Team:ETH Zurich/Wetlab/Chemostat Selection
From 2008.igem.org
|
Chemostat selectionGoalThe goal of our project is to find the minimal genome that is able to support survival of E. coli. In the previous section, we have introduced the method we want to apply for reducing the genome. However, manually selecting for those bacteria that have successfully reduced their genomes is obviously impossible. Therefore, we have to find a mechanism that automatically selects cells possessing a reduced genome.
IdeaOur idea is to set up a continuous bacterial culture in which cells possessing a reduced genome overgrow those maintaining more chromosomal DNA. However, bacteria with a smaller genome do not automatically grow faster than those containing more DNA. Therefore, we want to introduce a constraint that renders DNA synthesis the limiting factor of the growth rate. In this scenario, reducing chromosomal DNA would speed up replication leading to fastest growth of those cells with the smallest genome. Hence, we would not have to look for bacteria with reduced genomes, but let them find us!
MethodChemostatA chemostat is the instrument of choice for setting up a continuous culture. It is characterized by a continuous influx of medium, and an efflux of bacterial culture of the same volume. Since the volume of the continuous culture is kept constant, the growth rate of the population can be set by the dilution rate. Under these conditions, slowly growing (growth rate < dilution rate) cells will be washed out of the culture, while the fastest growing clone will take the lead and finally dominate the entire culture. A model of the selection mechanism can be found in Chemostat Selection. Here we also estimate the initial parameters for optimal selection.
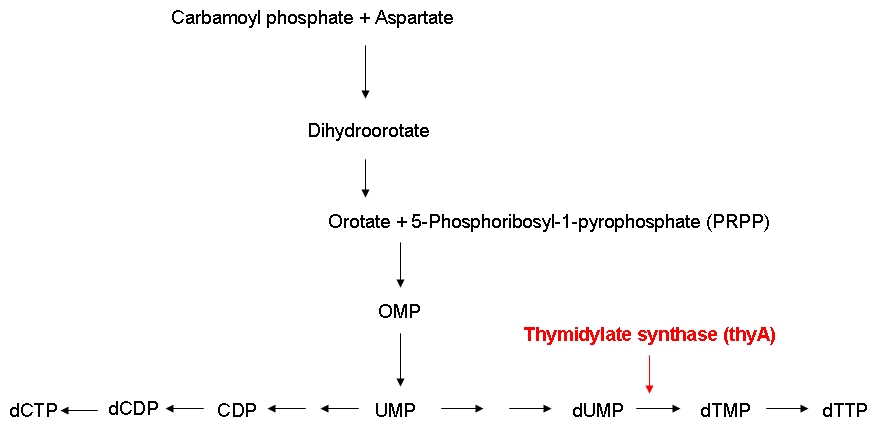
Growth constraintAs mentioned above, we want to introduce a constraint that renders DNA synthesis the limiting factor of the growth rate. DNA synthesis in bacterial cells is accomplished using the following biosynthetic pathway:
The thymidylate synthase catalyzes the methylation of dUMP to yield dTMP. Phosphorylation then converts dTMP into dTTP, one of the four building blocks of DNA. In contrast to the other three nucleotides DNA is made of, dTTP is exclusively used for DNA, but not for RNA synthesis. Therefore, impairing the activity of the thymidylate synthase should interfere quite specifically with DNA synthesis.
Lab resultsReferences(1) Escartin F., Skouloubris S., Liebl U., Myllykallio H. (2008): Flavin-dependent thymidylate synthase X limits chromosomal DNA replication. Proc Natl Acad Sci 22 105(29):9948-52. (2) Helmstetter C. (1996): Timing of synthetic activities in the cell cycle. Escherichia coli and Salmonella: Cellular and Molecular Biology. eds Neidhardt FC et al. (Am Soc Microbiol, Washington D. C.): 1627–1639. |
 "
"