Team:Newcastle University/Notebook
From 2008.igem.org
| Line 5: | Line 5: | ||
==Introduction== | ==Introduction== | ||
| - | Our | + | |
| + | The overall design was too complicated for the time and resources available so we devised a proof of concept Brick which represented an input node from our full design. This Brick demonstrates that it is possible to transfer two-component peptide sensing from one strain into another and have it function as it did in the first strain. We took the subtilin two component system from ''Bacillus subtilis'' ATCC6633 and integrated it into the chromosome of ''B. subtilis'' 168, the typical laboratory ''B. subtilis'' strain at the ''amyE'' locus. | ||
| + | |||
| + | The wet lab work involved: | ||
| + | * Cloning the synthesized Brick from pUC57 in ''E. coli'' into our ''Bacillus'' integration vectors (with GFP and mCherry respectively downstream of the Brick) | ||
| + | * Validation of the integration vector | ||
| + | * Transformation and integration into the chromosome of ''B. subtilis'' 168 at the ''amyE'' locus | ||
| + | * Validation of chromosomal integration | ||
| + | * Characterisation of the integrated parts and their response to subtilin. | ||
| + | |||
| + | Our subtilin sensor, part [http://partsregistry.org/Part:BBa_K104001 BBa_k104001] | ||
[[Image:biobrickk.gif|thumb|center|300px|Figure 1: Newcastle's iGEM team construct.]] | [[Image:biobrickk.gif|thumb|center|300px|Figure 1: Newcastle's iGEM team construct.]] | ||
| Line 16: | Line 26: | ||
* ''gfp'' (green fluorescence protein) - the marker being used to show activation of the ''spaRK'' system and therefore diagnosis of gram-positive bacteria by ''B. Subtilis'' | * ''gfp'' (green fluorescence protein) - the marker being used to show activation of the ''spaRK'' system and therefore diagnosis of gram-positive bacteria by ''B. Subtilis'' | ||
* ''spaS'' promotor - a strong promotor inducible by upstream activation of ''spaRK''. It can be placed in front any gene to regulate its activity. | * ''spaS'' promotor - a strong promotor inducible by upstream activation of ''spaRK''. It can be placed in front any gene to regulate its activity. | ||
| + | |||
| + | |||
| + | === Subcloning === | ||
| + | |||
'''Aim 1: clone the ''spaRK'' system from pUC57 into pJWV021 and transform into DH5 alpha competent ''E. coli''''' | '''Aim 1: clone the ''spaRK'' system from pUC57 into pJWV021 and transform into DH5 alpha competent ''E. coli''''' | ||
Revision as of 15:14, 28 October 2008
Newcastle University
GOLD MEDAL WINNER 2008
| Home | Team | Original Aims | Software | Modelling | Proof of Concept Brick | Wet Lab | Conclusions |
|---|
Home >> Wet Lab
Introduction
The overall design was too complicated for the time and resources available so we devised a proof of concept Brick which represented an input node from our full design. This Brick demonstrates that it is possible to transfer two-component peptide sensing from one strain into another and have it function as it did in the first strain. We took the subtilin two component system from Bacillus subtilis ATCC6633 and integrated it into the chromosome of B. subtilis 168, the typical laboratory B. subtilis strain at the amyE locus.
The wet lab work involved:
- Cloning the synthesized Brick from pUC57 in E. coli into our Bacillus integration vectors (with GFP and mCherry respectively downstream of the Brick)
- Validation of the integration vector
- Transformation and integration into the chromosome of B. subtilis 168 at the amyE locus
- Validation of chromosomal integration
- Characterisation of the integrated parts and their response to subtilin.
Our subtilin sensor, part BBa_k104001
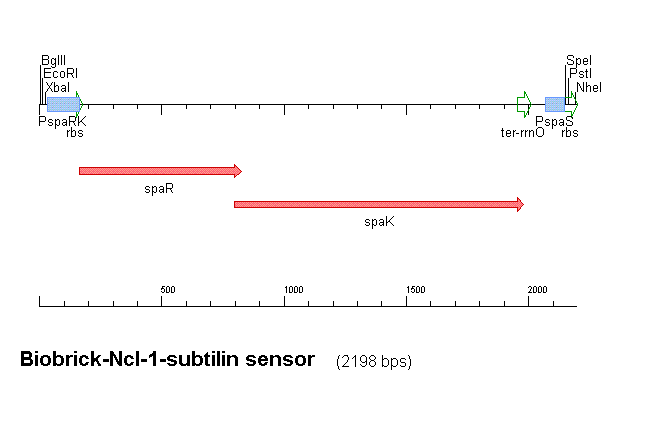
Fig 1 shows the construct which contains:
- spaRK promotor
- rrnB - rRNA binding site
- spaR (subtilin peptide antibiotic Regulation) - the 220 amino acid product of this gene usually regulates the downstream production of subtilin antibiotic. It has an N-terminal domain that can be phosphorylated and a C-terminal domian that has DNA binding properties [1]
- spaK (subtilin peptide antibiotic Kinase) - this gene codes for a 325 amino acid histadine kinase peptide that phosphorylates the N-terminus of spaR [2]. This activates the DNA binding ability of the C-terminus of spaR, which in turn initiates transcription of the downstream gene. In the case of our construct, this gene is gfp.
- gfp (green fluorescence protein) - the marker being used to show activation of the spaRK system and therefore diagnosis of gram-positive bacteria by B. Subtilis
- spaS promotor - a strong promotor inducible by upstream activation of spaRK. It can be placed in front any gene to regulate its activity.
Subcloning
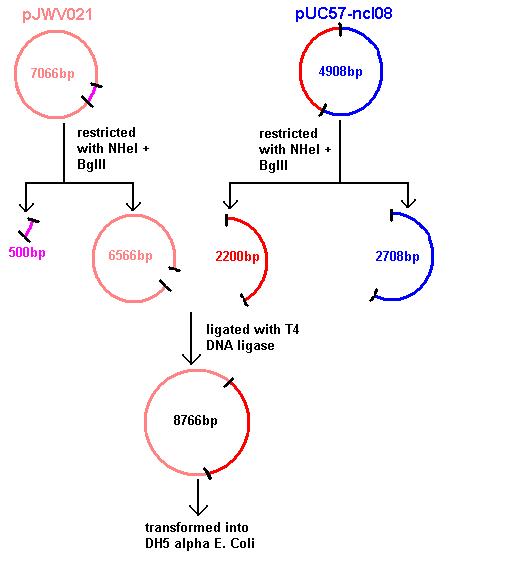
Aim 1: clone the spaRK system from pUC57 into pJWV021 and transform into DH5 alpha competent E. coli
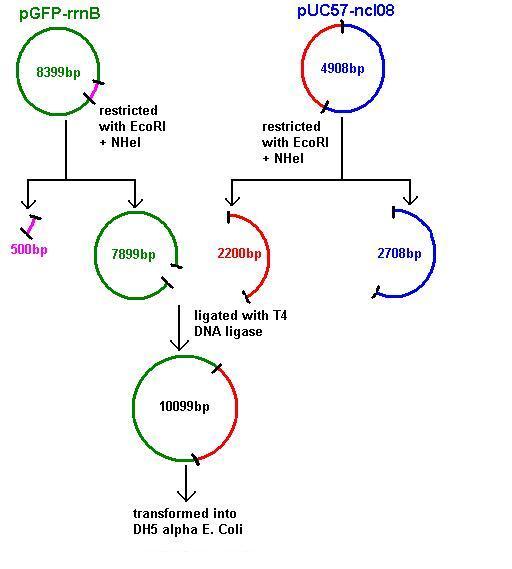
Aim 2: clone the spaRK system from pUC57 into pGFP-rrnB and transform into TOP10 competent E. coli
- The 2.2kb fragment (ncl08) contains the spaRK system and promotor-less gfp linked to this. This means that when spaR is activated, its positive regulatory effect on spaK will in turn activate gfp.
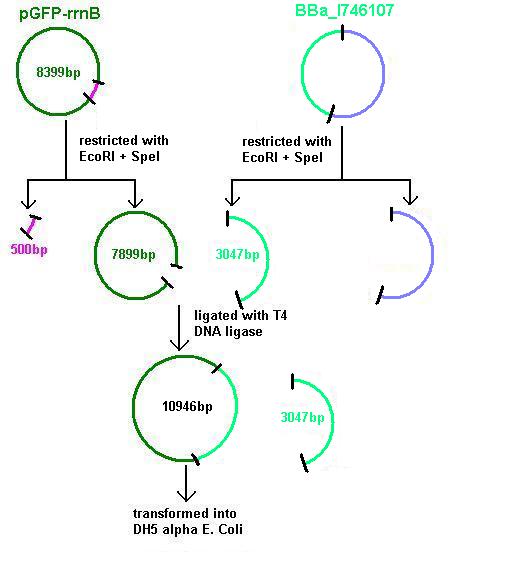
Aim 3: clone the agr system from the plasmid vector into pGFP-rrnB and transform into DH5 alpha competent E. coli
- BBa_I647107 contains a partial agr operon, which includes agrC and A but has agrB and D deleted. This allows coding for the receptor (C/A) but not for production of the quorum peptides themselves (B/D). Eventually this will be linked to the spaRK system in a B. subtilis vector.
Results
To analyse the results of the wet lab transformations of the inserts into B. subtilis, we used two methods: microscopy and flow cytometry.
Microscopy
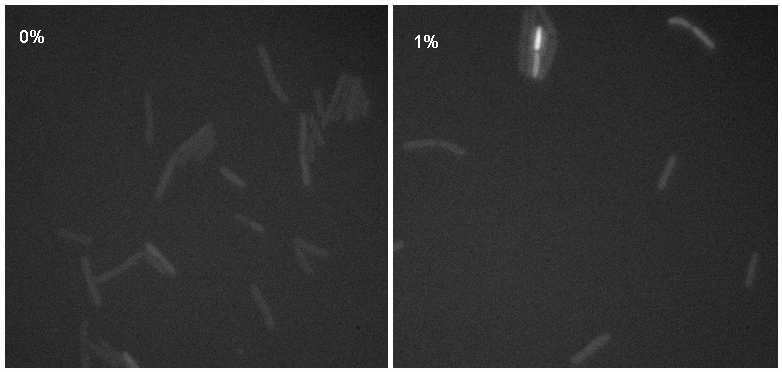
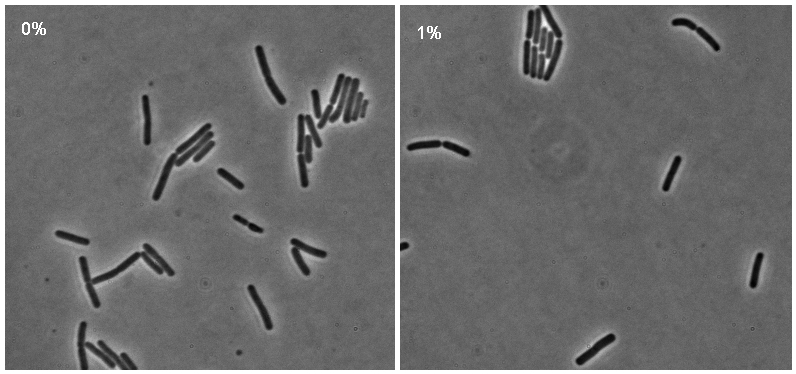
Microscopy work from 08.09.08 showed a difference in the level of flourescence of the iGEMgfp fluorescent cells (higher in 10% subtilin-induced cells compared to 0% subtilin-induced cells). However, there was little difference in the number of cells that fluoresced between the two cultures.
There was no difference in the number of fluorecent cells or the level of fluorescence between the 10% subtilin-induced and the 0% subtilin-induced iGEMcherry cells.
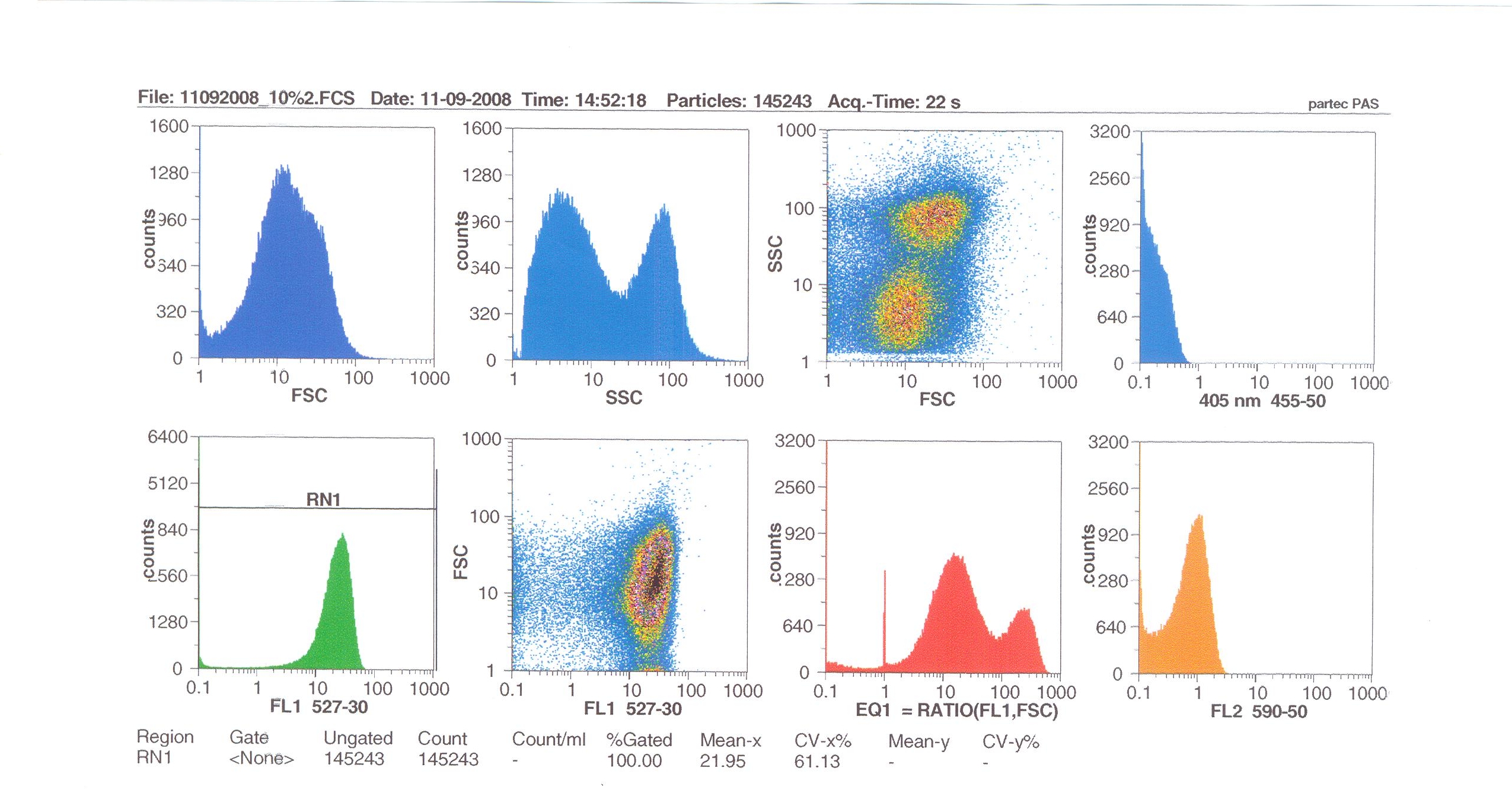
Flow cytometry
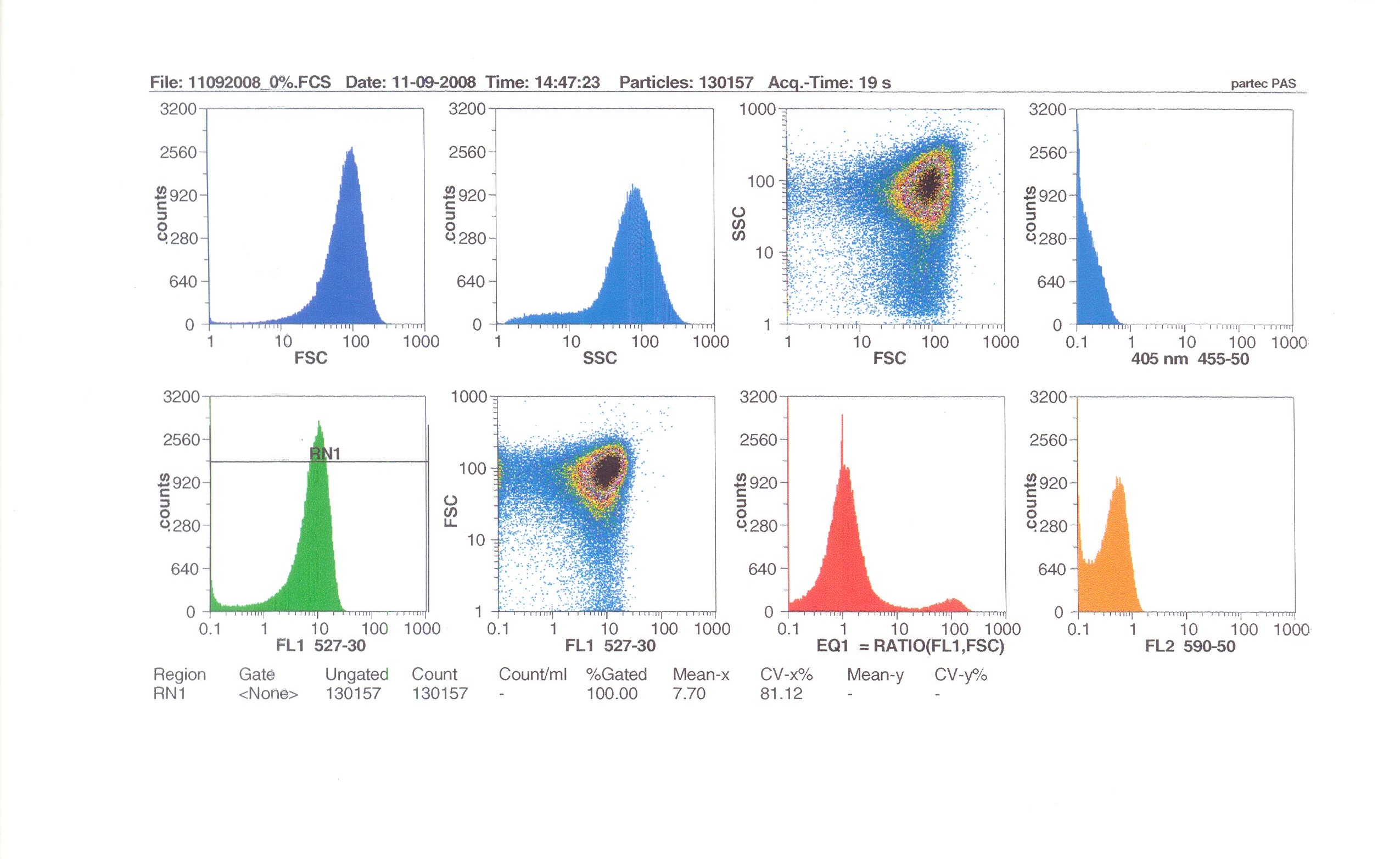
Flow cytometry measures cell fluorescence and light scattering, cell-by-cell, allowing us to quantify our results and present them in graphical form. A sample of our cells engineered Bacillus subtilis cells were injected into the machine, which hydro-dynamically focusses the fluid. Lasers of a particular wavelength are directed onto the stream of fluid, and each particle which passes through the light beam will cause the laser to scatter in a different way. Cells which absorb and then re-emit the light, result in fluorescence (a higher energy state).
The detectors in the machine measure the scattering of light and any fluoresence which occurs, and the data is displayed as histograms and dot plots.
Induction by 0% v/v subtilin (i.e in the absence of subtilin): mean fluoresence = 7.70
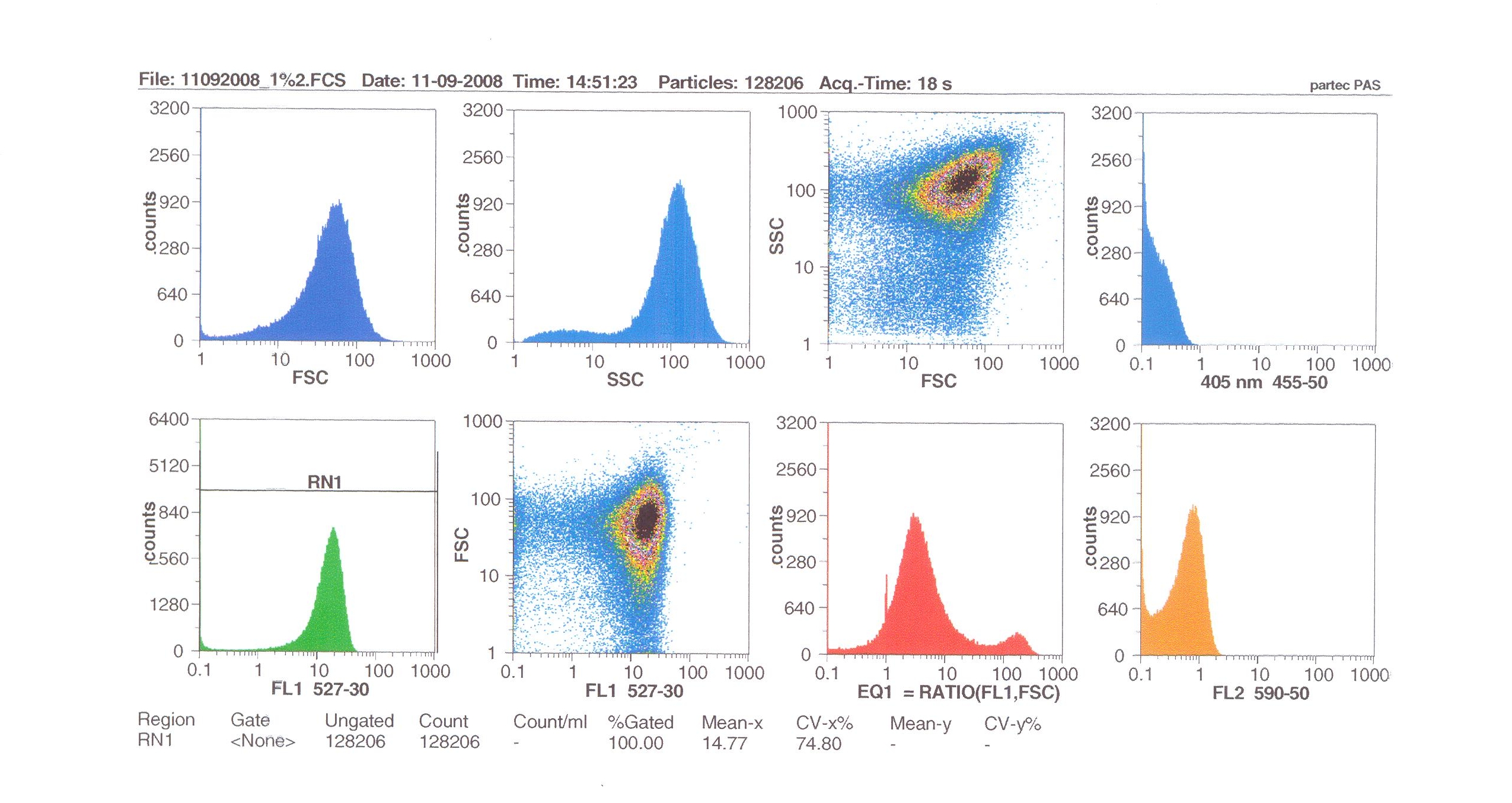
Induction by 1% v/v subtilin: mean fluoresence = 14.77
Induction by 10% v/v subtilin: the mean fluoresence = 21.95
These results show that the higher the concentration of subtilin, the more GFP is expressed.
At 10% induction, the subtilin began to cause some of the cells to lyse - this can be seen by the two distinct populations of cells - one being the cells which have lysed, and the other being the remainder of intact cells. Since subtilin is an antibiotic, this was a good indication that the subtilin we had collected that day was of good quality.
Wet Lab Journal
This contains the day-to-day progress of our wet lab team.
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"