Team:MIT
From 2008.igem.org
(→Lactobacillus Work) |
(→Lactobacillus Work) |
||
| Line 49: | Line 49: | ||
====Lactobacillus Work==== | ====Lactobacillus Work==== | ||
| - | We '''successfully transformed Lactobacillus delbruckii subsp. bulgaricus and Lactobacillus delbruckii subsp. lactis''' with a modified version of Serror et al's electrotransformation procedure. Our [http://openwetware.org/wiki/Lactobacillus_transformation '''transformation protocols'''] are found here. We also plan to submit these plasmids, with biobrick sites inserted in them, to the registry. These plasmids are [http://partsregistry.org/wiki/index.php?title=Part:BBa_K128008 '''pJK650'''] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K128007 '''pLEM415''']. | + | *We '''successfully transformed Lactobacillus delbruckii subsp. bulgaricus and Lactobacillus delbruckii subsp. lactis''' with a modified version of Serror et al's electrotransformation procedure. Our [http://openwetware.org/wiki/Lactobacillus_transformation '''transformation protocols'''] are found here. We also plan to submit these plasmids, with biobrick sites inserted in them, to the registry. These plasmids are [http://partsregistry.org/wiki/index.php?title=Part:BBa_K128008 '''pJK650'''] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K128007 '''pLEM415''']. |
| - | We are currently working on a method to effectively miniprep plasmid DNA from our ''Lactobacillus''. For information about our protocol for doing so, please click [http://openwetware.org/wiki/Lactobacillus_miniprep '''here''']. | + | *We are currently working on a method to effectively miniprep plasmid DNA from our ''Lactobacillus''. For information about our protocol for doing so, please click [http://openwetware.org/wiki/Lactobacillus_miniprep '''here''']. |
| - | We have built an expression system for Lactobacillus to secrete protein, including individual biobricked parts of a Lactobacillus [http://partsregistry.org/wiki/index.php?title=Part:BBa_K128006 '''promoter'''] and a Lactobacillus [http://partsregistry.org/Part:BBa_K128004 '''signal sequence''']. | + | *We have built an expression system for Lactobacillus to secrete protein, including individual biobricked parts of a Lactobacillus [http://partsregistry.org/wiki/index.php?title=Part:BBa_K128006 '''promoter'''] and a Lactobacillus [http://partsregistry.org/Part:BBa_K128004 '''signal sequence''']. |
| - | Please visit the results page of this wiki for full report of all our results and team-generated protocols!!!! | + | *Please visit the results page and experiments page of this wiki for full report of all our results and team-generated protocols!!!! |
==Acknowledgements== | ==Acknowledgements== | ||
Revision as of 03:58, 30 October 2008
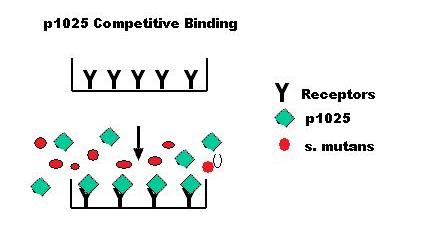
Biogurt: A Sustainable and Savory Drug Delivery SystemStreptococcus mutans, the main cause of dental caries, binds to glycoproteins on the teeth. A clinical study (Kelly CG et al.; Nature Biotechnol. 1999) isolated the 20aa functional segment (p1025) that S.mutans uses to attach to the teeth. p1025 competitively inhibits the binding of S.mutans, causing unharmful bacteria to grow in its place, preventing the recolonization of S.mutans for 90 days. We are engineering Lactobacillus bulgaricus, a bacteria commonly found in yogurt, to produce and secrete this peptide under a promoter activated by lactose. The peptide p1025 could simply be added to any food. But production of this peptide by L. bulgaricus is an independent process, so inserting the gene into live bacteria in yogurt will enable continuous production. Since a new batch of yogurt can be made using the bacteria from a small amount of the old batch, a continuous supply of teeth-cleaning yogurt will be available from the first successfully engineered batch. This could be the key to providing effective dental health care in underdeveloped rural communities, especially if yogurt is already an integral part of the diet. Also, the p1025 gene could be replaced by any other gene, so this same expression system could be used to produce other useful peptides. Yogurt with modified bacteria will provide a cheap, efficient, and delicious way to distribute vitamins, vaccines and more. ResultsCharacterization
Models
New Methods
Lactobacillus Work
AcknowledgementsGraduate AdvisersScott Carlson, Felix Moser, Chia-Yung Wu, Lav Varshney, Vikramaditya Yadav, Woo Chung, Rachel Hilmer, Robbier Barbero, Brian Cook, Jyoti Goda, Laure-Anne Ventouras Faculty Advisors:Drew Endy, Tom Knight OthersIsadora Deese, Dr. Pascale Serror at L'Institut National de la Recherche Agronomique, Dr. Bernhard Heinrich at the University of Kaiserslautern, Dr. Chris French at the University of Edinburgh, Dr. Daniel Smith at the Forsyth Institute, Joey Davis at the Sauer Lab at MIT, The Fink Lab at the Whitehead Institute, Emiko Bare at the Keating Lab at MIT, Jennifer Hou at the Baker Lab at MIT, The UROP office and Biological Engineering Department for financial support | ||||||||
|
| ||||||||
| Home | The Team | The Project | Experiments | Parts Submitted to the Registry | Modeling | Notebook |
|---|
 "
"