|
Home
The Team
Project Report
Parts
Modeling
Notebook
Safety
CoLABoration
|
_3D-modeling
DNA-Origami
Planning the input pattern on the DNA-Origami surface, we generated various 3D-models of the huge molecule using [ http://www.nanoengineer-1.com/content/ NanoEngineer], [http://pymol.sourceforge.net/ Pymol] and [http://spdbv.vital-it.ch/ SwissPdbViewer] (both freeware). Some of those models are shown here to give you an impression of the molecules we have engineered this year.
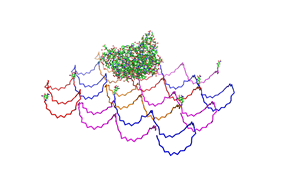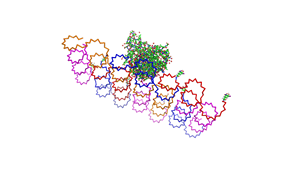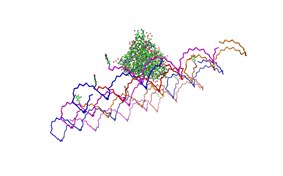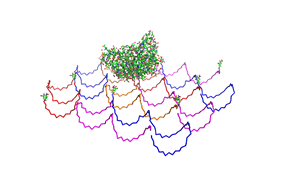 Fig.1: Some of the oligonucleotides that shape the single-stranded template fused to NIP-molecules and an anti-NIP-Fv-Fragment |
|
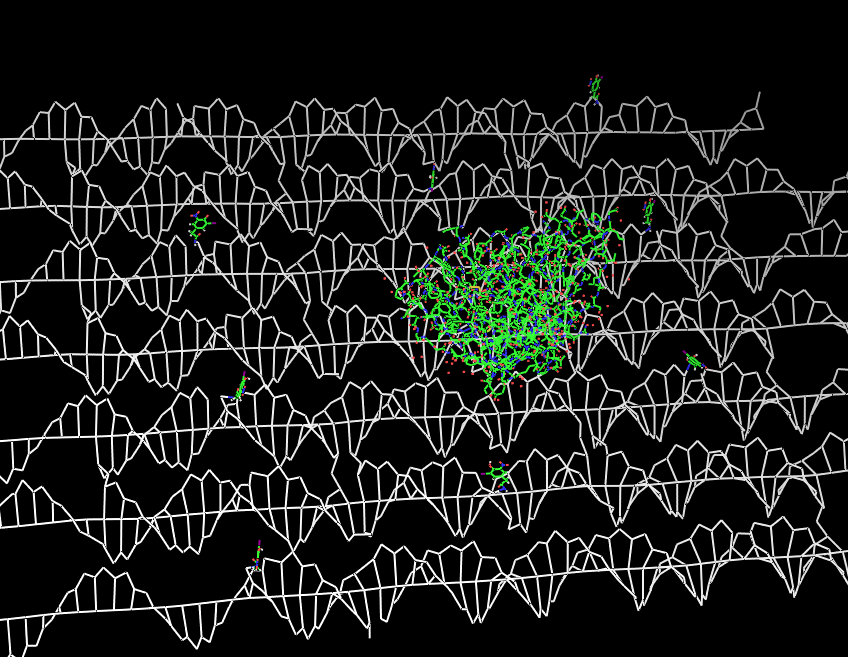 Fig.2: DNA-Origami with NIP-molecules and anti-NIP-fragment |
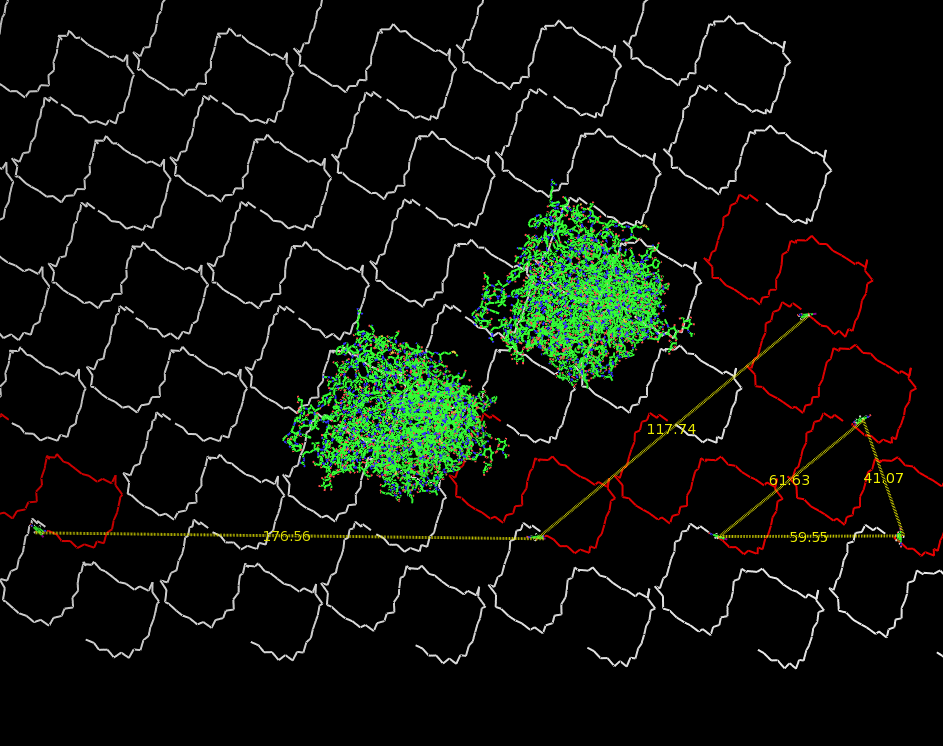 Fig.3: Oligo-pattern with scale and two fragments (distance estimation model) |
 Fig.4: Oligo-pattern with scale and two T-Cell-receptors (distance estimation model) |
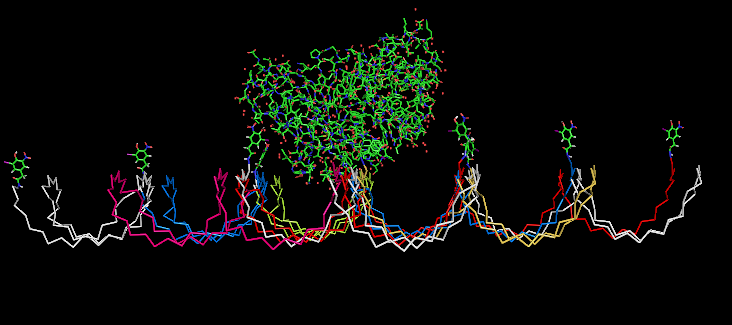 Fig.5: Oligo-pattern with NIP-molecules and Fab-fragment, binding accessibility estimation model
|
Fusion Proteins
The pictures below show models of the fusion proteins of our composite parts [http://partsregistry.org/wiki/index.php?title=Part:BBa_K157032 Bba_K157032] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K157033 Bba_K157033] code for (generated with "SwissPdbViewer"):
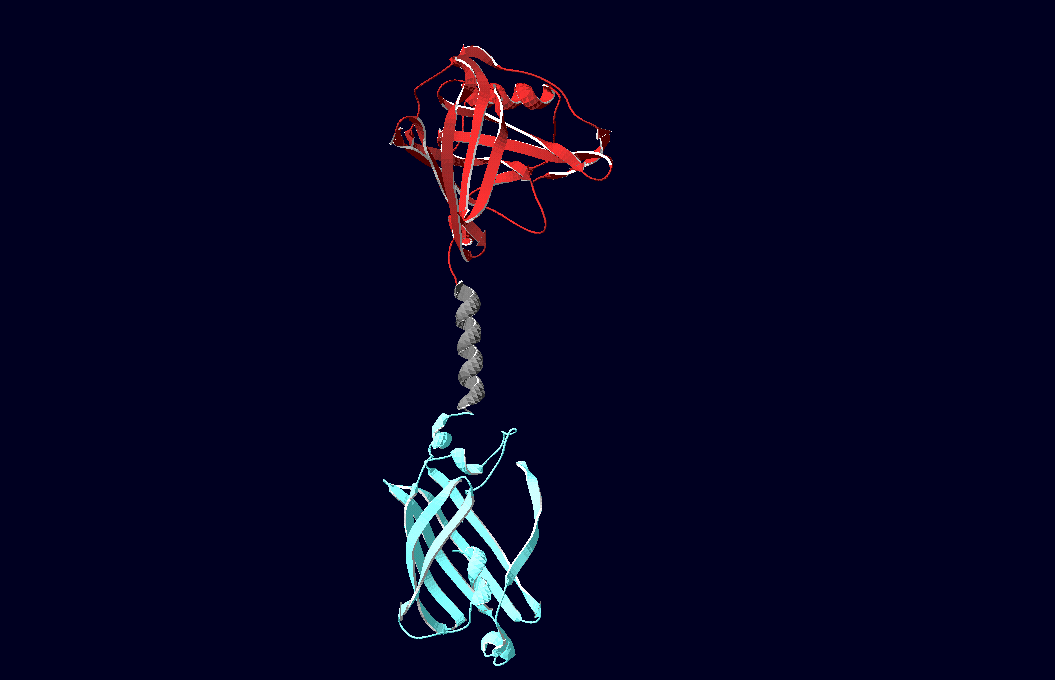 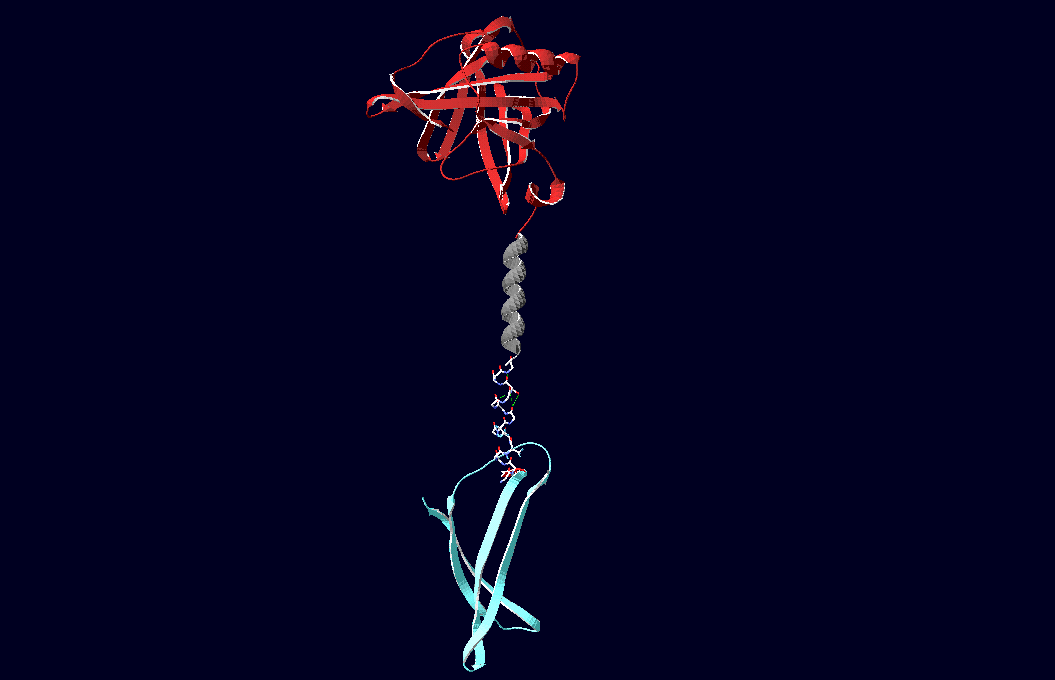
Fig.6: Red: LipocalinFluA, Grey: "Transmembrane region (BCR, no structural data; symbolized by helix), Blue: left: Split-Cerulean (CFP), N-terminal fragment; right: Split-Cerulean (CFP), C-terminal fragment
Following pdb-files have been used to generate the models:
Cerulean-CFP: 2Q57
Venus-YFP: 1MYW
Engineered Lipocalin: 1n0s
B1-8 Fv-Fragment:1a6w
|