IGEM:Cambridge/2008/Notebook/Voltage/Gene Design
From 2008.igem.org
(Difference between revisions)
(New page: =KdpF-C Biobrick= ==Gene Selection== *Kdp is a well documented P-Type K+ ATPase found naturally in E.coli, used to actively pump ions into the cell. *It consists of a 6-gene operon: F,A,B...) |
|||
| (19 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | =KdpF-C Biobrick= | + | {{Cambridge/Notitle}} |
| + | <div class=bodytable> | ||
| + | {{Cambridge08}} | ||
| + | {{Cambridge08voltage}} | ||
| + | <html> | ||
| + | <table style="background:#444444; padding:15px;"> | ||
| + | <table align=left width= border="0" style="background:#444444; padding:5px;"> | ||
| + | <tr> | ||
| + | <td style="width:100%; height: 400; padding-left: 15px;"> | ||
| + | <b class="b1f"></b><b class="b2f"></b><b class="b3f"></b><b | ||
| + | class="b4f"></b> | ||
| + | <div class="contentf"> | ||
| + | <div style="height: 400; background: white; line-height:170% padding: | ||
| + | 5px;"> | ||
| + | <div style="color: white; font: 2px;">x</div> | ||
| + | </html> | ||
| + | |||
| + | The two constructs that we designed are detailed bellow. KDP is the potassium importer and GluR0 is the glutamate gated potassium channel. Details of their construction can be found here: | ||
| + | |||
| + | [[IGEM:Cambridge/2008/Notebook/Voltage/BioBrick_Manipulation| KDP]] | ||
| + | |||
| + | [[IGEM:Cambridge/2008/Notebook/Voltage/GluR0_Manipulation| GluR0]] | ||
| + | |||
| + | |||
| + | =KdpF-C Biobrick- [http://partsregistry.org/Part:BBa_K090003 BBa_K090003]== | ||
==Gene Selection== | ==Gene Selection== | ||
| Line 5: | Line 29: | ||
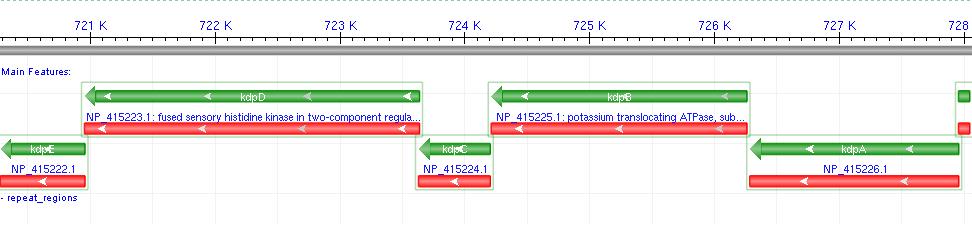
*It consists of a 6-gene operon: F,A,B,C,D,E Where F-C are the functional membrane protein subunits, and D-E comprises a bacterial 2-component regulatory system. | *It consists of a 6-gene operon: F,A,B,C,D,E Where F-C are the functional membrane protein subunits, and D-E comprises a bacterial 2-component regulatory system. | ||
| - | [[Image:kdpfull.JPG |800px| Native operon context from NCBI]] | + | [[Image:kdpfull.JPG |800px |Native operon context from NCBI]] |
*Literature shows that Kdp acts as a high-affinity transport system, and works most effectively at low external potassium concentrations, where a change in ion flux would be most likely to produce a measurable voltage difference. | *Literature shows that Kdp acts as a high-affinity transport system, and works most effectively at low external potassium concentrations, where a change in ion flux would be most likely to produce a measurable voltage difference. | ||
| Line 30: | Line 54: | ||
*. . . . . . . . . . . . . . . . . . . . . . . . | *. . . . . . . . . . . . . . . . . . . . . . . . | ||
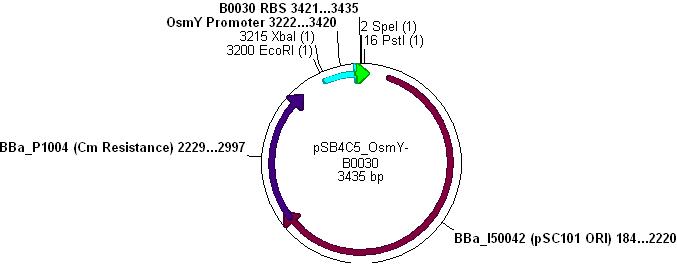
*Note: pSB4C5_Kdp biobrick plasmid has no promoter/RBS and so Kdp is not expressed in transformants. | *Note: pSB4C5_Kdp biobrick plasmid has no promoter/RBS and so Kdp is not expressed in transformants. | ||
| - | |||
| - | |||
=Promoter+RBS Biobrick= | =Promoter+RBS Biobrick= | ||
| Line 79: | Line 101: | ||
*Note: This promoter-RBS construct did not cause unwanted transcript problems because there are many double terminators scattered throughout the pSB4C5 backbone. | *Note: This promoter-RBS construct did not cause unwanted transcript problems because there are many double terminators scattered throughout the pSB4C5 backbone. | ||
| - | =Combination of Kdp, OsmY and B003x= | + | =Combination of Kdp, OsmY and B003x to make [http://partsregistry.org/Part:BBa_K090004 BBa_K090004]= |
*This will create a functional biobrick plasmid in which Kdp is overexpressed only in stationary phase of growing cells. | *This will create a functional biobrick plasmid in which Kdp is overexpressed only in stationary phase of growing cells. | ||
:*Cut pSB4C5-Kdp with XbaI and PstI | :*Cut pSB4C5-Kdp with XbaI and PstI | ||
| Line 85: | Line 107: | ||
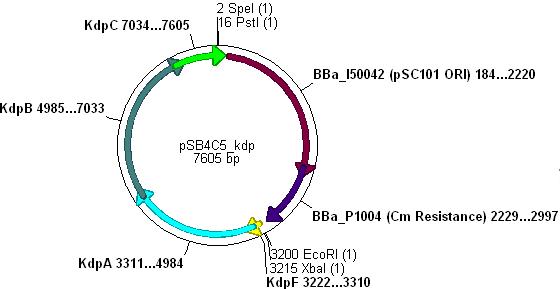
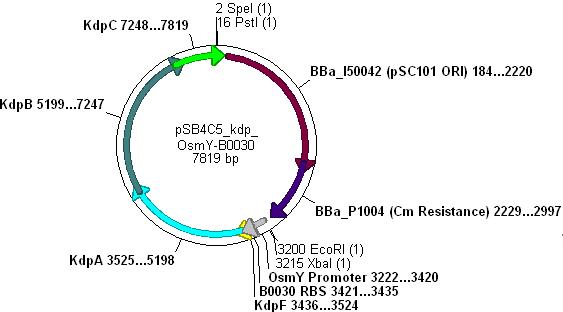
:*Ligation will form the following functional plasmid: | :*Ligation will form the following functional plasmid: | ||
| - | [[Image:psb4c5_osmy_b0030_kdp.JPG|400px]] | + | [[Image:psb4c5_osmy_b0030_kdp.JPG|400px]] |
| - | =GluR0 Biobrick= | + | =GluR0 Biobrick- [http://partsregistry.org/Part:BBa_K090002 BBa_K090002]= |
==Gene Selection== | ==Gene Selection== | ||
| Line 105: | Line 127: | ||
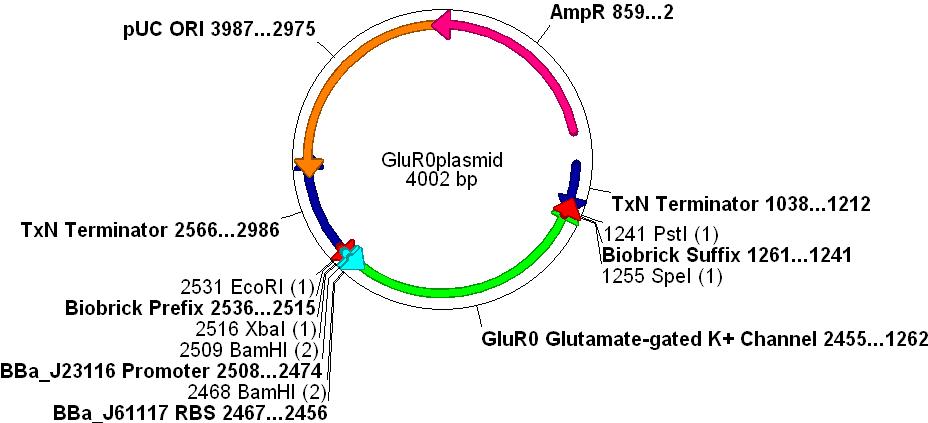
[[Image:Glur0plasmid.JPG |800px]] | [[Image:Glur0plasmid.JPG |800px]] | ||
| + | |||
| + | <html> | ||
| + | <div style="color: white; font: 2px;">x</div> | ||
| + | </div></div> | ||
| + | <b class="b4f"></b><b class="b3f"></b><b class="b2f"></b><b | ||
| + | class="b1f"></b> | ||
| + | </td></tr></table></table> | ||
| + | </html> | ||
Latest revision as of 22:30, 29 October 2008
|
x
The two constructs that we designed are detailed bellow. KDP is the potassium importer and GluR0 is the glutamate gated potassium channel. Details of their construction can be found here:
KdpF-C Biobrick- BBa_K090003=Gene Selection
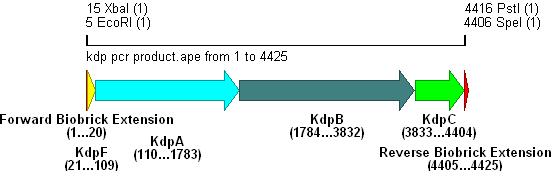
Amplification from E.coli MG1655
-Forward: ATAT GAATTC ATAT TCTAGA TGAGTGCAGGCGTGATAACCGGCGTATT
EcoRI XbaI
-Reverse: CTCT CTGCAG CTCT ACTAGT TTATTCATCAAGTTTATCCAGCGCCAGAT
PstI SpeI
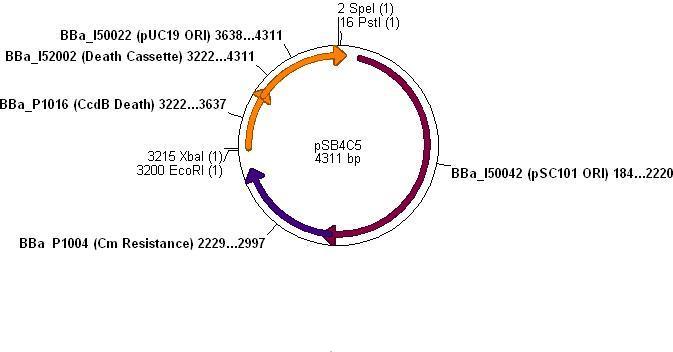
Integration into Vector
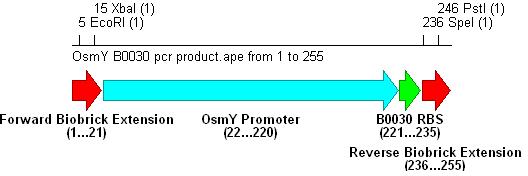
Promoter+RBS BiobrickPromoter and RBS SelectionPromoter
Ribosome Binding Site
Amplification from E.coli MG1655
-Forward: CTAT GAATTC ATAT TCTAGA GCTGGCACAGGAACGTTATCC (All OsmY-RBS constructs)
EcoRI XbaI
-B0030 Reverse: CGCG CTGCAG CTCT ACTAGT (TTTCTCCTCTTTAAT)TTGTTAAATATAGA
PstI SpeI B0030
-B0031 Reverse: CTCT CTGCAG CTCT ACTAGT (GGTTTCCTGTGTGA)TTGTTAAATATAGAT
PstI SpeI B0031
-B0032 Reverse: CTCT CTGCAG CTCT ACTAGT (CTTTCCTGTGTGA)TTGTTAAATATAGATCA
PstI SpeI B0032
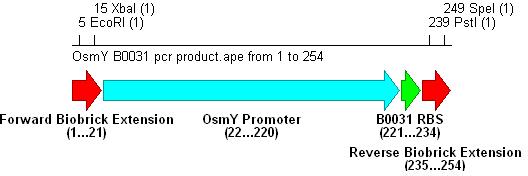 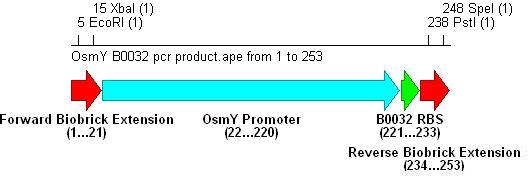
Integration into Vector
Combination of Kdp, OsmY and B003x to make BBa_K090004
GluR0 Biobrick- BBa_K090002Gene Selection
DNA Synthesis
x
|
 "
"