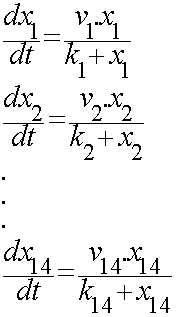Team:TUDelft/Color modeling
From 2008.igem.org
Bavandenberg (Talk | contribs) |
|||
| Line 3: | Line 3: | ||
{{Template:TUDelftiGEM2008_menu_home}} | {{Template:TUDelftiGEM2008_menu_home}} | ||
| - | |||
>> work in progress | >> work in progress | ||
| Line 26: | Line 25: | ||
... to be continued ;) | ... to be continued ;) | ||
| - | |||
| - | |||
| - | |||
| - | |||
{{Template:TUDelftiGEM2008_sidebar}} | {{Template:TUDelftiGEM2008_sidebar}} | ||
Revision as of 16:01, 25 August 2008
>> work in progress
Color Modeling
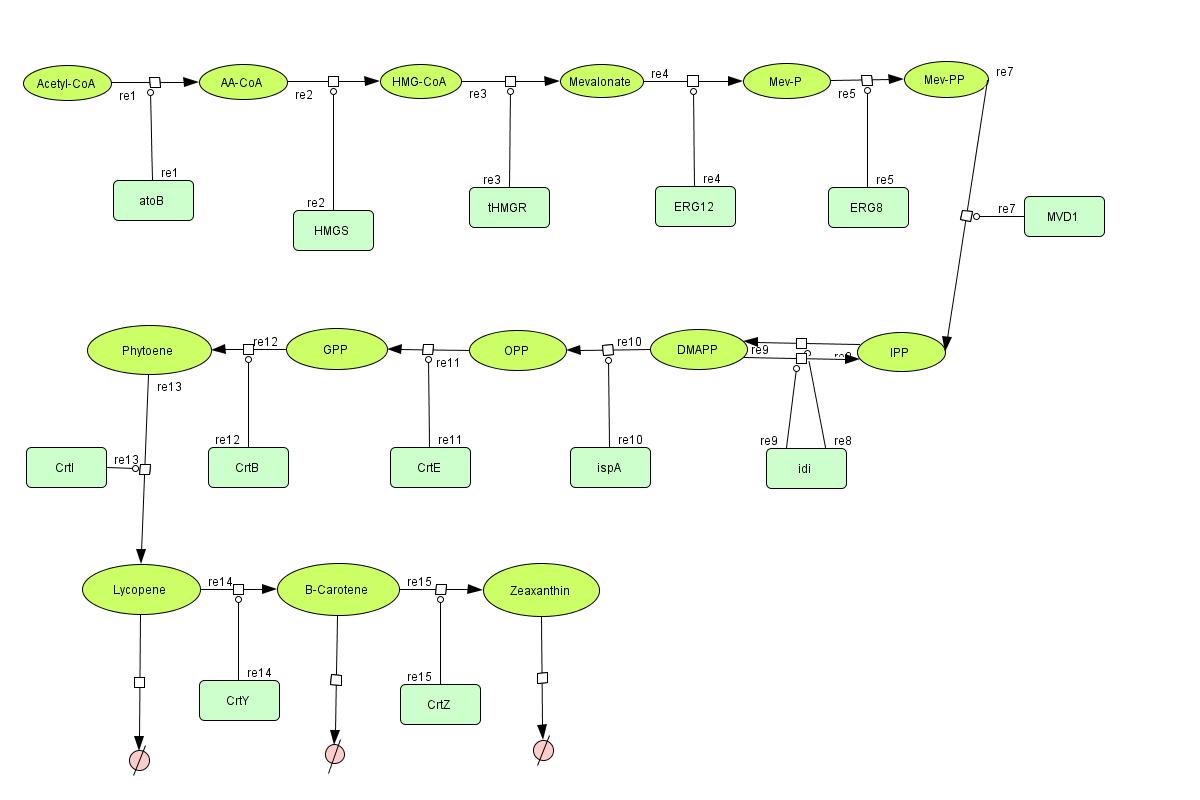
In the output we use mevalonate and GPP pathways together to produce red, orange and yellow colors. The biosynthetic model is bulit in CellDesigner™. There are 14 subtrates and 13 enzymes in total. The first subtrate is Acetyl-CoA which is provided to the pathway in a constant level. The last three subtrates are the color products and are consuming so we have degradation links for them. Lycopene, B-carotene and Zeaxanthin give red, orange and yellow colors respectively.
For the 14 reactions of enzyme-subtrate the Michaelis-Menten kinetics is applied and we have the mass action kinetics for the three degradations.
According to the kinetic laws there are 14 differential equations for enzyme-subtrate reactions and three for degradations which could be constructed as:
x1 to x14 are the subtrates.
... to be continued ;)
 "
"