Team:Wisconsin/Project
From 2008.igem.org
| Home | The Team | The Project | Parts Submitted to the Registry | Modeling | Notebook |
|---|
Abstract
The global fuel crisis impacts our economy, national security, and environment. The need for alternative fuels is of utmost importance. Team Wisconsin used Escherichia coli to produce biofuel precursors in an effort to find these alternative fuel sources. One project focused on using E. coli to efficiently produce sorbitol, a biofuel precursor. Using computer modeling, we determined a way to funnel glycolysis’ intermediates towards the production of sorbitol via sorbitol-6-phosphate dehydrogenase. Wisconsin’s second aim was to isolate high-energy precursors from the plant cell wall through E. coli mediated breakdown of cell wall lignin. This was achieved by inserting the gene encoding lignin peroxidase , found in the white rot fungus Phanerochaete chrysosporium, into genetically modified E. coli, capable of producing and exporting the enzyme. Both projects improve current methods in the production of alternative fuels via two different, unique routes, and have the potential to move sustainable biofuel research forward.
Introduction
In the U.S. currently, corn is used for ethanol and other biofuel production. However, only the kernel is used for fermentation despite the plant being cut at the base during harvest. The ratio of energy harnessed to energy put into ethanol production using corn is approximately 1.34:1, which is not a great ratio when you think about this method as a means for harnessing solar energy. Breakdown of cellulose for fermentation seemed the obvious next step for ethanol production as a biofuel since cellulose is the most abundant organic matter on earth (33% of plant matter). This seems to have been recently accomplished by several people with industrial production in the works. Due to the increased costs of fossil fuels and improvement in technology, it has now become cost effective to harness renewed energy whether it be directly through wind power or chemically through biofuel synthesis.
Figure 1: Map of forms of sustainable chemical production.
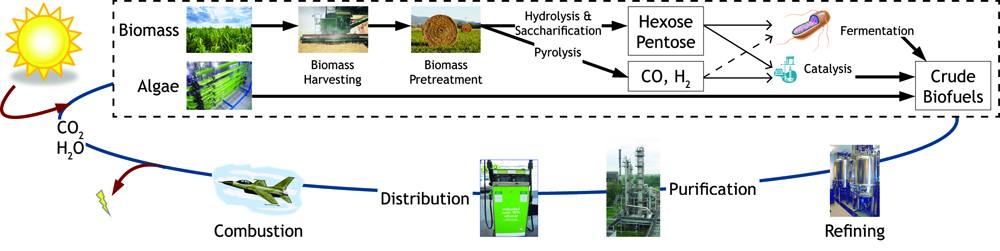
Figure 2: The biofuel cycle.
Overall Project
Fuel production has come to the forefront as an important political and biological issue. It has lead to innovative pursuits of renewable fuel as well as the controversial exploitation of natural resources. Currently ethanol is the commercial biofuel of choice; however, current production and distillation of ethanol in the United States is inefficient. With this problem in mind, the iGEM Wisconsin team is looking for alternative ways to make not only ethanol, but other biofuels through synthetic biology. We've designed the following two projects based around using E. coli to produce biofuels.
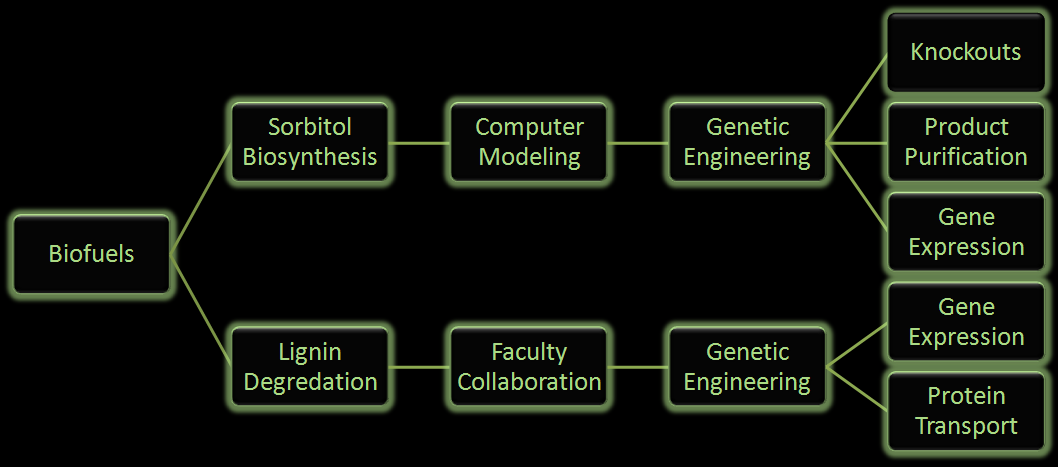
Figure 3: Flow chart illustrates the process our team followed in creating our genetically engineered cells.
One project focuses on using E. coli to produce sorbitol, a sugar alcohol, in large quantities for eventual commercial scale catalytic conversion to hydrocarbons. Along with producing sorbitol, we've modeled alterations in E. coli to make sorbitol production from a glycerol carbon source possible. Our aim is to modify a cell that will utilize glycerol, a byproduct from biodiesel production, and effectively create sorbitol.
In the other project we are attempting to use E. coli to break down lignin from plant matter. This would increase the efficiency of biofuel production from plant biomass. We are currently aiming to insert a fungal gene coding for lignin peroxidase into E. coli. Lignin breakdown will be made possible through the transport of lignin peroxidase out of the cell.
Lignin Peroxidase
Our project takes this initiative one step further by identifying and creating a means to break down lignin for fermentation as well. Lignin is the second most abundant organic substance on earth (30% of plant matter). Not only is it another largely untapped source of energy, it is one of the obstacles to cellulose fermentation efficiency. Lignin is a very complex phenyl-propane polymer that is covalently attached to cellulose. Lignin also happens to be one of the hardest organic molecules to degrade, causing challenges in cellulose fermentation. Lignin is bound to the outside of cellulose as a component of the cell wall. This hinders cellulose breakdown in nature by preventing access to cellulose as a substrate for enzymes. However, there are enzymes and organisms out there that can break down lignin. By adding genes necessary for the enzymatic breakdown of lignin to a microbe, it allows for the breakdown of lignin on a large scale, facilitating ethanol fermentation from organic matter. The white rot fungus, Phanerochaete chrysosporium, has the enzyme lignin peroxidase which seems to be promising for our purpose. If this is successful, not only will ethanol production from corn be much more efficient but other biological sources for ethanol could also be more feasible such as switch grass, wood scraps, yard waste, and really just about any plant matter readily available.
However, there are some difficulties with the implementation. Lignin is an extremely complex molecule formed by radical reactions, and it has proved very difficult to study. The mechanism of lignin peroxidase is not fully understood. Also, there could be other genes involved with lignin breakdown some of which may not even be known to the experts.
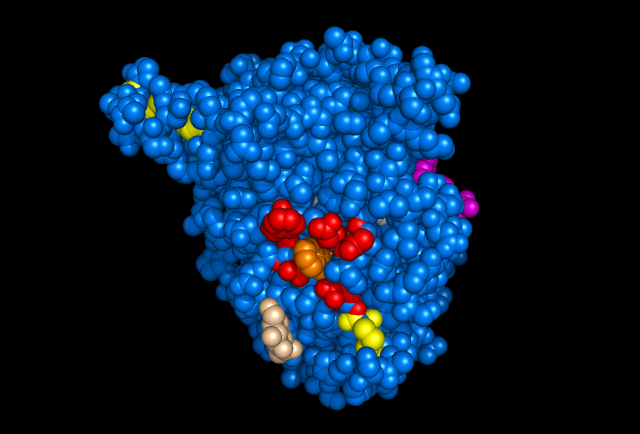
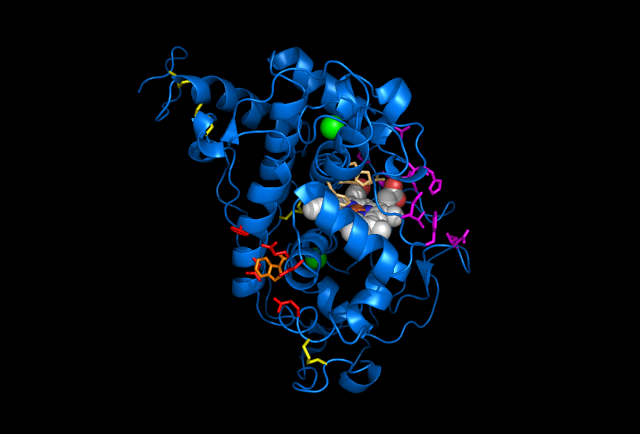
Figure 4: Images of Lignin peroxidase. Disulphide bonds (yellow), Ca2+ ions (green spheres), Heme access
channel (purple), Surface active cite (red) with hydroxylated typ171
(orange), Hydrogen peroxide active cite (tan wireframe, above heme),
Heme group (CPK coloring space fill.)
In researching for an appropriate assay in analyzing the amount of lignin peroxidase produced by our E. coli strain, we found that the veratryl alcohol assay, developed in 1984, is currently the most widely used. It works by viewing the absorbance spectrum of the reaction solution. As veratryl alcohol is oxidized by lignin peroxidase over time, the optical density (OD) and wavelength of the absorbance peak changes. By observing the disappearance of the substrate (the rate of change), we can determine the amount of active enzyme in the solution.
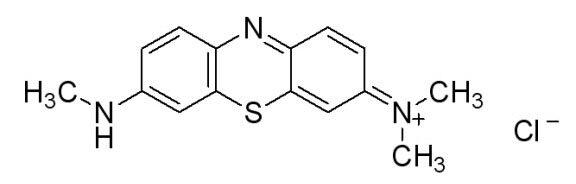
In doing additional research we came across a newer and seemingly more effective assay—the Azure B assay. Research showed that Azure B (a blue colored dye) has a higher specificity to lignin peroxidase compared to veratryl alcohol, as veratryl alcohol can be oxidized by fungal enyzmes other than lignin peroxidase. The Azure B assay works the same way as the veratryl alcohol assay and the disappearance of the dye is measured to determine the amount of lignin peroxidase. The reaction solution comprises of a tartrate buffer (50 mM at pH 4.5), the dye Azure B (32 μM), hydrogen peroxide (100 μM), and the enzyme to be measured, lignin peroxidase.
Figure 5: [http://www.sigmaaldrich.com/catalog/search/ProductDetail/SIGMA/A4043 Azure B. Sigma Aldrich.] (2008). Retrieved Oct 18.
Sorbitol Biosynthesis
Sorbitol anabolism is part of the glycolysis pathway in E.coli. In the pathway, interconversion between fructose and sorbitol is catalyzed by sorbitol dehydrogenase(SDH), a monomeric enzyme that uses NADH in the process. Natural expression of SDH is generally up-regulated only in the presence of sorbitol as a carbon source. The initial idea to stimulate sorbitol over-production was to simply insert a plasmid containing the gene which codes for SDH and grow the cells in an environment lacking sorbitol thus pushing the metabolic reaction governing fructose and sorbitol interconversion towards sorbitol.
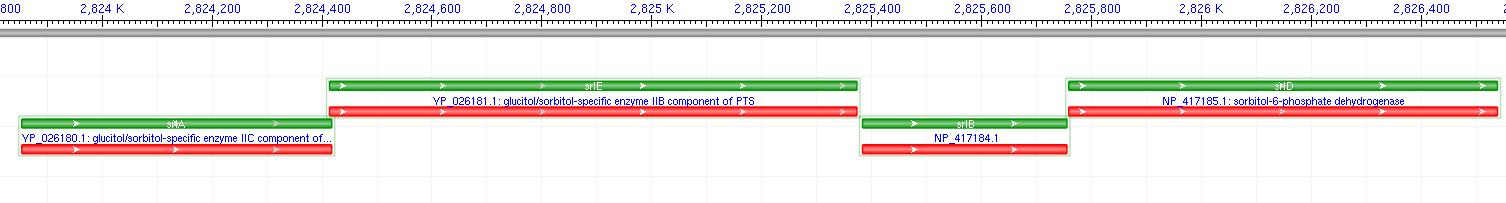
The gene encoding for SDH, srlD, is part of the srl Operon in E.coli (Operon pictured below). Information on the SDH assembly pathway and enzymatic activity is limited. Consequently we decided that insertion of both the srlD gene and srl operon into vectors would take more time initially but would potentially be beneficial due to the chance that the gene alone would not result in effective SDH production.
Increased cellular production of SDH would most likely increase the amount of sorbitol produced by the cell, however it could not alone account for the type of mass production that would make sorbitol biosynthesis commercially useful. In order to determine other methods of increasing sorbitol production, our group turned to computer modeling.
Phosphofructokinase Knockout
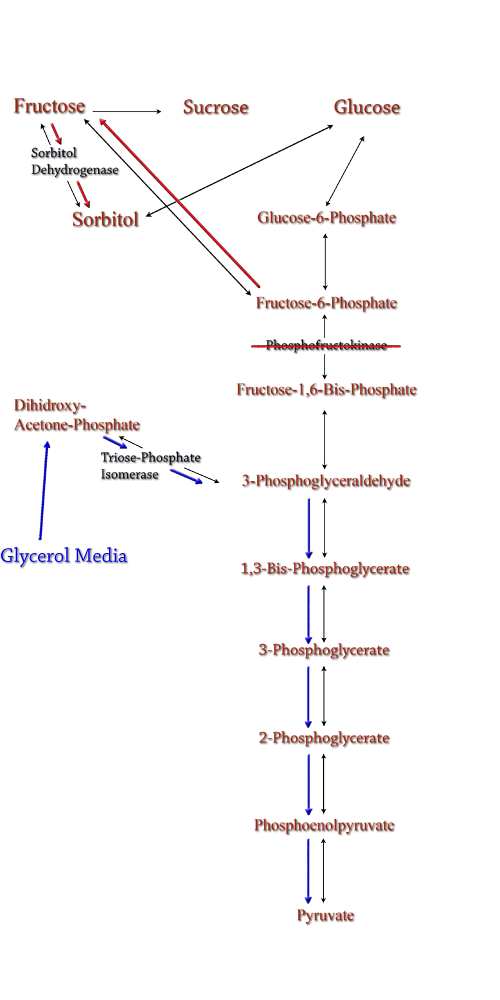
Figure 8: PFK Knockout in the metabolic pathway.
The idea behind a phosphofructokinase (PFK) knockout is to literally stop the glycolysis pathway after fructose-6-phosphate. The backup of metabolic products caused by the knockout should pool and form a favorable pathway for increased sorbitol production. The obvious problem with this procedure is that knocking out a part of the glycolysis pathway under normal circumstances kills the cell. This is also why modeling didn't prove useful as the program showed the cell would be dead. We set out to determine how to bypass this normal cellular function in order to use it to increase sorbitol production.
Experiments have been successfully done to create a viable PFK chromosomal knockouts that contain inducible plasmids encoded with the pfk gene. After obtaining this strain we have done countless growth curves with varying media to create ideal conditions for cell growth. After the optimal media is obtained, the addition of a plasmid containing srlD or the srl operon will be introduced to create what we hope will be this metabolic pathway(right).
Triose Phosphate Isomerase Knockout
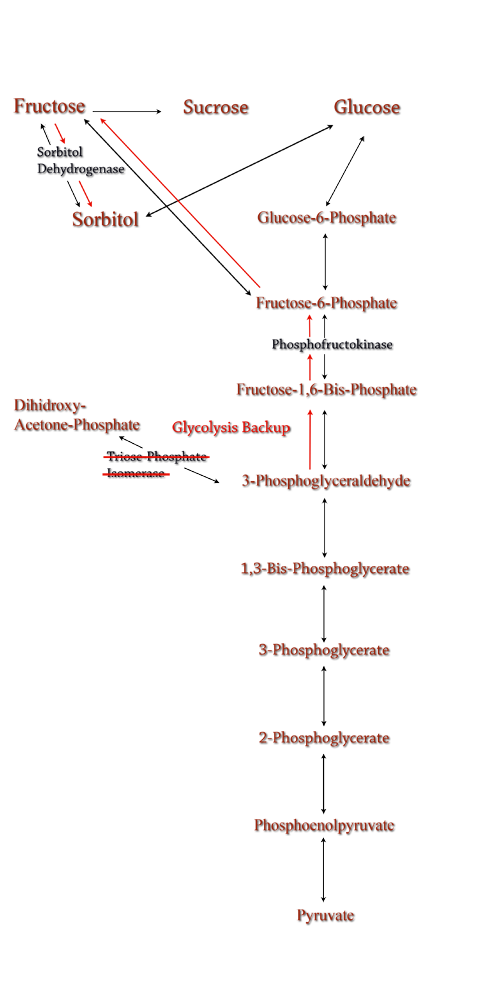
Figure 7: TPI Knockout in the metabolic pathway.
Computer modeling was done to determine gene knockouts that increased cellular flux towards sorbitol production. The model was given to us by Dr. Jenni Reed of University of Wisconsin-Madison. More extensive information about modeling can be found at our modeling page. Through the use of our model we determined that knocking out the gene encoding for triose phosphate isomerase would be the most effective at increasing sorbitol production when combined with srl gene up-regulation.
Triose phosphate isomerase(TPI) is another enzyme in E.coli's glycolysis pathway. Its function is to catalyze the interconversion of 3-Phosphoglyceraldehyde to Dihidroxy-acetone-phosphate and back. Knocking out the gene which encodes TPI stops this interconversion, a process which removes products of glycolysis from the cellular reaction mixture shifting the overall reaction downwards towards pyruvate. In essence, knocking out the TPI gene backs up glycolysis and produces a shift towards sorbitol (Pictured right).
TPI is a nonessential gene and as such we were able to obtain it from the Keio collection, a collection of non-essential single gene knockouts from the E.coli K-12 derivative strain BW25113. The increased cellular flux modeled in a TPI knockout combined with increased intracellular SDH production from a plasmid containing either the srlD gene or srl operon will hopefully result in a large scale production of cellular sorbitol.
Even with the promising modeling results, our group worked on alternative routes for increased cellular sorbitol. One of these routes, knocking out phosphofructokinase, was an additional project that our group undertook.
Approach
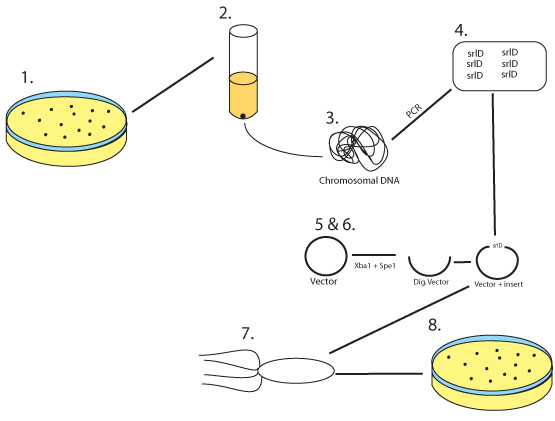
Figure 8: 1. Wild Type E.coli MG1655 is grown on an LB plate
2. A colony is selected and grown overnight
3. The chromosomal DNA is harvested from the over night culture
4. srlD gene is cloned and amplified via a PCR reaction adding restriction sites for Xba1 and Spe1 at either end
5. The biobrick vector pSB1AK3 as well as the srlD PCR product are digested with Xba1 and Spe1
6. The two digestion products are then ligated together
7. The ligation product is then transformed into competent E.coli
8. E. coli is then plated on LB plates containing Amp/Kan to select for transformants
References
Guzman L., Belin D., CarsonM., and Beckwith L. (1995). Tight regulation, modulation and high-level expression by vectors containing the arabinose pBAD promoter. Journal of Bacteriology. 177(14): 4121-4130.
Lorenzo Nissen, Gaspar Pérez-Martínez, María J. Yebra (2005) Sorbitol synthesis by an engineered Lactobacillus casei strain expressing a sorbitol-6-phosphate dehydrogenase gene within the lactose operon �FEMS Microbiology Letters 249 (1) , 177–183 doi:10.1016/j.femsle.2005.06.010
Roe, A. J., C. O'Byrne, D. McLaggan, and I. R. Booth. 2002. Inhibition of Escherichia coli growth by acetic acid: a problem with methionine biosynthesis and homocysteine toxicity. Microbiology 148:2215-2222.
Ladero, V., Ramos, A., Wiersma, A., Goffin, P., Schanck, A., Kleerebezem, M., et al. (2007) High-level production of the low-calorie sugar sorbitol by Lactobacillus plantarum through metabolic engineering. Appl Environ Microbiol 73: 1864–1872.
Metzger J (2006) Production of liquid hydrocarbons from biomass. Angew. Chem. Int. Ed. 2006, 45, 696-698.
Lovingshimera, M. R., Siegeleb, D., & Reinharta, G. D. (2006). Construction of an inducible, pfkA and pfkB deficient strain of Escherichia coli for the expression and purification of phosphofructokinase from bacterial sources. Protein Expression and Purification , 46 (2), 475-482.
Karacaoğlan, V., & Özer, I. (2005). Steady-state kinetic properties of sorbitol dehydrogenase from chicken liver. Comparative Biochemistry and Physiology , 140, 309-312.
Archibald, F. S. A new assay for lignin-type peroxidases employing the dye azure B. Applied and Environmental Microbiology, 58(9), 3110-3116.
 "
"