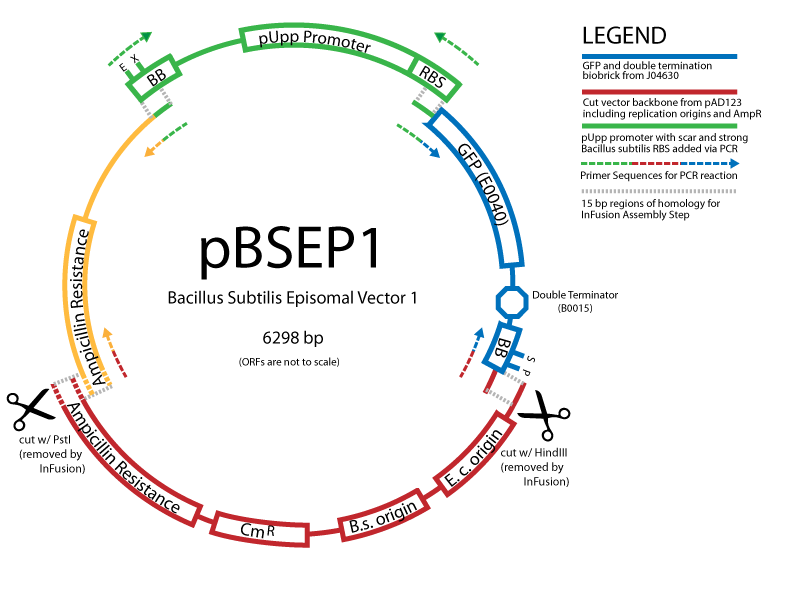
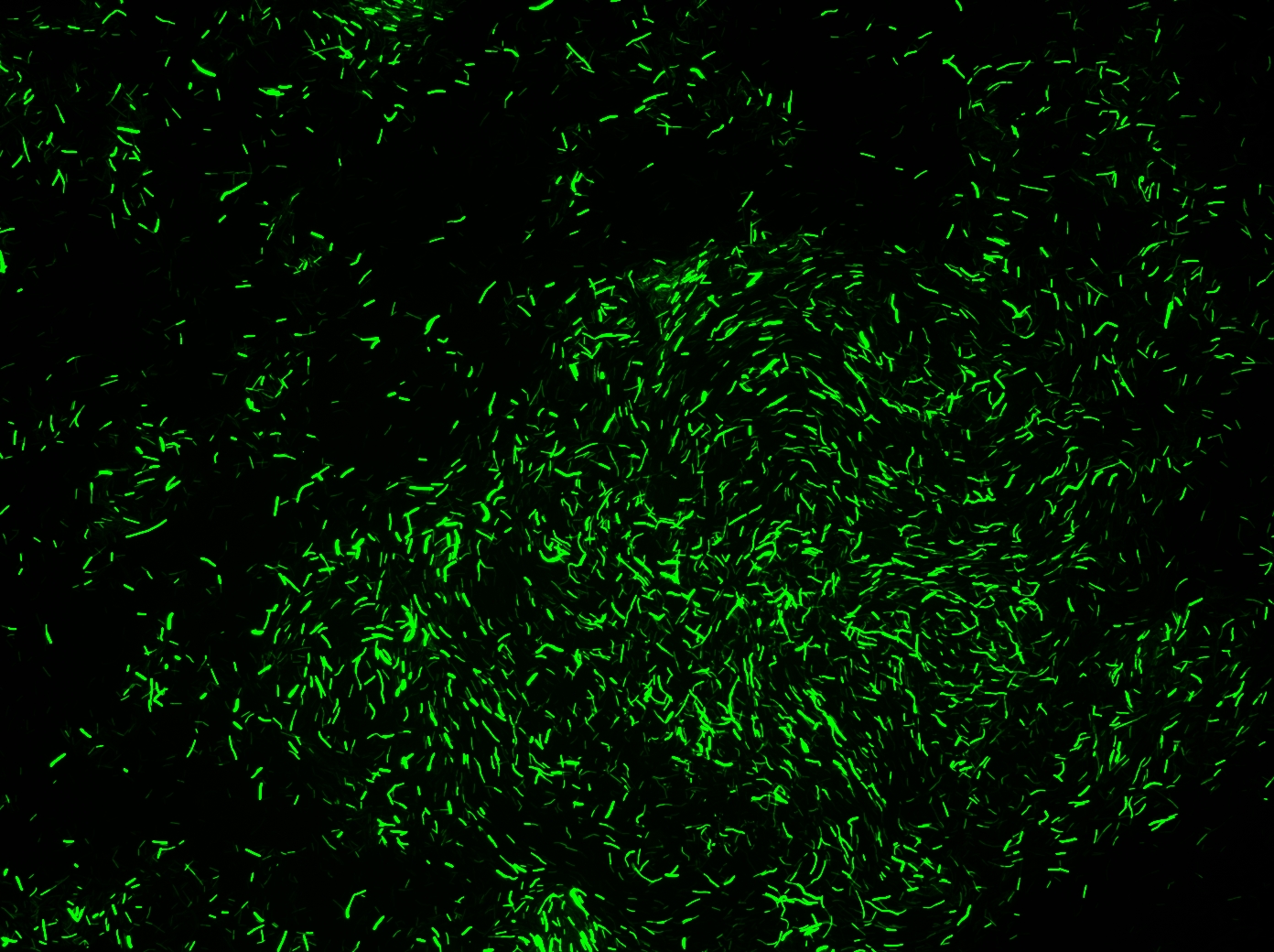
Successful transformation of BacillusTransformation with episomal vectorsECE166 : High-level constitutive expression of a Green Fluorescent Protein (with the promoter Pupp) Transformation with integration vectorsECE153 and ECE112 : Transformation in IA751 : Amylase test B. subtilis Promoters Designed: Biobricks submitted:
Information about the B. subtilis vectors can be found here The Schematic:
x
|
 "
"