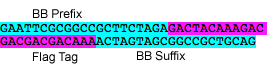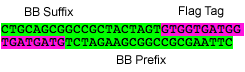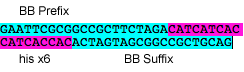Team:MIT/Notebook/sequences
From 2008.igem.org
| Home | The Team | The Project | Parts Submitted to the Registry | Modeling | Notebook |
|---|
Primers
Include sequence, usage, temperatures, concentration, start date.
P1: Forward Primer for Promoter
- foward sequence: CCG CTT CTA GAG TAA TAC GAC TCA CTA TAG GGA ATA CAA GCT ACT TGT TCT TTT TGC ATA CTA GAG ATT AAA GAG GAG AAA TAC TAG ATG CAG AAG AAA AAA TCC GC
- length = 107 bp
- GC content = 37.4 %
- melting temp = 68.9 ºC
- concentration:
- original:
- dilute:
P2: Reverse Primer for Peptide
- reverse sequence: GTT CTT CTC CTT TAC GCA TGC TGC CCC CGC CGC TGC CGC C
- length = 40 bp
- GC content = 67.5 %
- melting temp = 74.7 ºC
- concentration:
- original:
- dilute:
TEV Protease cut site
- Amino Acid Sequence : Glu-Asn-Leu-Tyr-Phe-Gln-Gly (Cuts between Gln-Gly)
- Base Pair Sequence: GAAAACCTGTACTTCCAGGGT
P3: Forward Primer for GFP
- forward sequence: GGC GGC AGC GGC GGG GGC AGC ATG CGT AAA GGA GAA GAA C
- length = 40 bp
- GC content = 67.5 %
- melting temp = 74.7 ºC
- concentration:
- original:
- dilute: 5μM?
P4: Reverse Primer for GFP
- sequence: CTG CAG CGG CCG CTA CTA GTA AGA GAA TAT AAA AAG CCA GAT TAT TAA TCC GGC TTT TTT ATT ATT TTT ATT AGT GGT GAT GGT GAT GAT GTT TGT ATA GTT CAT CCA TGC
- length = 111 bp
- GC content % = 36.0 %
- melting temp = 69.6 ºC
- concentration =
P5: Reverse Primer for GFP without Terminator
- sequence: CTG CAG CGG CCG CTA CTA GTA TTA TTA GTG GTG ATG GTG ATG ATG TTT GTA TAG TTC ATC CAT GC
- length = 65bp
- GC content = 44.6 %
- melting temp = 68.5 ºC
- concentration:
P6: Forward Primer for BioBrick FLAG tag
- sequence: GAATTCGCGGCCGCTTCTAGAGACTACAAAGACGACGACGACAAAACTAGTAGCGGCCGCTGCAG
- length = 65bp
- GC content = 55.4%
- melting temp = 72.1 ºC
- concentration =
P7: Reverse Primer for BioBrick FLAG tag
- sequence: CTGCAGCGGCCGCTACTAGTTTTGTCGTCGTCGTCTTTGTAGTCTCTAGAAGCGGCCGCGAATTC
- length = 65bp
- GC content = 55.4%
- melting temp = 72.1 ºC
- concentration =
P8: Forward Primer for BioBrick HA tag
- sequence: GAATTCGCGGCCGCTTCTAGATACCCGTACGACGTTCCGGACTACGCTACTAGTAGCGGCCGCTGCAG
- length = 68bp
- GC content = 60.3%
- melting temp = 73.4 ºC
- concentration =
P9: Reverse Primer for BioBrick HA tag
- sequence:CTGCAGCGGCCGCTACTAGTAGCGTAGTCCGGAACGTCGTACGGGTATCTAGAAGCGGCCGCGAATTC
- length = 68bp
- GC content = 60.3%
- melting temp = 73.4 ºC
- concentration =
P10: Forward Primer for BioBrick His tag
- sequence: GAATTCGCGGCCGCTTCTAGACATCATCACCATCACCACACTAGTAGCGGCCGCTGCAG
- length = 59bp
- GC content = 57.6%
- melting temp = 72.7 ºC
- concentration =
P11: Reverse Primer for BioBrick His tag
- sequence: CTGCAGCGGCCGCTACTAGTGTGGTGATGGTGATGATGTCTAGAAGCGGCCGCGAATTC
- length = 59bp
- GC content = 57.6%
- melting temp = 72.7 ºC
- concentration =
P12: Forward Primer for BioBrick TEV protease cut site
- sequence: GAATTCGCGGCCGCTTCTAGAGAAAACCTGTACTTCCAGGGTACTAGTAGCGGCCGCTGCAG
- length = 62bp
- GC content = 56.5%
- melting temp = 72.2 ºC
- concentration =
P13: Reverse Primer for BioBrick TEV protease cut site
- sequence: CTGCAGCGGCCGCTACTAGTACCCTGGAAGTACAGGTTTTCTCTAGAAGCGGCCGCGAATTC
- length = 62bp
- GC content = 56.5%
- melting temp = 72.2 ºC
- concentration =
P14: Forward Primer for BioBrick pRT B l.bulgaricus signal sequence
- sequence = GAATTCGCGGCCGCTTCTAGAATGCAGAAGAAAAAATCCGC
- length = 41bp
- GC content = 48.8%
- melting temp = 67.1 ºC
- concentration =
P15: Reverse Primer for BioBrick pRT B l.bulgaricus signal sequence
- sequence = CTGCAGCGGCCGCTACTAGTGGTAACCGGAGCCGTTTCTTG
- length = 41bp
- GC content = 61%
- melting temp = 71.3ºC
- concentration =
Plasmids
Include sequence, BioBrick #, functional features, digestion map.
PCR Products
Include sequence, template and primers, functional features.
Other
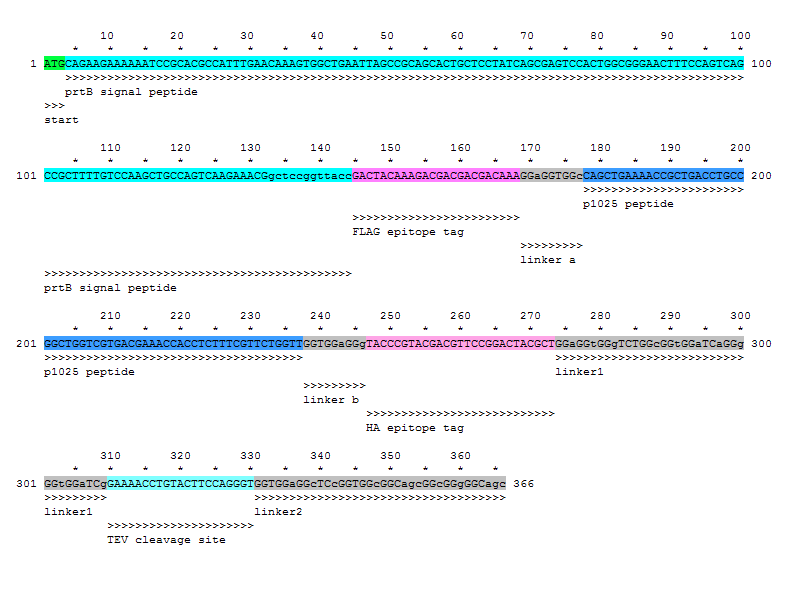
DNA sequence to synthesize - epitope/p1025/linker (6/17/08)
- The signal peptide sequence (141 bp) can be found at [http://www.ncbi.nlm.nih.gov/sites/entrez?dispmax=1&db=nucleotide&doptcmdl=graph&term=104773257 the NCBI genome database]
- 6/17, lengthened signal peptide so that it will cleave properly
- silent mutations in linkers to reduce repetition and to ensure specificity of primers (ex. GGC/GGA/GGG instead of GGT coding for Gly)
sequence: ATGCAGAAGAAAAAATCCGCACGCCATTTGAACAAAGTGGCTGAATTAGCCGCAGCACTGCTCC TATCAGCGAGTCCACTGGCGGGAACTTTCCAGTCAGCCGCTTTTGTCCAAGCTGCCAGTCAAGA AACGgctccggttaccGACTACAAAGACGACGACGACAAAGGaGGTGGcCAGCTGAAAACCGCT GACCTGCCGGCTGGTCGTGACGAAACCACCTCTTTCGTTCTGGTTGGTGGaGGgTACCCGTACG ACGTTCCGGACTACGCTGGaGGtGGgTCTGGcGGtGGaTCaGGgGGtGGaTCgGAAAACCTGTA CTTCCAGGGTGGTGGaGGcTCcGGTGGcGGCagcGGcGGgGGCagc
Whole Sequence
CCGCTTCTAGAGTAATACGACTCACTATAGGGAATACAAG CTACTTGTTCTTTTTGCATACTAGAGATTAAAGAGGAGAA ATACTAGATGCAGAAGAAAAAATCCGCACGCCATTTGAAC AAAGTGGCTGAATTAGCCGCAGCACTGCTCCTATCAGCGA GTCCACTGGCGGGAACTTTCCAGTCAGCCGCTTTTGTCCA AGCTGCCAGTCAAGAAACGGCTCCGGTTACCGACTACAAA GACGACGACGACAAAGGaGGTGGcCAGCTGAAAACCGCTG ACCTGCCGGCTGGTCGTGACGAAACCACCTCTTTCGTTCT GGTTGGTGGaGGgTACCCGTACGACGTTCCGGACTACGCT GGaGGtGGgTCTGGcGGtGGaTCaGGgGGtGGaTCgGAAA ACCTGTACTTCCAGGGTGGTGGaGGcTCcGGTGGcGGCag cGGcGGgGGCagcATGCGTAAAGGAGAAGAACTTTTCACT GGAGTTGTCCCAATTCTTGTTGAATTAGATGGTGATGTTA ATGGGCACAAATTTTCTGTCAGTGGAGAGGGTGAAGGTGA TGCAACATACGGAAAACTTACCCTTAAATTTATTTGCACT ACTGGAAAACTACCTGTTCCATGGCCAACACTTGTCACTA CTTTCGGTTATGGTGTTCAATGCTTTGCGAGATACCCAGA TCATATGAAACAGCATGACTTTTTCAAGAGTGCCATGCCC GAAGGTTATGTACAGGAAAGAACTATATTTTTCAAAGATG ACGGGAACTACAAGACACGTGCTGAAGTCAAGTTTGAAGG TGATACCCTTGTTAATAGAATCGAGTTAAAAGGTATTGAT TTTAAAGAAGATGGAAACATTCTTGGACACAAATTGGAAT ACAACTATAACTCACACAATGTATACATCATGGCAGACAA ACAAAAGAATGGAATCAAAGTTAACTTCAAAATTAGACAC AACATTGAAGATGGAAGCGTTCAACTAGCAGACCATTATC AACAAAATACTCCAATTGGCGATGGCCCTGTCCTTTTACC AGACAACCATTACCTGTCCACACAATCTGCCCTTTCGAAA GATCCCAACGAAAAGAGAGACCACATGGTCCTTCTTGAGT TTGTAACAGCTGCTGGGATTACACATGGCATGGATGAACT ATACAAACATCATCACCATCACCACTAATAAAAATAATAA AAAAGCCGGATTAATAATCTGGCTTTTTATATTCTCTTAC TAGTAGCGGCCGCTGCAGGATTA
| Home | The Team | The Project | Parts Submitted to the Registry | Modeling | Notebook |
|---|
 "
"