Constructs
Saccharomyces cerevisiae is widely used for baking and brewing, a versatile eukaryotic model system, and is particularly useful for synthesizing metabolites under fermentation conditions. Such microaerobic conditions prevent the air oxidation of bioreactive compounds and are optimal for the in vivo synthesis of resveratrol. To achieve our project and expand the synthetic biology toolbox for programming yeast, we have introduced into the iGem registry BioBricks encoding 3 yeast promoters, 3 yeast terminators, a two micron origin of replication, 2 selectable markers, 2 enzymes, and a yeast integration plasmid. In addition, we have generated seven constructs using these parts. Furthermore, we have submitted two additional parts representing a foundational tool, including a gene encoding an amber suppressed RFP biobrick for screening of SupF+ (Amber suppressor) genotype and an amber suppressor tRNA biobrick.
Yeast Promoters
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122000 BBa_K122000] |
pPGK1 |
This is the 1500 bp upstream of the PGK1 coding region in an industrial yeast strain. Constitutive promoter. |
1497 |
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122002 BBa_K122002] |
pADH1 |
700bp upstream of ADH1 promoter region containing RBS. Constitutive promoter. |
701 |
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122001 BBa_K122017] |
pGAL1 + tetO |
The GAL1 promoter which is highly repressed by glucose. An additional tetracycline operator site was included upstream of the RBS to allow repression by tetR. |
484 |
Yeast Terminators
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122003 BBa_K122003] |
tCYC1 |
300bp downstream the CYC1 coding region in a standard yeast strain. |
300 |
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122004 BBa_K122004] |
tADH1 |
300bp downstream the ADH1 coding region in a standard yeast strain. |
300 |
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122013 BBa_K122013] |
tPGK1 |
1000bp downstream the PGK1 coding region in an industrial yeast strain. |
1000 |
Selectable Markers
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122001 BBa_K122018] |
ZeoR |
Zeocin Resistance Gene |
300 |
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122008 BBa_K122008] |
BleoR |
Bleocin Resistance Gene under pTet promoter |
800ish |
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122014 BBa_K122014] |
ORI+HisTag |
2 Micron ORI and Histadine Tag |
<9000 |
Project Specific Constructs
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122001 BBa_K122001] |
[pGAL1][tetO][ZeoR] |
 |
874 |
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122005 BBa_K122005] |
Tyrosine Ammonia Lyase |
 |
1933 |
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122010 BBa_K122010] |
4CL:STS |
 |
4000 |
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122012 BBa_K122012] |
[pPGK1][4CL:STS][tCYC1] |
 |
5497 |
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122015 BBa_K122015] |
[pGAL1][tetO][ZeoR][tADH1] |
 |
1175 |
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122021 BBa_K122021] |
[pADH1][TAL][tPGK1] |
 |
2651 |
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122016 BBa_K122019] |
[pPGK1][4CL:STS][tCYC1][pGAL1][tetO][ZeoR][tADH1] |
 |
1824 |
Additional Bacterial Parts
Novel Zero Leak Inverter ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K122007 K122007] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122006 K122006])
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122007 K122007] |
The supF construct, an amber suppressor tRNA, allows for read-through at native amber (TAG) stop codons. |
211 |
| [http://partsregistry.org/wiki/index.php?title=Part:BBa_K122006 K122006] |
Point Mutation of RFP(13521) with incorporation of an amber stop codon at the chemophore. |
923 |
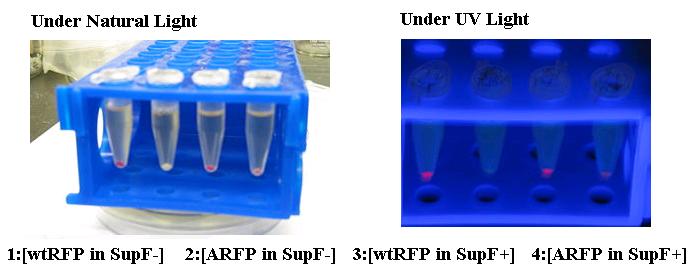
Through incorporation of amber stop codons or point mutations of tyrosine codons (TAC) to TAG within the coding region, a genetic circuit can be used to add an additional level of regulation and determine whether a full protein or partial peptide with be synthesized. This design has characteristically high signal-to-noise ratio, with virtually no leakage.
The Amber suppressed Red Florescent Protein (ARFP) has no virtually expression in SupF- (2) cells while visually apparent levels of red pigment is present in SupF+ cells (4). ARFP expression was compared against wtRFP expression in both cell types (1 and 3).
|
 "
"