Team:Guelph/Results
From 2008.igem.org
m |
m |
||
| (27 intermediate revisions not shown) | |||
| Line 16: | Line 16: | ||
|} | |} | ||
| + | == '''Synthetic Operon''' == | ||
| + | |||
| + | We got the pDSK-GFPuv plasmid from the Noble Foundation and signed an MTA for its use in the iGEM competition. We PCRed the strong 250 bp consitutive promoter from this plasmid (its the 16S ribosomal promoter from the chloroplast of an herbicide resistant type of Amaranthus weed) and put it into the promoter testing device, BBa_E0240 and it does indeed prove to be a strong promoter: | ||
| + | |||
| + | {|align="center" | ||
| + | |[[Image:PSBAinpSB1A2-E0240.jpg|300px]] | ||
| + | | | ||
| + | |} | ||
| + | |||
| + | Knowing we had at least this biobricked strong promoter to work with, we went on building the synthetic operon in the pSB1A2 biobrick plasmid. Using the Phusion proofreading taq kindly provided to us by NEB and primers from IDT, we PCR amplified the crtE, B, and Y genes from ''Erwinia uredevora'' that were shared with us by members of iGEM Minnesota. Biobricked crt I was sent to us by iGEM Edinburgh. These were sequenced and test digested to confirm insert size. The gel photo showing Eco/Pst digests of the constructs in pSB1A2 is below. | ||
| + | |||
| + | |||
| + | {|align="center" | ||
| + | |[[Image:Crtdigest.jpg|500px]] | ||
| + | | | ||
| + | |} | ||
| + | |||
| + | |||
| + | |||
| + | Next, we ligated crt B to E, and crt Y to I. E and I were cut with SpeI and Pst, while B and Y were cut with XbaI and PstI. To cap off the operon, we used the same strategy; to restrict and ligate the biobrick GFP (plus terminators and rbs) from BBa_E0240 onto the end of the operon - you can see the approximately 1000 bp size increase on the gel above. We also ligated the 1.5 kb frdBCD operon to this GFP in vitro, then PCR fished for the frdBCD plus GFP using a frdBCD fwd primer and VR primer. This PCR product was cut with XbaI and PstI and put onto the end of the CrtE,B,I,Y operon but was not thoroughly tested by October 29th. | ||
| + | |||
| + | |||
| + | The last cloning steps were to put promoters onto the constructs, test them for carotenoid production and GFP fluorescence. We definately didn't get a chance to test the operon with frdBCD in it, but crtEBCD with just GFP at the end did work (it seems). Both the constitutive promoter, PSBA and the Biobrick arabinose inducible promoter (BBa_R0080) were put in control of this operon, with similar results (GFP expression and orange pigment production). | ||
| + | |||
| + | |||
| + | {|align="center" | ||
| + | |[[Image:synop.jpg|400px]] | ||
| + | |[[Image:synopgfp.jpg|400px]] | ||
| + | | | ||
| + | |} | ||
| + | |||
| + | |||
| + | Testing these constructs in ''E. coli'' Nissler 1917 will be attempted in anaerobic conditions after this website freezes, but lactobacilli have proven to be beyond the time frame of the possible for us this year. This quick intestinal simulation will hopefully make it on our presentation and poster, but the website won't show it. Talk to us at the Jamboree to see if we got good results or not! | ||
| + | |||
| + | {|align="center" | ||
| + | |[[Image:guelph intestine.jpg|300px]] | ||
| + | | | ||
| + | |} | ||
| + | |||
| + | |||
| + | == '''RNAi inducing corn endophytes (BIGS)''' == | ||
| + | |||
| Line 24: | Line 66: | ||
We plan to do wet lab modelling. By tagging our microbes with simple GFP constructs and counting colony forming units per set sample fresh weight, we hope to be able to observe microbial survival with our construct and predict how much beta carotene or RNAi might be produced and released. | We plan to do wet lab modelling. By tagging our microbes with simple GFP constructs and counting colony forming units per set sample fresh weight, we hope to be able to observe microbial survival with our construct and predict how much beta carotene or RNAi might be produced and released. | ||
| - | In corn, we are doing this using a corn endophyte called Klebsiella pneumonii. Plants were either injected with | + | In corn, we are doing this using a corn endophyte called ''Klebsiella pneumonii'' strain 342. Using protocols for E. coli electroporation, these microbes were transformed with the gram negative broad host range GFP plasmid GFPuv. Plants were either injected twice with 10 ul of bacterial suspension or dipped in it at an early stage. Two or four weeks later 500 mg of tissue were harvested, ground in sterilized mortars, resuspended in 500 ul of sodium phosphate buffer, and 50 ul of the dilution was spread on half of a kanamycin LB plate. The results are depicted graphically below. |
| + | |||
| + | {|align="center" | ||
| + | |[[Image:KP342 GFP assay in corn.jpg|700px]] | ||
| + | | | ||
| + | |} | ||
| + | |||
| + | We injected 20 ul of a bacterial OD of 0.85 we estimated to amount to be 1.07 million microbes based on colony forming unit counts (CFU). The above plates show a one fold dilution of homogenized plant tissue. This suggests that corn leaf tissue was harbouring 12 microbes per milligram of tissue (both root types had a similar number), while stem tissue near the site of injection had a number closer to 500. It is difficult to extrapolate accurately how many microbes might exist in the plant, know whether these are dispersed survivors from the injection or recent generations, but at the very least this suggests the microbial population we injected spread systematically and are stabily transcribing a transgene on a plasmid inside this plant for a significant period of time. | ||
| + | |||
| + | |||
| + | We observed similar result in ''Arabidopsis thaliana'', which may indicate this would be a useful transgene delivery tool for ''Arabidopsis'' research. | ||
| + | |||
| + | |||
| + | {|align="center" | ||
| + | |[[Image:arab342.jpg|400px]] | ||
| + | | | ||
| + | |} | ||
| + | |||
| + | RNAi constructs against TB1 and GFP were successfully made and tested by PCR for successful electroporation into Kp342. The 750 bp TB1 construct (two 250 bp TB1 fragments oriented sense and antisense around the central 100 bp ''Zea mays'' Actin1 intron) was cut with NdeI and PstI and ligated into the NdeI and PstI cut pDSK-GFPuv, trading places with the plasmid's GFP. For the GFP RNAi construct, the 1000 bp biobrick from BBa_E0240 was used, which is a composite part containing rbs and terminators. Here the PSBA promoter was introduced infront of the construct in the pSB1A2 plasmid, and the Eco/Pst casssette then removed and inserted into the pDK-GFPuv plasmid. | ||
| + | |||
| + | The TB1 RNAi construct was successfully electroporated into ''Klebsiella pneumonii'' 342 using common electroporation protocols for ''E. coli'' which was verified by colony PCR. This RNAi signal producing endophyte was then injected into corn plants to acertain it potential to silence a plant gene inhibiting branch formation. These plants were grown alongside tb1 mutant corn, and GFP tagged kp342 for two weeks, at which time they were to be injected again. | ||
| + | |||
| + | {|align="center" | ||
| + | |[[Image:Injectcorn.jpg|500px]] | ||
| + | | | ||
| + | |} | ||
| + | |||
| + | At this time the plants have still not reached a stage where phenotypes appear to be visible - see the below photo which was taken 2 weeks post injection. :-( Also, the GFP RNAi construct in pDSK-GFPuv while maintained in ''E. coli'', was not successfully introduced into kp342 despite three attempts, so the trial on GFP silencing proceeded with ''E. coli'' instead. Work and monitoring will continue up until and after the jamboree - BIGS will not be silenced!!! | ||
| + | |||
| + | {|align="center" | ||
| + | |[[Image:tb1plants.jpg|500px]] | ||
| + | | | ||
| + | |} | ||
| - | + | Black and white corn and tb1 mutant image reproduced from Nature Reviews Genetics 3, 11-21 (January 2002). | |
Latest revision as of 07:35, 30 October 2008
| Home | The Team | The Project | Parts | Notebook | Results | Links |
|---|
Synthetic Operon
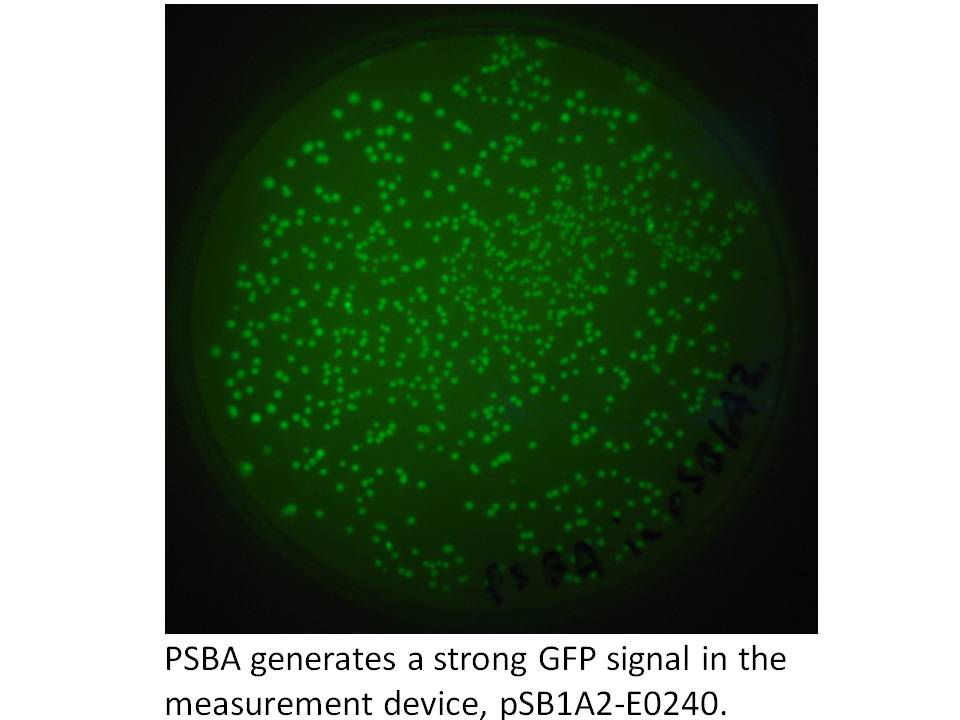
We got the pDSK-GFPuv plasmid from the Noble Foundation and signed an MTA for its use in the iGEM competition. We PCRed the strong 250 bp consitutive promoter from this plasmid (its the 16S ribosomal promoter from the chloroplast of an herbicide resistant type of Amaranthus weed) and put it into the promoter testing device, BBa_E0240 and it does indeed prove to be a strong promoter:
|
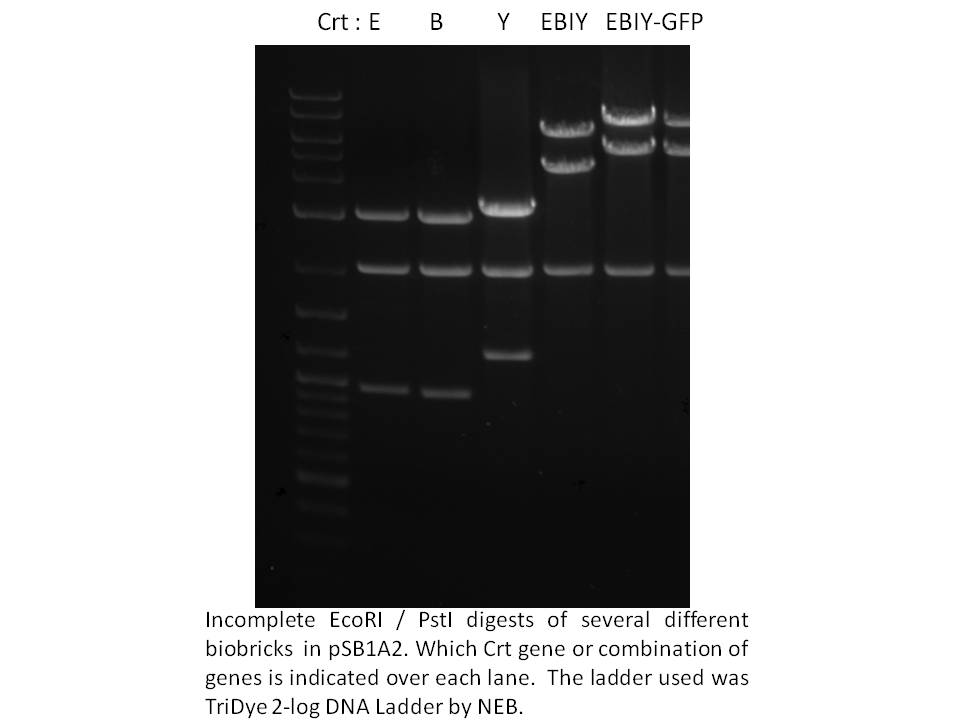
Knowing we had at least this biobricked strong promoter to work with, we went on building the synthetic operon in the pSB1A2 biobrick plasmid. Using the Phusion proofreading taq kindly provided to us by NEB and primers from IDT, we PCR amplified the crtE, B, and Y genes from Erwinia uredevora that were shared with us by members of iGEM Minnesota. Biobricked crt I was sent to us by iGEM Edinburgh. These were sequenced and test digested to confirm insert size. The gel photo showing Eco/Pst digests of the constructs in pSB1A2 is below.
|
Next, we ligated crt B to E, and crt Y to I. E and I were cut with SpeI and Pst, while B and Y were cut with XbaI and PstI. To cap off the operon, we used the same strategy; to restrict and ligate the biobrick GFP (plus terminators and rbs) from BBa_E0240 onto the end of the operon - you can see the approximately 1000 bp size increase on the gel above. We also ligated the 1.5 kb frdBCD operon to this GFP in vitro, then PCR fished for the frdBCD plus GFP using a frdBCD fwd primer and VR primer. This PCR product was cut with XbaI and PstI and put onto the end of the CrtE,B,I,Y operon but was not thoroughly tested by October 29th.
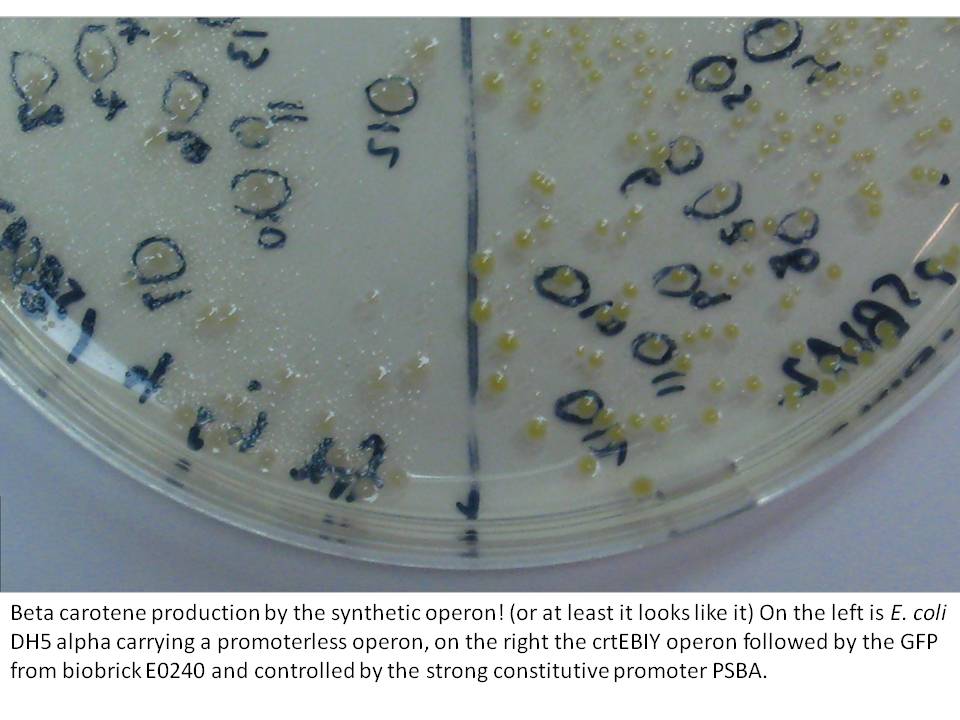
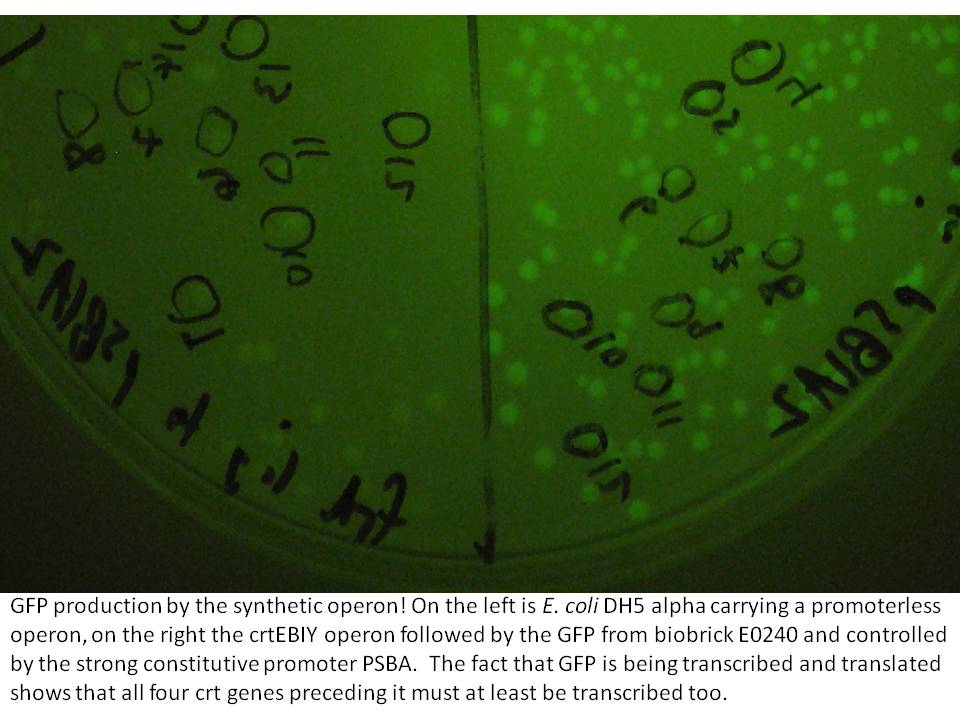
The last cloning steps were to put promoters onto the constructs, test them for carotenoid production and GFP fluorescence. We definately didn't get a chance to test the operon with frdBCD in it, but crtEBCD with just GFP at the end did work (it seems). Both the constitutive promoter, PSBA and the Biobrick arabinose inducible promoter (BBa_R0080) were put in control of this operon, with similar results (GFP expression and orange pigment production).
| 
|
Testing these constructs in E. coli Nissler 1917 will be attempted in anaerobic conditions after this website freezes, but lactobacilli have proven to be beyond the time frame of the possible for us this year. This quick intestinal simulation will hopefully make it on our presentation and poster, but the website won't show it. Talk to us at the Jamboree to see if we got good results or not!
|
RNAi inducing corn endophytes (BIGS)
Making projections based on measurements on endosymbiont and endophyte numbers within a functioning biological system using GFP
We plan to do wet lab modelling. By tagging our microbes with simple GFP constructs and counting colony forming units per set sample fresh weight, we hope to be able to observe microbial survival with our construct and predict how much beta carotene or RNAi might be produced and released.
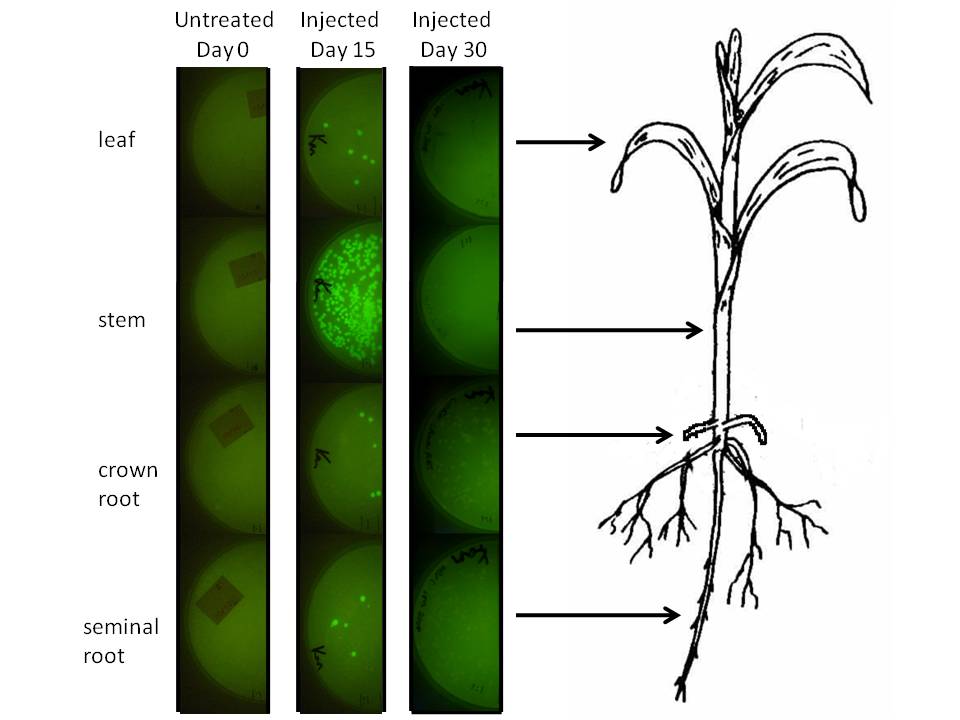
In corn, we are doing this using a corn endophyte called Klebsiella pneumonii strain 342. Using protocols for E. coli electroporation, these microbes were transformed with the gram negative broad host range GFP plasmid GFPuv. Plants were either injected twice with 10 ul of bacterial suspension or dipped in it at an early stage. Two or four weeks later 500 mg of tissue were harvested, ground in sterilized mortars, resuspended in 500 ul of sodium phosphate buffer, and 50 ul of the dilution was spread on half of a kanamycin LB plate. The results are depicted graphically below.
|
We injected 20 ul of a bacterial OD of 0.85 we estimated to amount to be 1.07 million microbes based on colony forming unit counts (CFU). The above plates show a one fold dilution of homogenized plant tissue. This suggests that corn leaf tissue was harbouring 12 microbes per milligram of tissue (both root types had a similar number), while stem tissue near the site of injection had a number closer to 500. It is difficult to extrapolate accurately how many microbes might exist in the plant, know whether these are dispersed survivors from the injection or recent generations, but at the very least this suggests the microbial population we injected spread systematically and are stabily transcribing a transgene on a plasmid inside this plant for a significant period of time.
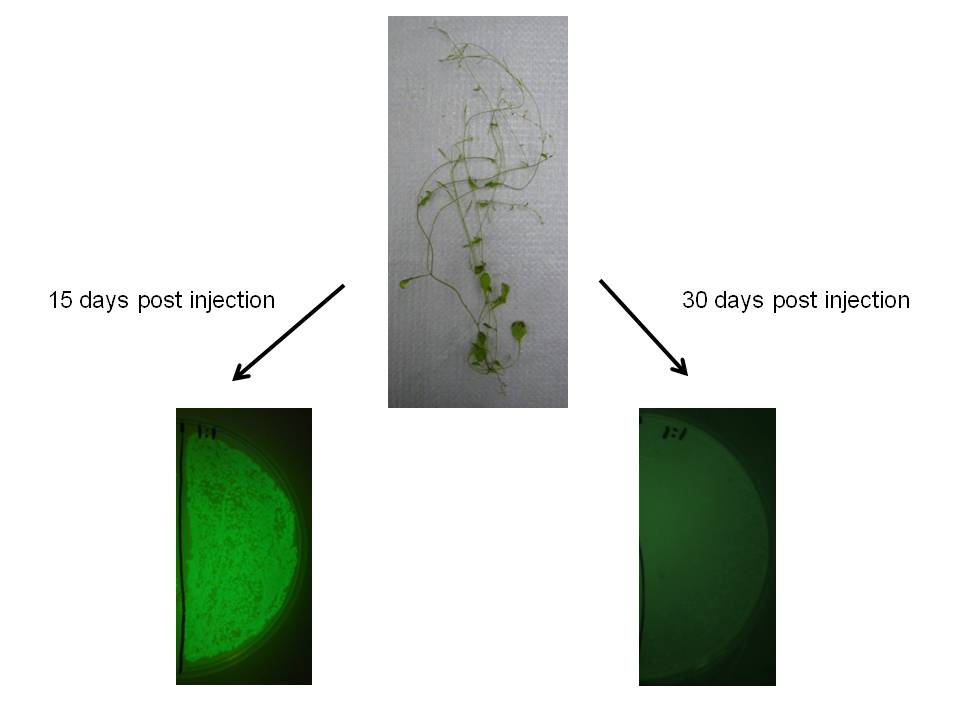
We observed similar result in Arabidopsis thaliana, which may indicate this would be a useful transgene delivery tool for Arabidopsis research.
|
RNAi constructs against TB1 and GFP were successfully made and tested by PCR for successful electroporation into Kp342. The 750 bp TB1 construct (two 250 bp TB1 fragments oriented sense and antisense around the central 100 bp Zea mays Actin1 intron) was cut with NdeI and PstI and ligated into the NdeI and PstI cut pDSK-GFPuv, trading places with the plasmid's GFP. For the GFP RNAi construct, the 1000 bp biobrick from BBa_E0240 was used, which is a composite part containing rbs and terminators. Here the PSBA promoter was introduced infront of the construct in the pSB1A2 plasmid, and the Eco/Pst casssette then removed and inserted into the pDK-GFPuv plasmid.
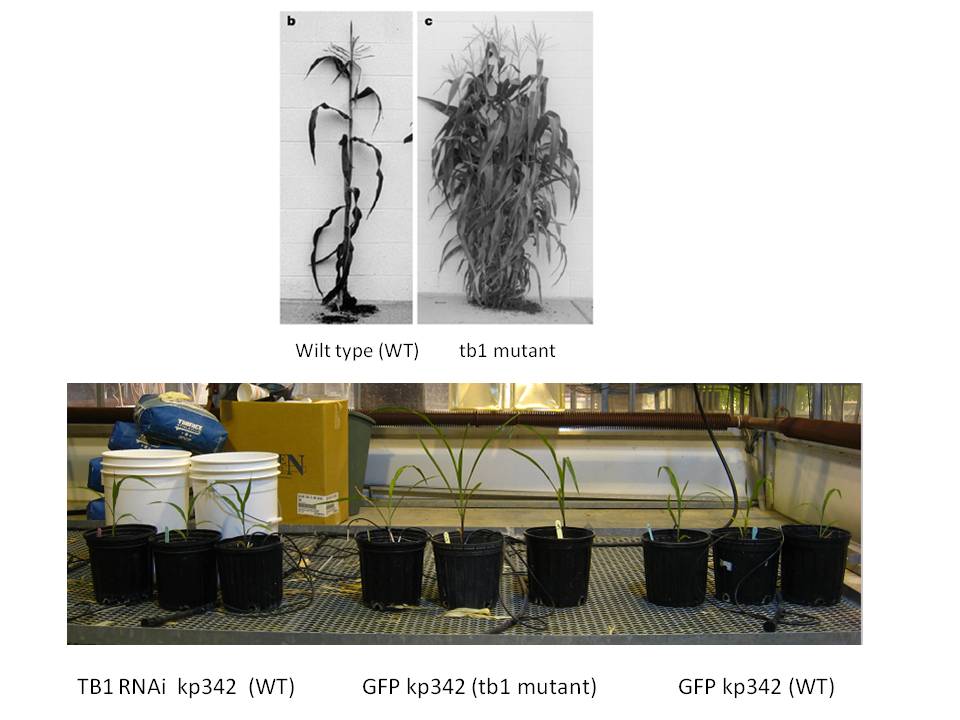
The TB1 RNAi construct was successfully electroporated into Klebsiella pneumonii 342 using common electroporation protocols for E. coli which was verified by colony PCR. This RNAi signal producing endophyte was then injected into corn plants to acertain it potential to silence a plant gene inhibiting branch formation. These plants were grown alongside tb1 mutant corn, and GFP tagged kp342 for two weeks, at which time they were to be injected again.

|
At this time the plants have still not reached a stage where phenotypes appear to be visible - see the below photo which was taken 2 weeks post injection. :-( Also, the GFP RNAi construct in pDSK-GFPuv while maintained in E. coli, was not successfully introduced into kp342 despite three attempts, so the trial on GFP silencing proceeded with E. coli instead. Work and monitoring will continue up until and after the jamboree - BIGS will not be silenced!!!

|
Black and white corn and tb1 mutant image reproduced from Nature Reviews Genetics 3, 11-21 (January 2002).
 "
"