Team:ETH Zurich/Modeling/Genome Static Analysis
From 2008.igem.org
Restriction Enzyme AnalysisThis section presents the computational investigation we performed in order to understand which restriction enzymes are optimal when used in our minimal genome approach, in order to find out which restriction enzymes cut the genome in fragments that most probably will lead to find the minimal genome in our reduction approach. Important is to note that this is a "statical" analysis, meaning that we do not include in the evaluation of the restriction enzyme optimality any prevision regarding to the effects its cutting patterns can have on cell physiology or cell system behaviour. We addressed questions regarding the cell system response after genome reduction using more advanced modelling techniques (a genome scale model) in the Genome Scale Analysis section. We focus here only on the insights that can be obtained using three kinds of "statical" information:
Using computational tools and the above mentioned information we are interested in asking (and answering) the following questions:
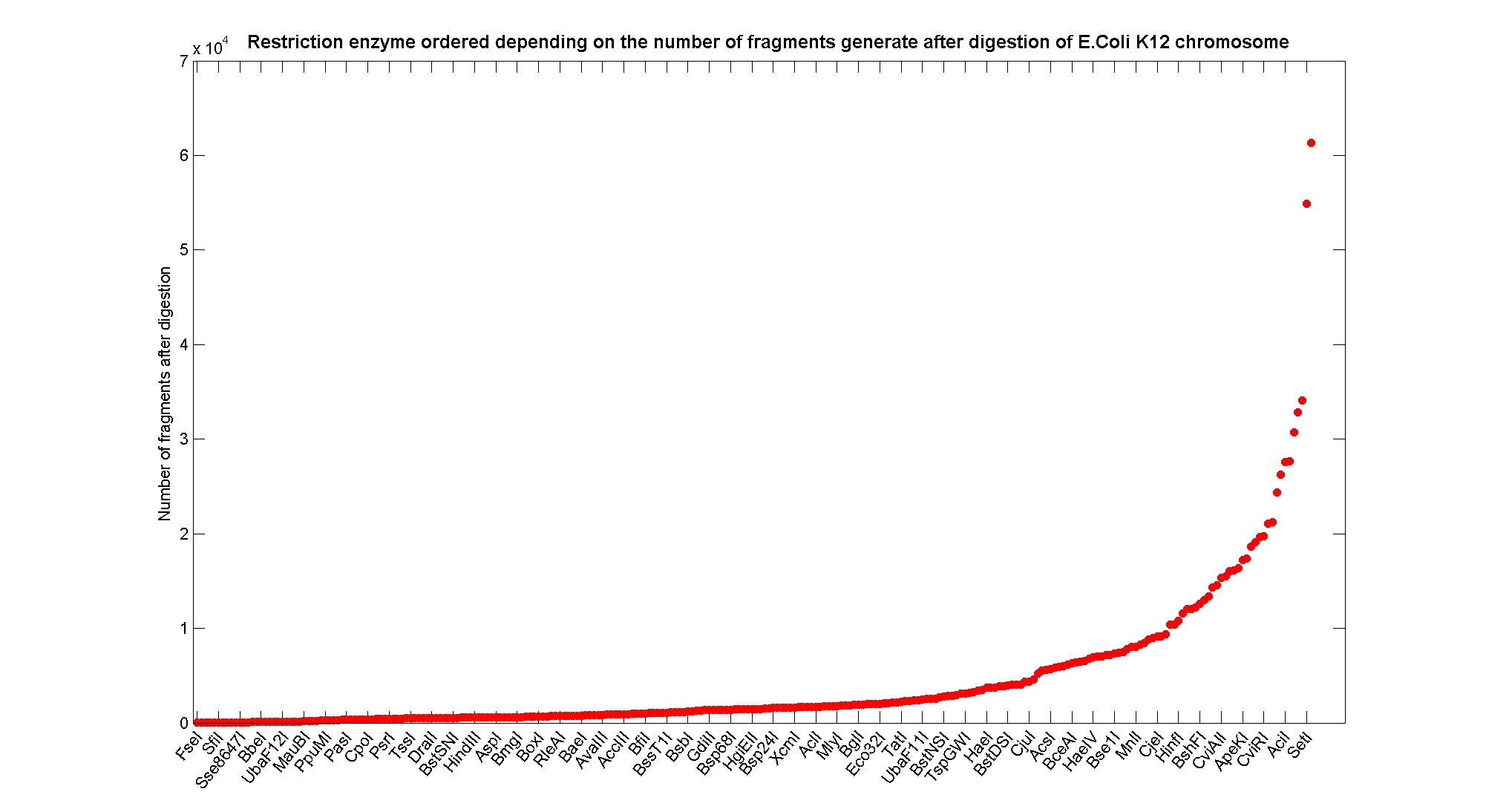
Available restricion enzymes and digestion simulationAs source for the restriction enzyme to consider, we used the REBASE database. We found 713 restriction enzymes that spawn from 4 up to 13 cutters, some with complete specific recognition sites and some with unspeficisity properties. Since some of the restriction enzymes present the same recognition site sequence, we grouped them together as a single entity to be tested (216 groups). We downloaded the genome and annotation information regarding E.Coli K12 MG1655 from GenBank® database. We then simulated the digestion of E.Coli chromosome sequentially for each group of restriction enzymes and performed statistical analysis on the fragment pattern obtained. The following pictures summarize the distribution of the available enzymes regarding to their frequency of cutting (number of fragments after digestion): It is possible to note that there is a huge number of restriction enzymes that digest the chromosome in few to high number of fragments (up to 10000 fragments) and relatively fewer that generate a very high number of fragments. Analysing the gene content of the fragmentsResult TableUsing our digestion simulation code (that can be downloade from our donwload page) we produced a table with the statistic data for each and all the restriciton enzyme. The complete table can be consulted here. |
 "
"