Team:Peking University/Project
From 2008.igem.org

| Home | The Team | The Project | Parts Submitted to the Registry | Modeling | Misc&Fun | Our OWW | Notebook |
|---|
Contents |
Project Abstract
A Genetic Circuit for Directed Evolution in vivo
Directed evolution is a powerful tool for answering scientific questions or constructing novel biological systems. Here we present a simple genetic circuit for in vivo directed evolution which comprises minimal elements for random mutation and artificial selection. We engineer yeast to generate the DNA mutator hAID, an essential protein in adaptive immunity, and target it specifically to a gene of interest. The target gene will be mutated and consequently promptly evolves. By linking the expression of hAID repressor LacI and favorite gene functionality, the mutation rate inversely correlates between the functionality of the desired gene and hAID. This circuit may be adopted for in vivo evolution in eukaryotic system on genetically encoded targets. It has a variety of potential applications in academic and industrial contexts, theoretically most inter-molecular interaction that involves proteins and RNAs.
Genetic Circuit
|
Project Background
Over millions of years, evolution has created finely tuned networks of proteins capable of producing incredible outputs because of their optimized interactions with each other. The early efforts of synthetic biologists to recreate these networks artificially has given us a new respect for just how well-optimized natural systems really are; our efforts have in general appeared clumsy next to natural examples.
However, natural evolution is a very slow process, governed by subtle selective pressures and long generation times. Therefore, scientists have developed several strategies for directed evolution in the lab, breaking it down into mutation (various PCR based techniques, shuffling, mutator strains) and selection (display technologies, etc.) These methods do cut down on time, and there have been some notable successes by researches using in-vitro directed evolution. Nevertheless, running the necessary high-throughput screens and mutagenesis protocols requires a great deal of manpower, expertise, and time.
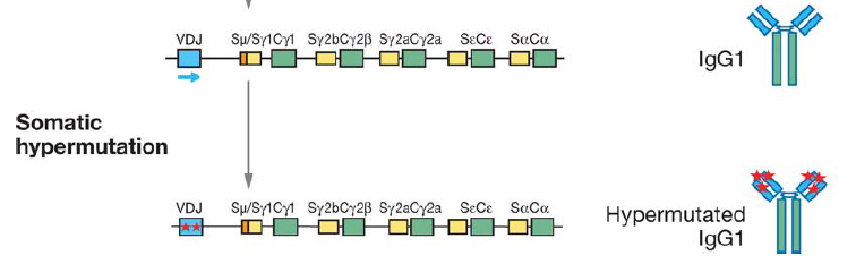
It would seem that nature has beaten us to the punch once again, however, as a natural mechanism for rapid evolution already exists: the immune system's adaptive immunity. It is well known that B-cells are capable of producing a large pool of diverse antibodies upon antigenic stimuli, a fact which underlies our own ability to survive infection. Recent work has revealed that an enzyme called AID (activation induced cytidine deaminase) is the workhorse behind this immune capacity. AID facilitates three processes crucial to antibody diversity: somatic hypermutation (SHM), class switch recombination (CSR), and gene conversion.
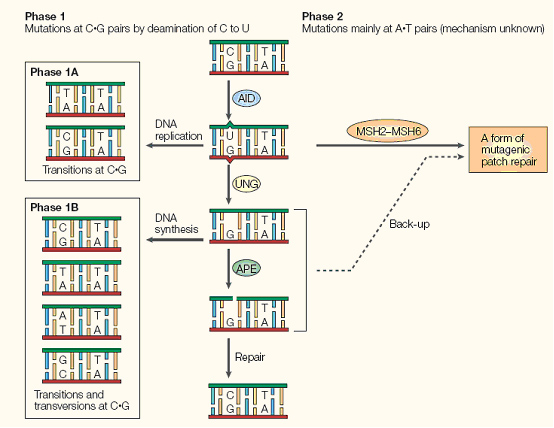
Briefly, AID converts cytidine to uracil by oxidizing the amino group to a carbonyl, resulting in a Watson-Crick mismatch. DNA repair pathways then are activated the remove the mismatch, resulting in a changed coding sequence of the hypervariable region that AID is targeted to. Because of its mutagenic properties, AID must be carefully targeted to avoid B-cell lymphocytoma.
AID has been shown to be functional in other cells besides human B-cells. In 2005, Youri Pavlov's group expressed human AID in yeast, and found that AID maintained a high mutation rate even in this heterologous host.
(Neuberger, M.S., et al., Somatic hypermutation at A.T pairs: polymerase error versus dUTP incorporation. Nat Rev Immunol, 2005. 5(2): p. 171-8.PubMed)
Inspired by this work, we hope to engineer a genetic circuit in yeast that will govern the directed evolution process from mutation to selection. Compared to human B-cells, yeast is cheap and reproduces incredibly fast. Moreover, yeast is a very well characterized eukaryotic model with many of the post-translational abilities found in human cells, allowing us to do in vivo analysis of protein function without having to extract and purify. Our system allows for a wide variety of substrates, essentially any intermolecular interaction that involves proteins.
Project Details
Project Proposal
Evolution could be dissociated into two parallel processes: random mutation, and natural selection. Whilst its role in life history is well understood and accepted, evolution is also evident across individual life process. In particular, adaptive immunity system adopts evolution strategy to produce antibodies for novel antigens. Artificial evolution method could be a powerful tool for answering scientific questions or engineering novel biological systems. Via systems biology approach, here we present a simple genetic circuit consisting functional elements for random mutation and artificial selection. This circuit may perform in vivo evolution on virtually any genetically encoded targets, with potential applications in academic and industrial contexts.
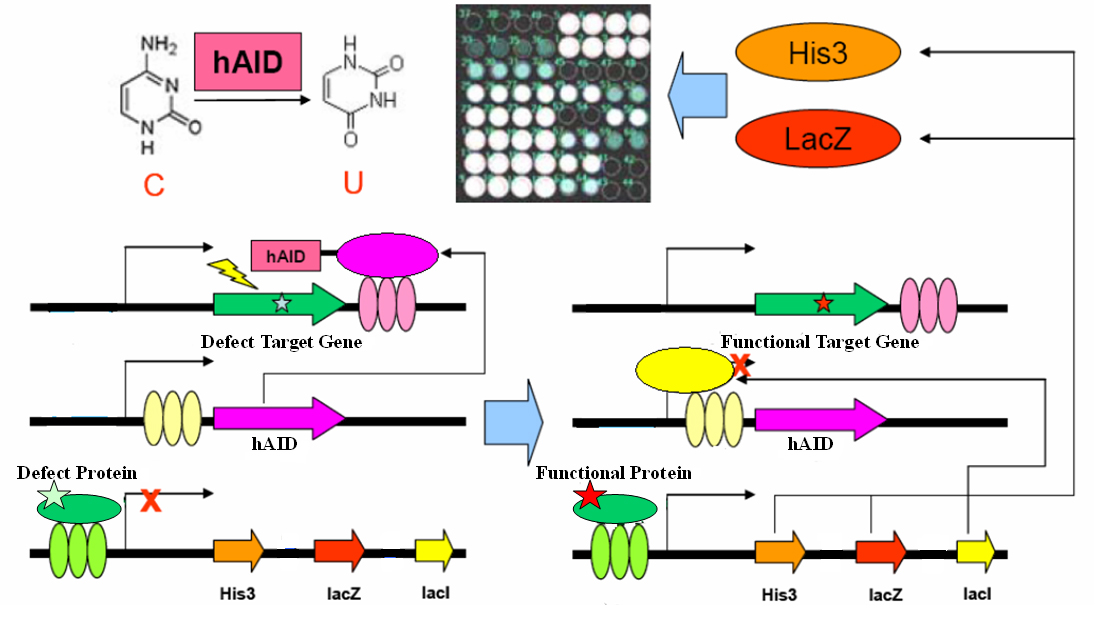
The gene encoding the core element in adaptive immunity, hAID, is fused with the LexA-DBD domain with flexible linker. The fusion gene hAID-LexA is inserted into a yeast ESC expression cassette at 3’ of tandem LacO elements. The target gene (your favorite gene, yfg) is inserted into a yeast ESC expression cassette, with its 3’UTR containing tandem LexO elements. An inducible promoter response to yfg activity drives expression of His3, LacZ and LacI in cistrone spanned by IRES. These three plasmids together form a genetic circuit for in vivo evolution.
For clearness, here we present one very simple example in yeast one hybrid: a mutated Gal4 gene is inserted in the target cassette. The selector cassette contains UAS to drive expression of His3, LacZ and LacI. All three plasmids are transfected or knock-in-ed into gal4-, his3- yeast strain with proper selection tags. Initially, we culture the yeast in complete YPD medium. Defect Gal4 product cannot bind to UAS, hence hAID-LexA is constitutively expressed, recruited to LexO sites of the target, and mutates the Gal4 conding sequence. Once Gal4 mutation is reversed, it binds to UAS to drive LacI expression, which represses hAID-LexA. We plate the yeast into Trp-, His-, Leu-, 3AT+, XGal+ plate and select for the large, blue colonies. Sequencing the colonies then gives us the activated Gal4 gene sequences.
The power of this system could be best revealed in the above case: since selection is not required for initiation and attenuation of mutagenesis, there is virtually no need to perform selective pressure titration.
The system could be readily adopted to in vivo evolution of any kind of protein-protein and protein-nucleic acid interaction: for this we simply adopt a yeast two hybrid-like approach, evolving a functional protein linked to Gal4 AD which could bind to the "bait" protein linked to Gal4 DBD. We could also use the yeast three hybrid system to study RNA-protein interaction, or use yeast one hybrid to evolve specific protein that binds to given promotor.
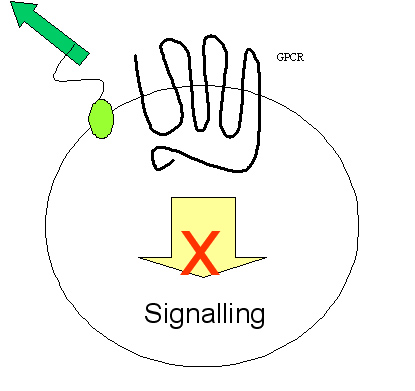
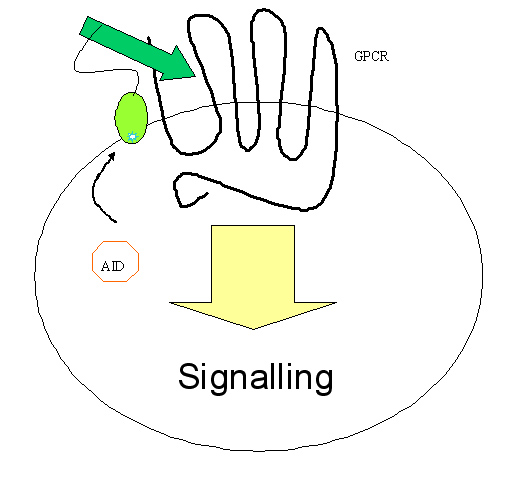
Yeast shows several advantages in studying eukaryotic process, for it folds and modifies the eukaryotic protein accurately, whilst bacteria does not. Another advantage of yeast is that cell wall blocks intracellular communication between transmembrane proteins, and the selection happens completely within the cell, therefore blocking possible false positives. We plan to screen for extracellular activator of certain eukaryotic transmembrane protein, for example, novel peptidergic ligand for GPCR and antibody-antigen interaction.
We have been working on further improvements of the mutation system, by improving the enzyme to archieve directed, "hotspot-less", and evenly distributed mutagenesis, by utilizing the overwhelming power of yeast genetics, and by harnessing novel protein chemistry.
//Last modified: ZY 20080704
The Experiments
Plasmids Construction
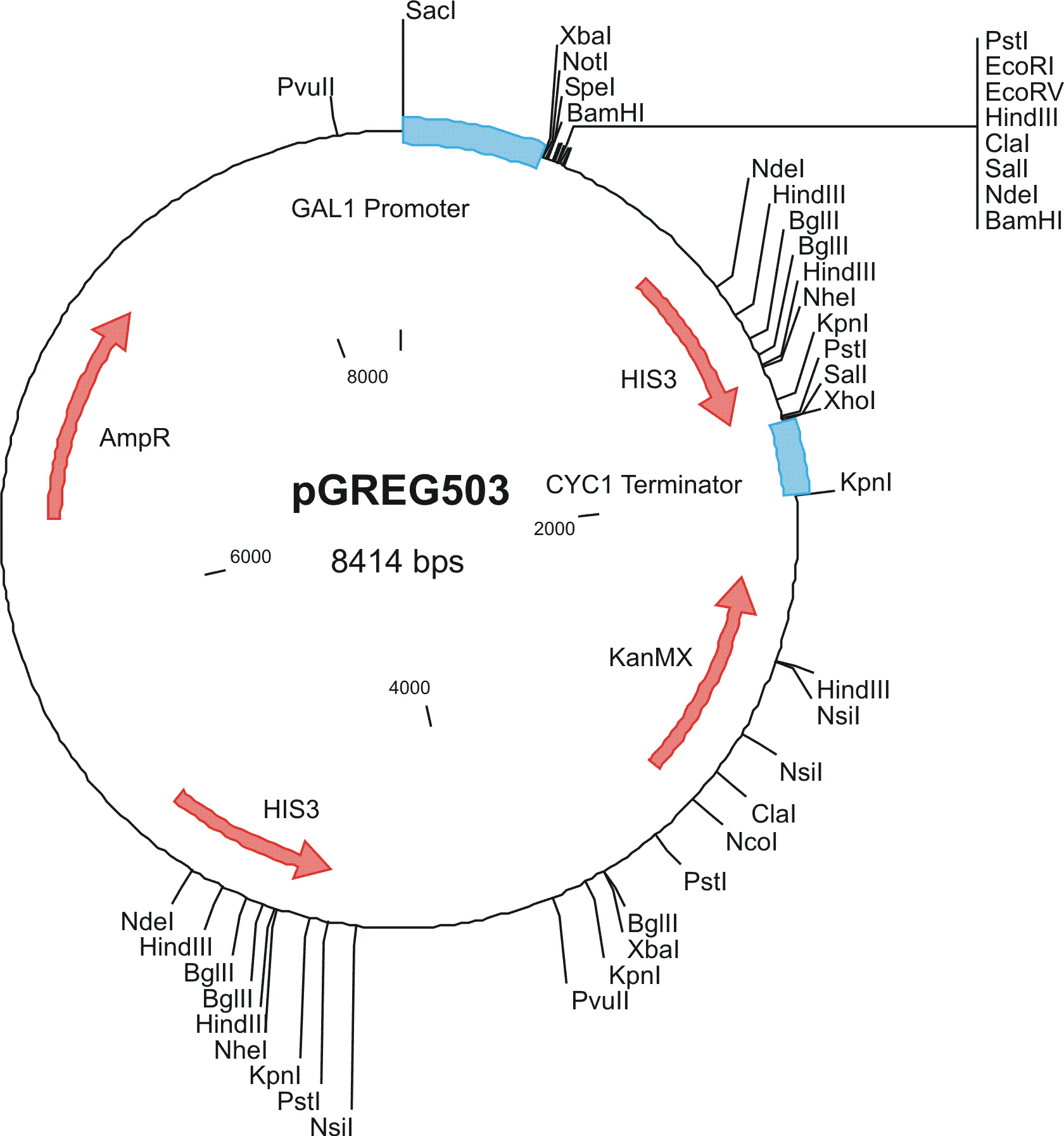
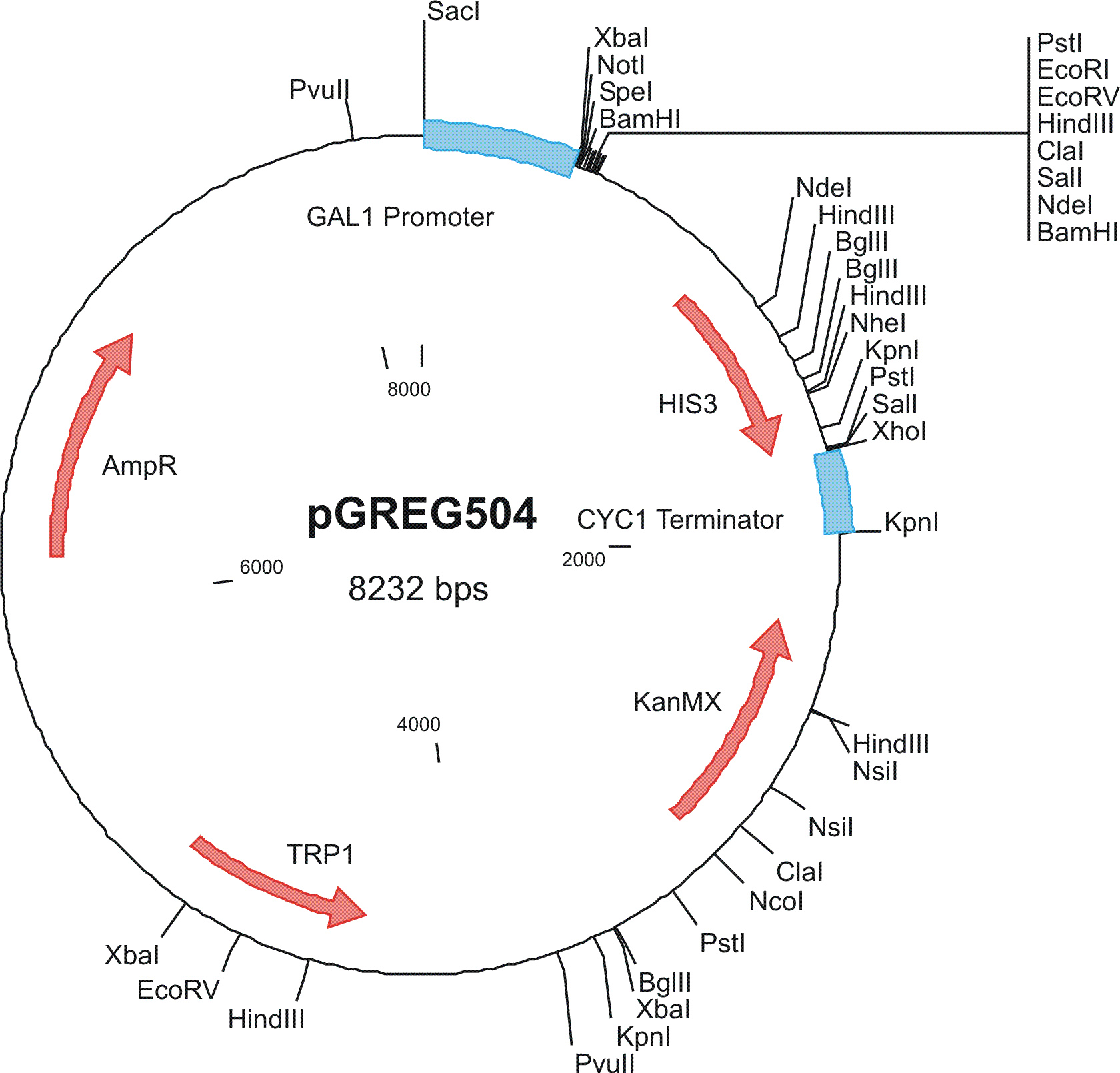
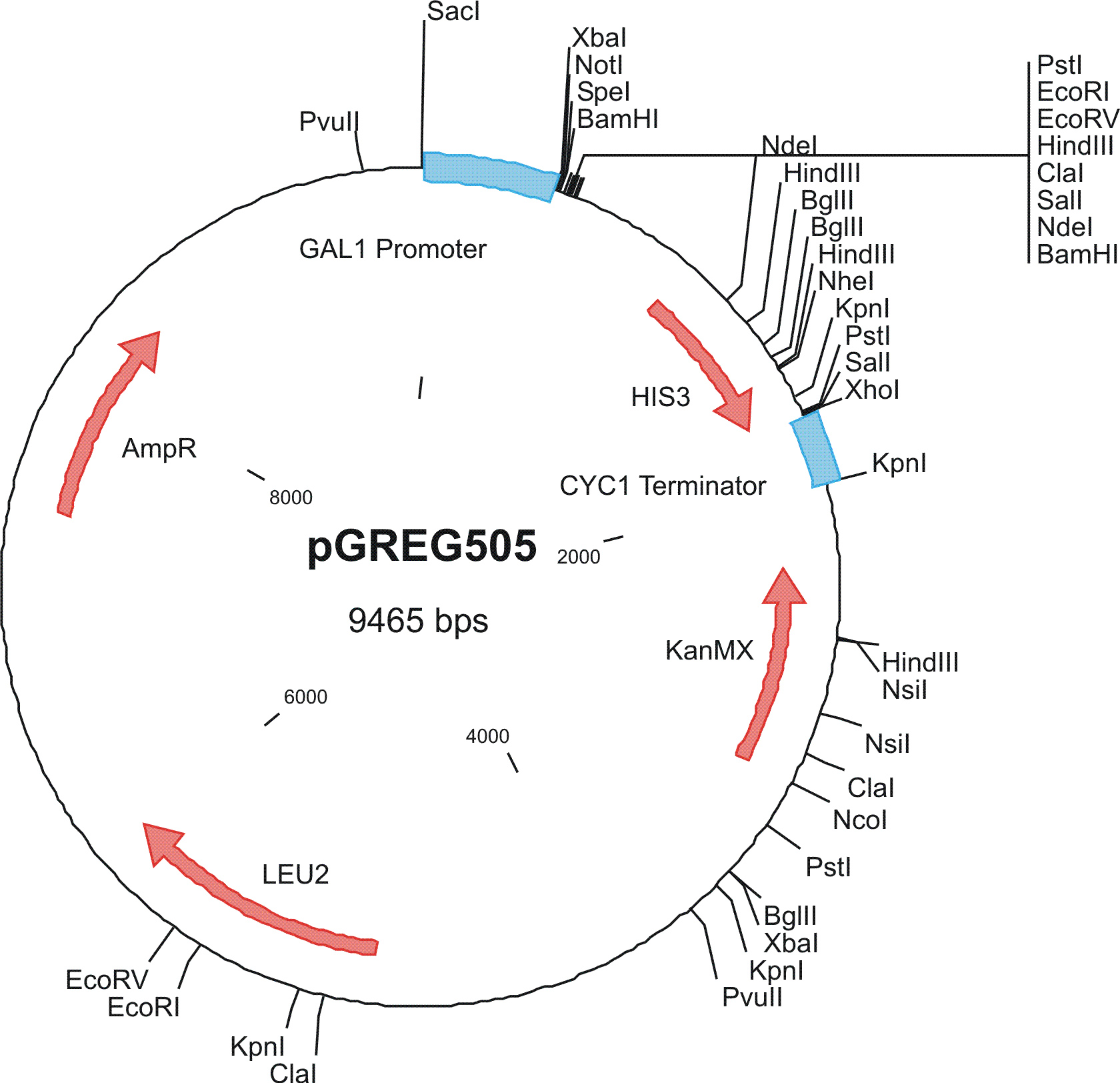
To construct the genetic circuits above, we have used the pGREG series of vectors which were hosted in budding yeast Saccharomyces cerevisiae AH109. pGREG503, the expression plasmid with HIS3 auxotroph resistance was used to construct the UAS-lacI component. LacI was under the galactose inducible promoter GAL1. The promoters of pGREG504 and pGREG505 were replaced by pADH-lac and pACT respectively. pADH-lac is pADH based promoter which is sensitive to lacI, genes under this promoter will be inhibited by lacI. pACT is a constitutively expressing promoter derived from the promoter of actin in budding yeast. The construct hAID-linkers-lexA was under the pADH-lac promoter and the GAL4-lexO construct was engineered under the pACT promoter. Notably, in the genome of the yeast strain AH109, it has GAL1-HIS3 and GAL1-lacZ constructs. PGREG504-hAID-linkers-lexA DBD was constructed via the following Figure.
| 
| 
|
To construct plasmid on pGREG503, we have used the versatile PCR-based homologous recombination system: the PCR product of lacI was directly co-transformed with pGREG503 and it was automatically recombined to the site flanking the 3’ of the promoter. Other constructs were accomplished with standard protocols.
Yeast Strains Preparation
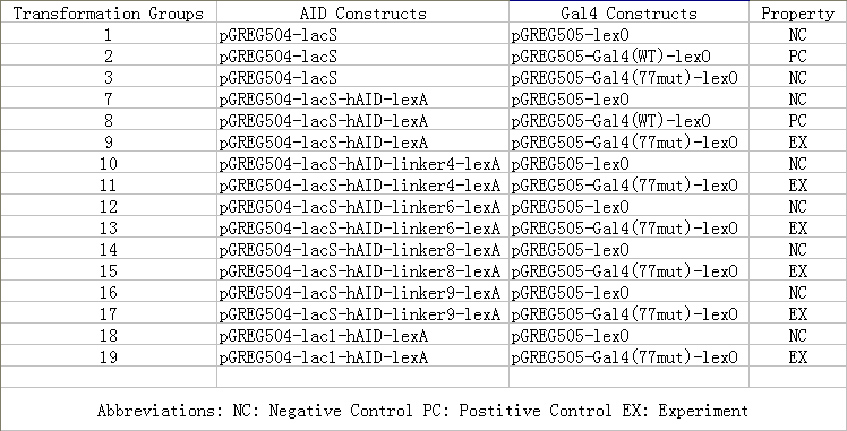
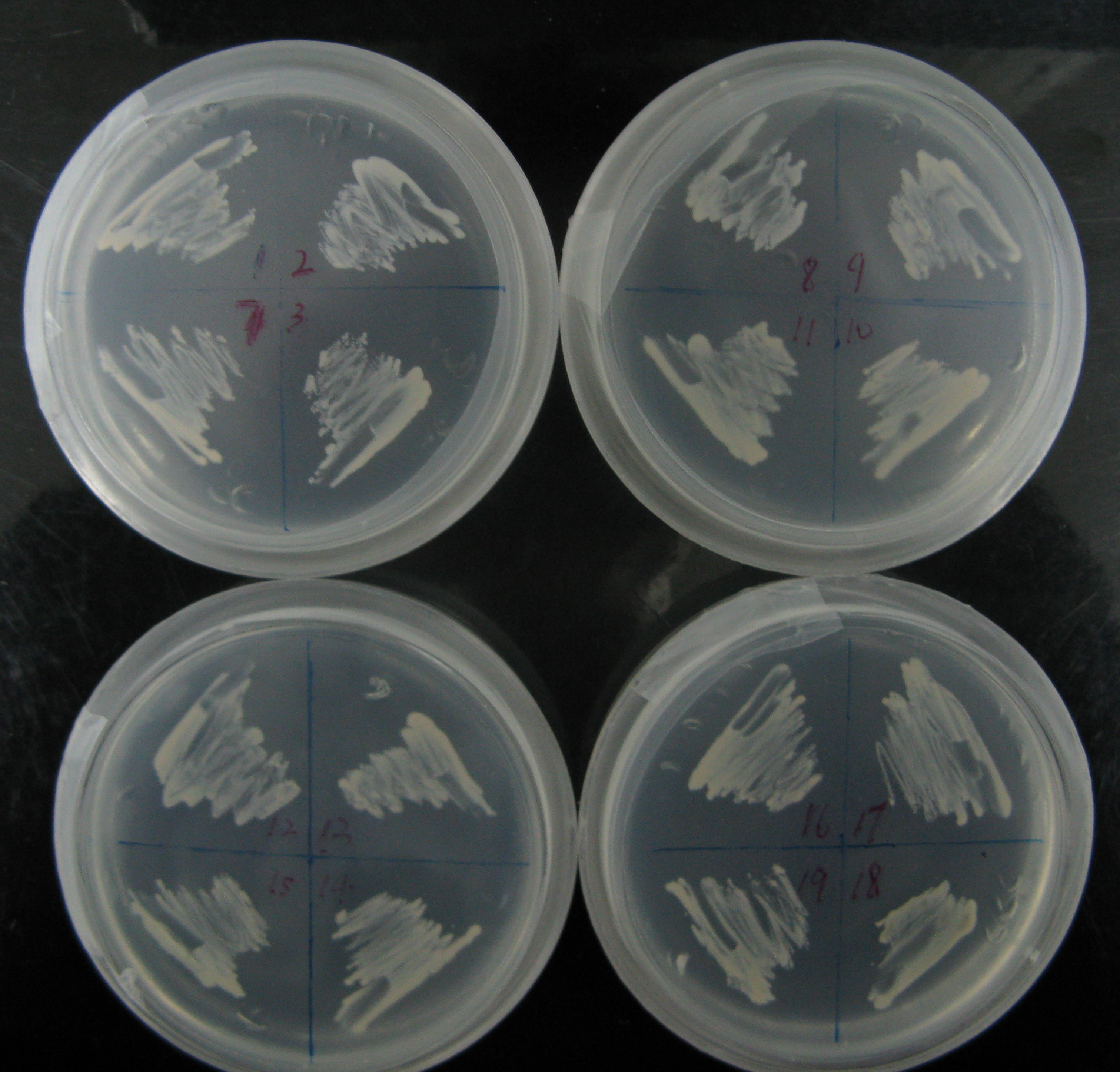
Among more than 50 plasmids we have constructed, we have chosen the above 16 groups for the following yeast screening task. For the AID construct which is on pGREG504, we have pGREG-lacS (note: lacS is one type of pADH-lac promoters) as a negative control and those with lac1, another pADH-lac promoter. A set of four linkers were tested in order to mostly maintain the original activity of hAID. For the GAL4 constructs, many spot mutations were triggered in order to disrupt function of wildtype GAL4 but only a spot mutation at 77-position was selected to carry out the further yeast screening process. Wildtype gal4 in pGREG505 and plasmids without gal4 gene were also used as positive and negative controls.
As the above table listed out, 16 strains of yeast were constructed via co-transformation with AID construct and GAL4 construct.
 "
"