Team:Caltech/Project
From 2008.igem.org
Caltechigem (Talk | contribs) |
Caltechigem (Talk | contribs) |
||
| Line 23: | Line 23: | ||
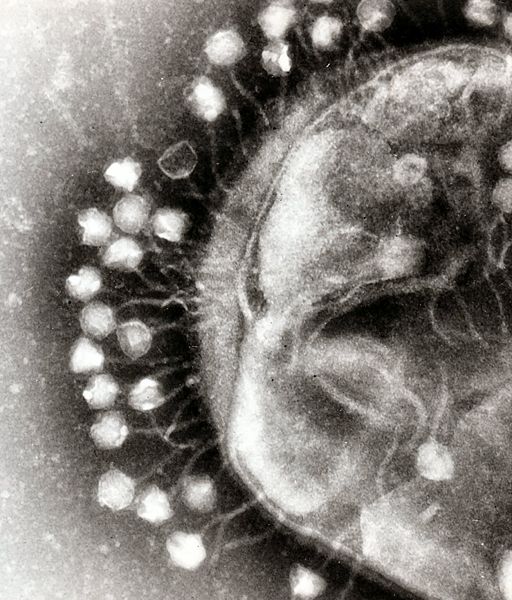
Another aspect of bacterial pathogen defense for our probiotic is to produce bacteriophages that would rapidly infect and wipe out all of the pathogens. We examined two methods to approach phage production, differentiated by the type of phage used. The first method uses the bacteriophage λ, which targets ''Escherichia coli''. The other method explores the use of a temperate bacteriophage from ''Bacillus subtilis''. The latter method, if successful, can be adapted to temperate bacteriophages of any bacterial strain. | Another aspect of bacterial pathogen defense for our probiotic is to produce bacteriophages that would rapidly infect and wipe out all of the pathogens. We examined two methods to approach phage production, differentiated by the type of phage used. The first method uses the bacteriophage λ, which targets ''Escherichia coli''. The other method explores the use of a temperate bacteriophage from ''Bacillus subtilis''. The latter method, if successful, can be adapted to temperate bacteriophages of any bacterial strain. | ||
| - | Bacteriophage λ is a temperate phage with an ''E. coli'' host. λ infects ''E. coli'' through the lamB receptor, where absence of this receptor prevents λ infection. We will | + | Bacteriophage λ is a temperate phage with an ''E. coli'' host. λ infects ''E. coli'' through the lamB receptor, where absence of this receptor prevents λ infection. We will take advantage of this property of bacteriophage λ to engineer an ''E. coli'' strain that is resistant to the phage, but harbors a phage in its genome and releases it to destroy susceptible pathogenic ''E. coli''. |
The second approach is more versatile, and will enable targeting of a broad range of pathogenic bacteria. The goal is to construct a phasmid out of the genome of a temperate bacteriophage. A phasmid is built by combining an ''E. coli'' plasmid origin of replication with a linear phage genome and then circularizing the DNA. The phasmid can then replicate and be maintained as a plasmid within ''E. coli''; however, when transferred to its native host, the phage phage will be induced. | The second approach is more versatile, and will enable targeting of a broad range of pathogenic bacteria. The goal is to construct a phasmid out of the genome of a temperate bacteriophage. A phasmid is built by combining an ''E. coli'' plasmid origin of replication with a linear phage genome and then circularizing the DNA. The phasmid can then replicate and be maintained as a plasmid within ''E. coli''; however, when transferred to its native host, the phage phage will be induced. | ||
| Line 41: | Line 41: | ||
{{!}}-valign="top" | {{!}}-valign="top" | ||
{{!}} | {{!}} | ||
| - | Folate, a term which encompasses the various forms of the vitamin B9, is an essential vitamin involved in everyday cell functions such as DNA replication. Unable to naturally produce folate, humans must obtain it from vegetables or folate-supplements. In regions with little or no access to these foods, folate deficiencies can cause serious birth defects. One possible solution to alleviate the effects of folate deficiency is to engineer a strain of gut microbes to produce bioavailable folate directly in the colon. | + | Folate, a term which encompasses the various forms of the vitamin B9, is an essential vitamin involved in everyday cell functions such as DNA replication. Unable to naturally produce folate, humans must obtain it from vegetables or folate-supplements. In regions with little or no access to these foods, folate deficiencies can cause serious birth defects. One possible solution to alleviate the effects of folate deficiency is to engineer a strain of gut microbes to produce bioavailable folate directly in the colon. We tested a total of four heterologous genes, two from the folate biosynthesis gene cluster and two from the paraaminobenzoic acid (pABA) synthesis pathway. Using standardized genetic sequences, folate biosynthesis genes extracted from the ''Lactoccocus lactis'' genome were cloned into Biobricks plasmids, transformed into ''Escherichia coli'' and overexpressed. We measured the effects of overexpression |
| - | + | in terms of total folate and paraaminobenzoic acid levels. PABA, an intermediate in folate synthesis, was detected using [[Team:Caltech/Protocols/PABA_HPLC_assay|<font style="color:#BB4400">high performance liquid chromatography</font>]] (HPLC). Folate detection was achieved via a [[Team:Caltech/Protocols/Folate_assay|<font style="color:#BB4400">microbiological assay</font>]]. A measurable increase in folate production in ''E. coli'' provides proof-of-concept for both the feasibility of engineering overproduction of folate in ''E. coli'', as well as using standardized genetic components to do so. | |
{{!}}{{navimg|xsize=193|ysize=288|image=Folate_foods.jpg|link=Team:Caltech/Project/Vitamins}} | {{!}}{{navimg|xsize=193|ysize=288|image=Folate_foods.jpg|link=Team:Caltech/Project/Vitamins}} | ||
{{!}}} | {{!}}} | ||
Revision as of 07:00, 25 October 2008
|
People
|
Subprojects
Note: Click on the subproject title or picture for a detailed description of the subproject
Oxidative Burst
Phage Pathogen Defense
Lactose Intolerance
Vitamin Production
Population Variation
|
 "
"