Team:TUDelft/Color modeling
From 2008.igem.org
(Difference between revisions)
| Line 9: | Line 9: | ||
For the 14 reactions of enzyme-subtrate the Michaelis-Menten kinetics is applied and we have the mass action kinetics for the three degradations. | For the 14 reactions of enzyme-subtrate the Michaelis-Menten kinetics is applied and we have the mass action kinetics for the three degradations. | ||
| - | [[Image:Mevalonate.jpg | | + | |
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | [[Image:Mevalonate.jpg |500px]] | ||
| + | |||
| + | Color Pathway | ||
</div> | </div> | ||
{{Template:TUDelftiGEM2008_sidebar}} | {{Template:TUDelftiGEM2008_sidebar}} | ||
Revision as of 13:36, 21 August 2008
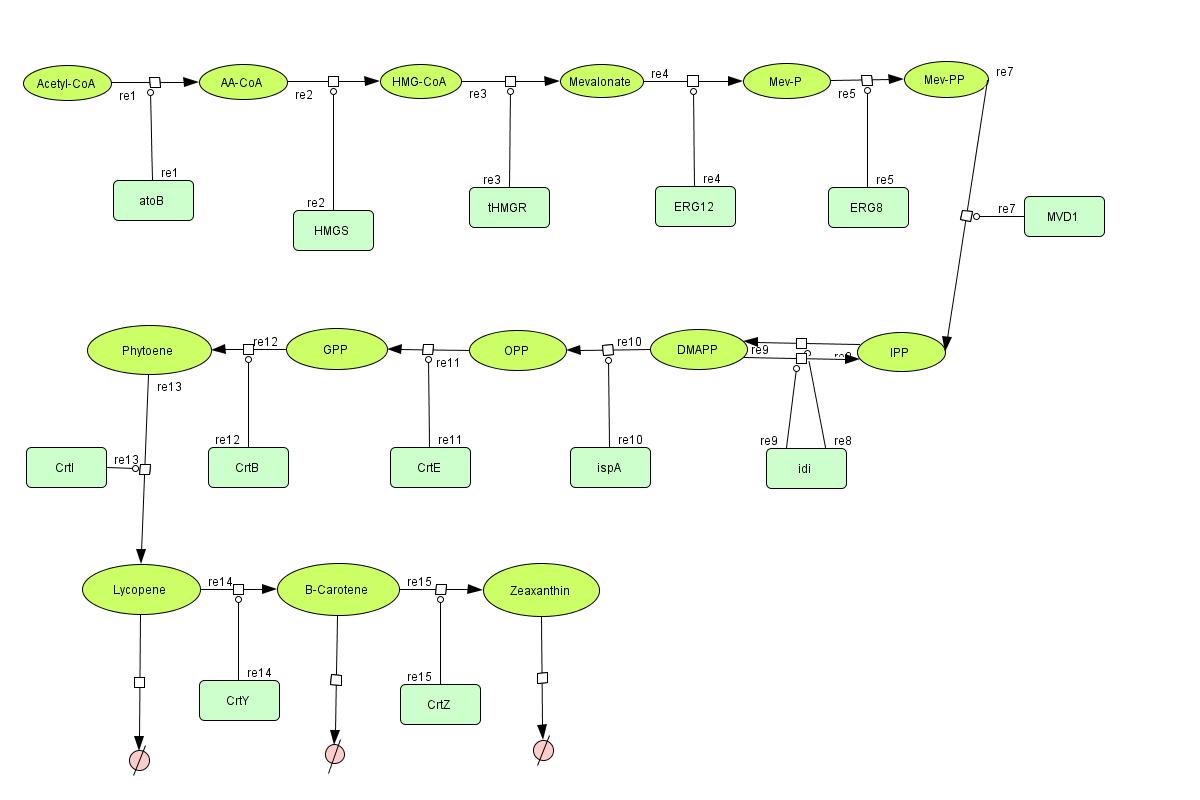
In the output we use mevalonate and GPP pathways together to produce red, orange and yellow colors. The biosynthetic model is bulit in CellDesigner™. There are 14 subtrates and 13 enzymes in total. The last three subtrates are the color products and are consuming so we have degradation links for them. Lycopene, B-carotene and Zeaxanthin give red, orange and yellow colors respectively. For the 14 reactions of enzyme-subtrate the Michaelis-Menten kinetics is applied and we have the mass action kinetics for the three degradations.
Color Pathway
 "
"