Team:Newcastle University
From 2008.igem.org
m |
|||
| (74 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | |||
| - | |||
| - | |||
| - | |||
{{:Team:Newcastle_University/Template:Header}} | {{:Team:Newcastle_University/Template:Header}} | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
<div id="maincol"> | <div id="maincol"> | ||
| - | + | '''Welcome!''' | |
| - | + | This year is Newcastle's first year participating in the iGEM competition! We are a small [[Team:Newcastle_University/Team|team]] of six students, three supervisors, and seven advisors, but we had high ambitions for our first year. Recognizing the problem of antibiotic-resistant bacteria and poor methods of diagnosis in developing countries, we aim to use synthetic biology and bioinformatics to engineer a simple, safe, fast, and reliable biological diagnostic system to identify bacteria. | |
| - | + | {{:Team:Newcastle University/Template:BlankBox | |
| - | + | |boxtype=bluebox | |
| - | + | |title=Overview | |
| - | {{:Team:Newcastle University/Template: | + | |detail-text= |
| - | | | + | We aimed to develop a diagnostic biosensor for detecting pathogens. We wanted this to be cheaply and readily available for deployment in areas where access to medical resources, such as refrigeration and sophisticated laboratories, is limited or absent. We chose to use [[Team:Newcastle_University/Bacillus_subtilis|''Bacillus subtilis'']] as a method of delivery due to its ability to sporulate. The sensor bacteria could then be dried down as spores, which are very stable and extremely resilient to hostile environmental conditions, and rehydrated when required. The ambient temperature of much of the developing world is ideal for the growth of ''Bacillus spp.'' without the use of incubation equipment. |
| - | | | + | |
| - | | | + | |
| - | + | ||
| - | | | + | |
| - | + | ||
| - | + | Gram-positive bacteria communicate using quorum communication peptides. Research has shown that these peptides are extremely strain-specific. We chose to engineer ''B. subtilis'' 168 to detect four Gram-positive pathogens by their quorum communication peptides. The different combinations of quorum communication peptides would be sensed by the engineered bacterium, and this signal converted into a visual output as fluorescent proteins such as mCherry, GFP, CFP and YFP. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| + | Mapping multiple inputs to three output states is a multiplexing problem. The design of the genetic circuitry to do this is non-trivial and is not feasible manually. We therefore chose to use a biological implementation of an artificial neural network (ANN). Our team members wrote, designed and implemented a complete suite of tools that allowed the design and simulation of regulatory networks. We aimed to use this software to design an appropriate gene regulatory network that behaved as an ANN. An essential part of the approach was the use of computational evolution to design circuits with predictable behaviour even when the details of the required topology are unknown in advance. Models were encoded in the modelling language CellML because of its ability to model individual virtual parts that can be easily assembled, in a bottom up fashion, into a circuit which can be simulated. We aimed to translate this model into a sequence that could be implemented as BioBricks. | ||
| - | + | [[Team:Newcastle_University/Original Aims|Learn more about our project]] }} | |
| - | + | ||
| - | + | ||
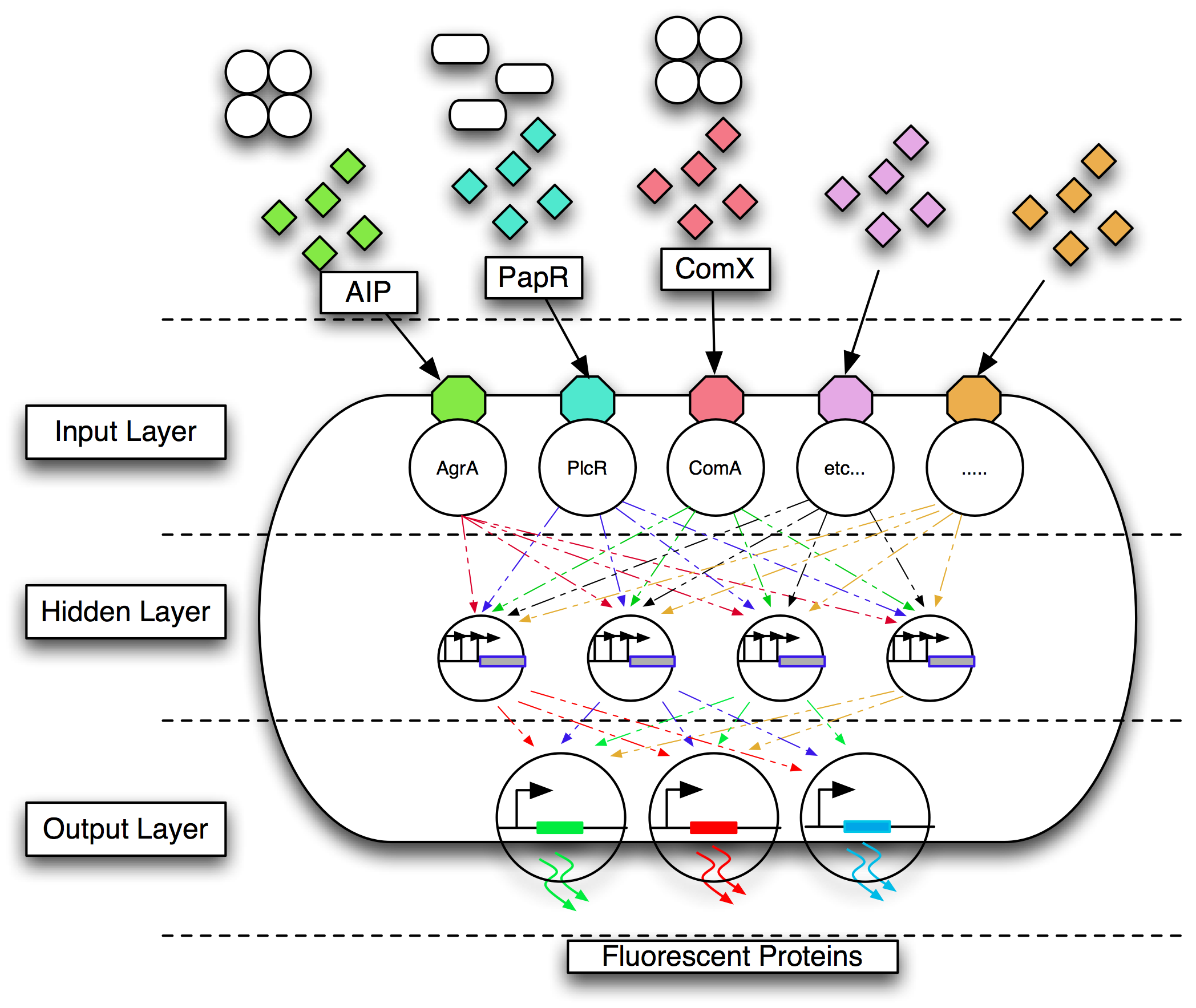
| - | + | [[Image:Bug.png|center|500px]] | |
| - | + | <center>Conceptual overview of the implementation of an ANN in a bacterium.</center> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | {{:Team:Newcastle University/Template: | + | {{:Team:Newcastle University/Template:BlankBox |
|boxtype=bluebox | |boxtype=bluebox | ||
| - | |title= | + | |title=Achievements |
| - | |detail-text= | + | |detail-text=<ul> |
| - | + | ||
| - | + | <li>We designed and implemented a software framework for the computational design of genetic regulatory circuits. | |
| - | | | + | <li>We designed, modelled and submitted a [http://partsregistry.org/Part:BBa_K104001 working standard BioBrick] for sensing the quorum communication peptide subtilin, that [[Team:Newcastle University/Notebook|works as expected]]. |
| - | | | + | <li>Sent information and developed a [[Team:Newcastle_University/Bacillus_subtilis|''B. subtilis'' website]] to help the Cambridge University team. |
| - | + | <li>Took the Cambridge 2007 [http://partsregistry.org/wiki/index.php?title=Part:BBa_I746107 BBa_I746107] AIP-inducible promoter P2 and GFP reporter, cloned it into an integration vector and successfully integrated it into the chromosome of 168, ready for further characterization.</li> | |
| - | + | <li>Developed and documented a new technical standard using CellML for the bottom up modelling, simulation and composition of BioBrick parts, from virtual parts. | |
| + | </ul> | ||
| + | }} | ||
| - | + | == Sponsors == | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | We gratefully thank our sponsors: | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | {{:Team:Newcastle University/Sponsors}} | |
| - | + | {{:Team:Newcastle University/HomeSideBar}} | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Latest revision as of 22:14, 29 October 2008
Newcastle University
GOLD MEDAL WINNER 2008
| Home | Team | Original Aims | Software | Modelling | Proof of Concept Brick | Wet Lab | Conclusions |
|---|
Welcome! This year is Newcastle's first year participating in the iGEM competition! We are a small team of six students, three supervisors, and seven advisors, but we had high ambitions for our first year. Recognizing the problem of antibiotic-resistant bacteria and poor methods of diagnosis in developing countries, we aim to use synthetic biology and bioinformatics to engineer a simple, safe, fast, and reliable biological diagnostic system to identify bacteria.
Overview
We aimed to develop a diagnostic biosensor for detecting pathogens. We wanted this to be cheaply and readily available for deployment in areas where access to medical resources, such as refrigeration and sophisticated laboratories, is limited or absent. We chose to use Bacillus subtilis as a method of delivery due to its ability to sporulate. The sensor bacteria could then be dried down as spores, which are very stable and extremely resilient to hostile environmental conditions, and rehydrated when required. The ambient temperature of much of the developing world is ideal for the growth of Bacillus spp. without the use of incubation equipment.
Gram-positive bacteria communicate using quorum communication peptides. Research has shown that these peptides are extremely strain-specific. We chose to engineer B. subtilis 168 to detect four Gram-positive pathogens by their quorum communication peptides. The different combinations of quorum communication peptides would be sensed by the engineered bacterium, and this signal converted into a visual output as fluorescent proteins such as mCherry, GFP, CFP and YFP.
Mapping multiple inputs to three output states is a multiplexing problem. The design of the genetic circuitry to do this is non-trivial and is not feasible manually. We therefore chose to use a biological implementation of an artificial neural network (ANN). Our team members wrote, designed and implemented a complete suite of tools that allowed the design and simulation of regulatory networks. We aimed to use this software to design an appropriate gene regulatory network that behaved as an ANN. An essential part of the approach was the use of computational evolution to design circuits with predictable behaviour even when the details of the required topology are unknown in advance. Models were encoded in the modelling language CellML because of its ability to model individual virtual parts that can be easily assembled, in a bottom up fashion, into a circuit which can be simulated. We aimed to translate this model into a sequence that could be implemented as BioBricks.
Achievements
- We designed and implemented a software framework for the computational design of genetic regulatory circuits.
- We designed, modelled and submitted a [http://partsregistry.org/Part:BBa_K104001 working standard BioBrick] for sensing the quorum communication peptide subtilin, that works as expected.
- Sent information and developed a B. subtilis website to help the Cambridge University team.
- Took the Cambridge 2007 [http://partsregistry.org/wiki/index.php?title=Part:BBa_I746107 BBa_I746107] AIP-inducible promoter P2 and GFP reporter, cloned it into an integration vector and successfully integrated it into the chromosome of 168, ready for further characterization.
- Developed and documented a new technical standard using CellML for the bottom up modelling, simulation and composition of BioBrick parts, from virtual parts.
Sponsors
We gratefully thank our sponsors:

|

|

|

|

|
 "
"