Team:Wisconsin/Project
From 2008.igem.org
| Home | The Team | The Project | Parts Submitted to the Registry | Modeling | Notebook |
|---|
| Overall Project |
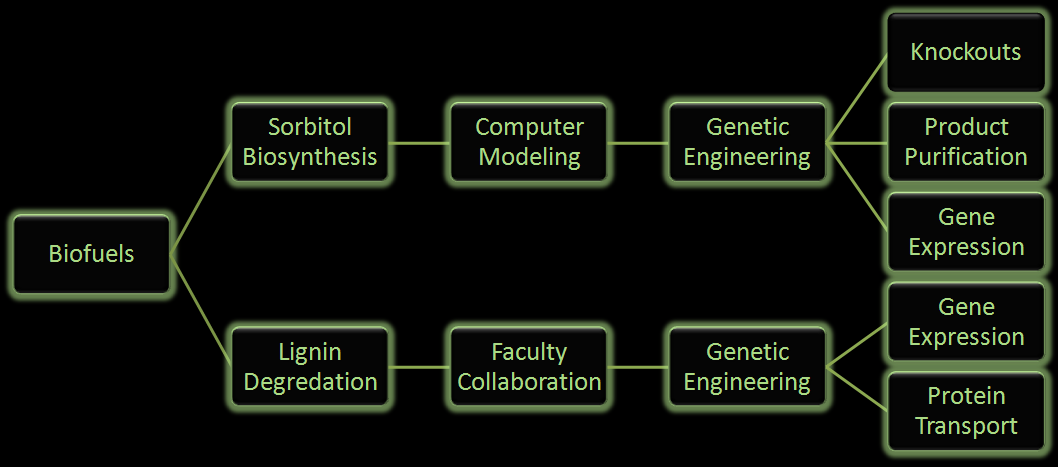
| Fuel consumption has come to the forefront as an important political and biological issue that has lead to innovative pursuits of renewable fuel as well as the controversial exploitation of natural resources. Currently ethanol is the commercial biofuel of choice; however, currently production and distillation of ethanol is inefficient. With this problem in mind, iGEM Wisconsin has looked for alternative ways to make not only ethanol, but other biofuels through synthetic biology. We've designed the following two projects using E. coli to produce biofuels in innovative ways: |
|
| One project focuses on using E. coli to produce sorbitol, a sugar alcohol, in large quantities for eventual commercial scale catalytic conversion to hydrocarbons. Along with producing sorbitol, we've modeled alterations in E. coli to make sorbitol production from a glycerol carbon source possible. Our aim is to modify a cell that will utilize glycerol, a byproduct from biodiesel production, and effectively create sorbitol.
In the second project we will be attempting to use E. coli to break down lignin from plant matter into usable biofuels. We are currently aiming to insert fungal genes coding for lignin peroxidase into E. coli. Lignin breakdown will be made possible through the transport of lignin peroxidase out of the cell. To achieve this, protein transporters will be added to the cell.
|
| Sorbitol Biosynthesis |
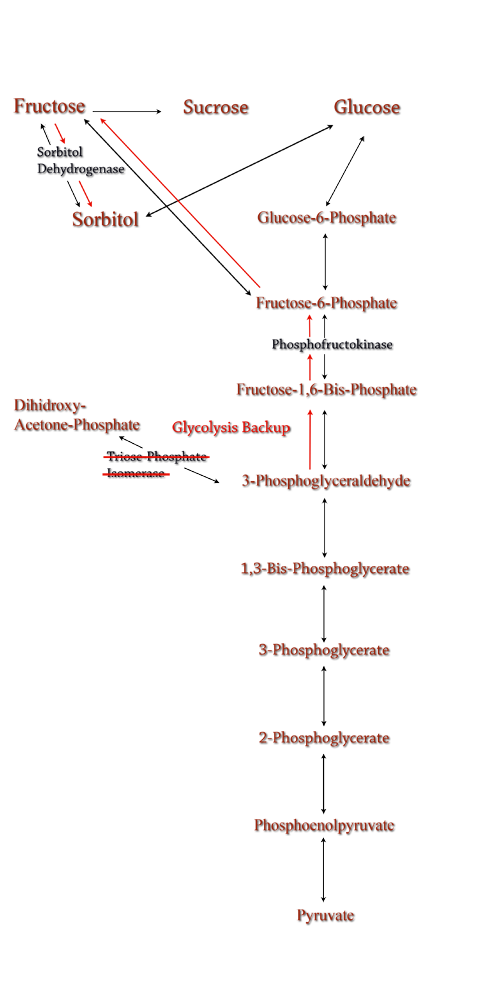
| Sorbitol anabolism is part of the glycolysis pathway in E.coli. In the pathway, interconversion between fructose and sorbitol is catalyzed by sorbitol dehydrogenase, a monomeric enzyme that uses NADH in the process. Natural expression of sorbitol dehydrogenase is generally up-regulated only in the presence of sorbitol as a carbon source. The initial idea to stimulate sorbitol over-production was to simply up-regulate the gene encoding for sorbitol dehydrogenase in an environment lacking sorbitol thus pushing the reaction governing fructose and sorbitol interconversion towards sorbitol.
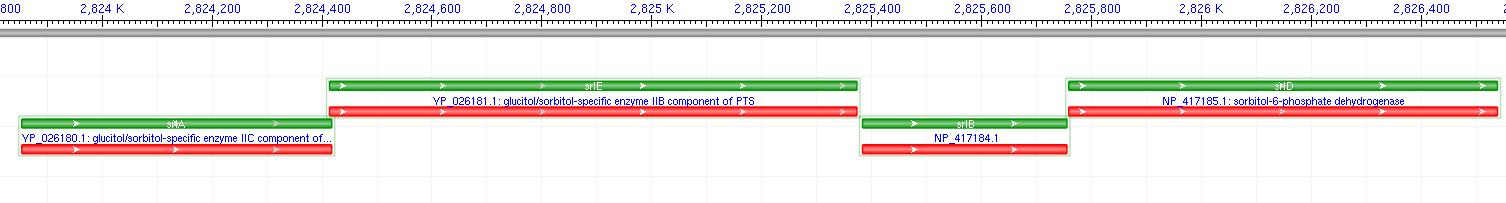
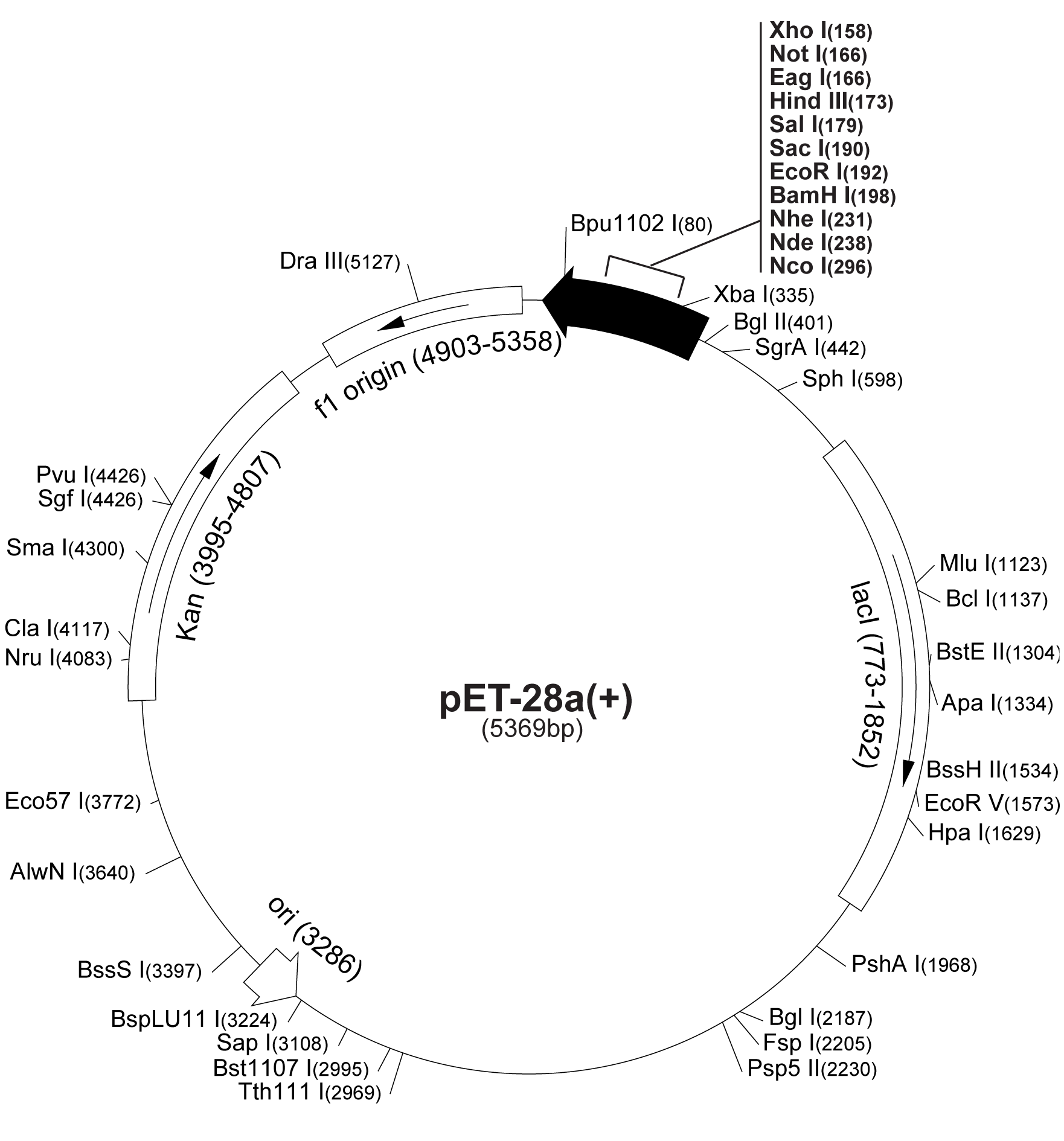
The gene encoding for sorbitol dehydrogenase(SDH), srlD, is part of the srl Operon in E.coli (Operon pictured below). Information on the SDH assembly pathway and enzymatic activity is limited. Consequently we decided that insertion of both the srlD gene and srl operon into vectors would take more time initially but would potentially be beneficial due to the chance that the gene alone would not result in effective SDH production. Increased cellular production of SDH would most likely increase the amount of sorbitol produced by the cell, however it could not alone account for the type of mass production that would make sorbitol biosynthesis commercially useful. In order to determine other methods of increasing sorbitol production, our group turned to computer modeling.
|
| Triose Phosphate Isomerase Knockout |
|
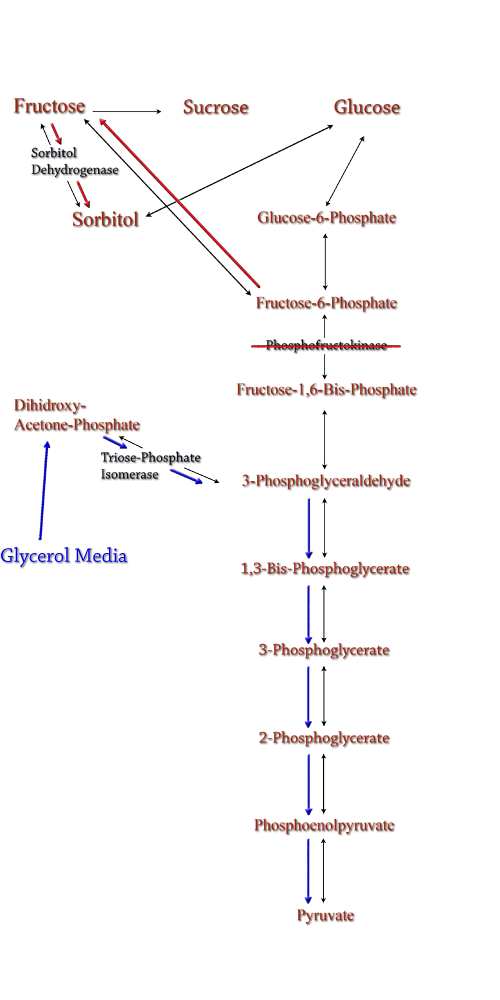
Computer modeling was done to determine gene knockouts that increased cellular flux towards sorbitol production. The model was given to us by Dr. Jenni Reed of University of Wisconsin-Madison. More extensive information about modeling can be found at our modeling page. Through the use of our model we determined that knocking out the gene encoding for triose phosphate isomerase would be the most effective at increasing sorbitol production when combined with srl gene up-regulation. Triose phosphate isomerase(TPI) is another enzyme in E.coli's glycolysis pathway. It's function is to catalyze the interconversion of 3-Phosphoglyceraldehyde to Dihidroxy-acetone-phosphate and back. Knocking out the gene which encodes TPI |
 "
"