Team:Bologna/Wetlab
From 2008.igem.org
| HOME | PROJECT | TEAM | SOFTWARE | MODELING | WET LAB | LAB-BOOK | SUBMITTED PARTS | BIOSAFETY AND PROTOCOLS |
|---|
Contents |
LAB Experiment
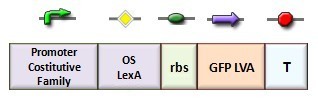
In our genetic bi-stable, the amount of molecules produced to switch the system from the two possible steady state is ruled by LacI and TetR protein. The production of LacI molecules by UV induction can be tested replacing the LacI gene by GFP, so it is possible to have a relation of the strength of LacI synthesis measuring the value of its fluorescence. BBa_K079049 and BBa_K079050 are two new construct submitted to the registry by Bologna’s Igem Team 2008. These UV test circuits can be presented schematically by the following sequence:
- Promoter Constitutive Family: BBa_J23118 (1429 strength ) or BBa_23100 (2547 strength);
- LexA Operator Site Binding (K079040 ) ;
- Rbs with Green fluorescence Protein LVA and Terminator (BBa_I763020)
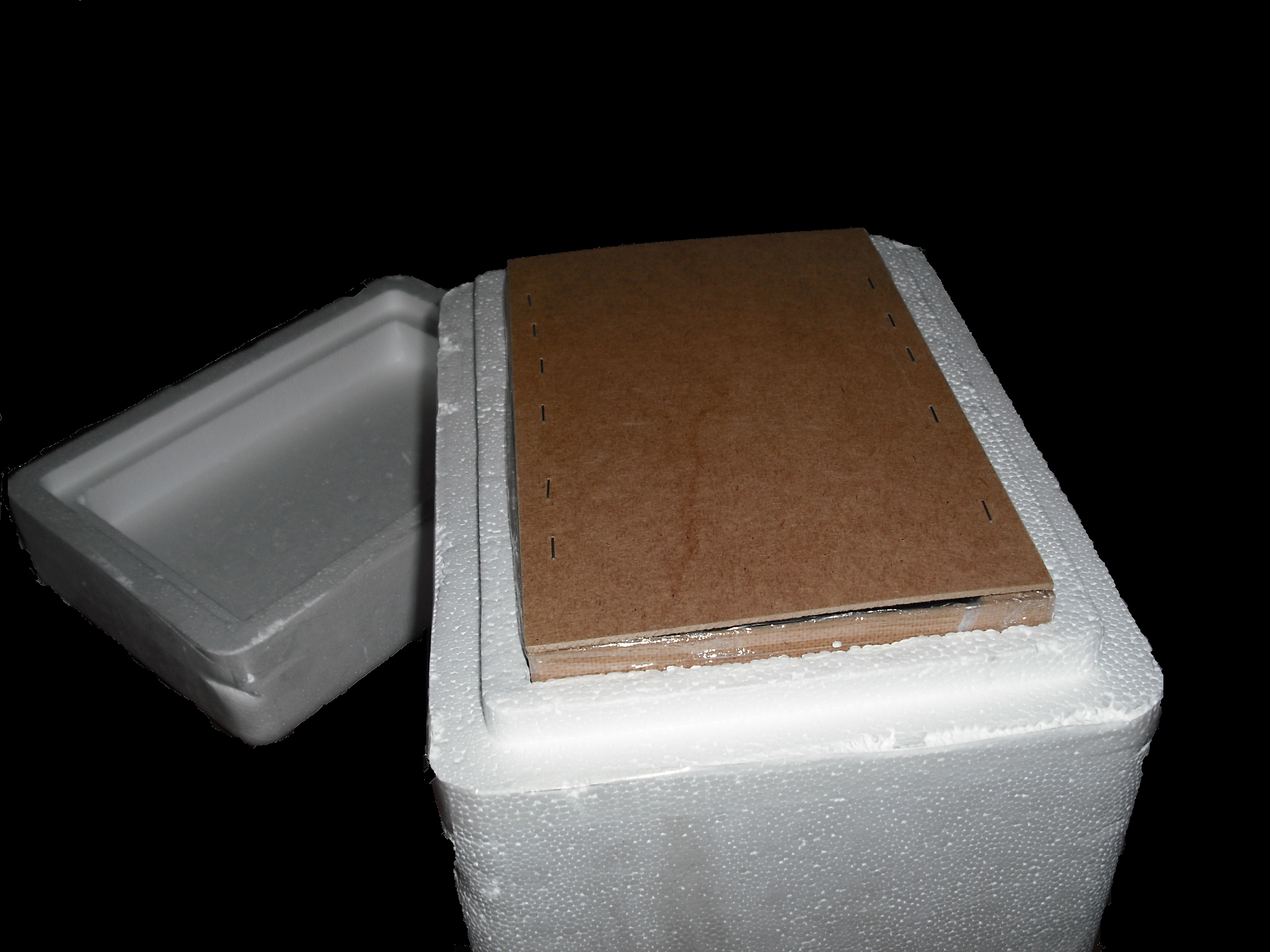
In order to generate the induction by Uv, a box with UV lamp was realized.
Homemade UV Illuminator
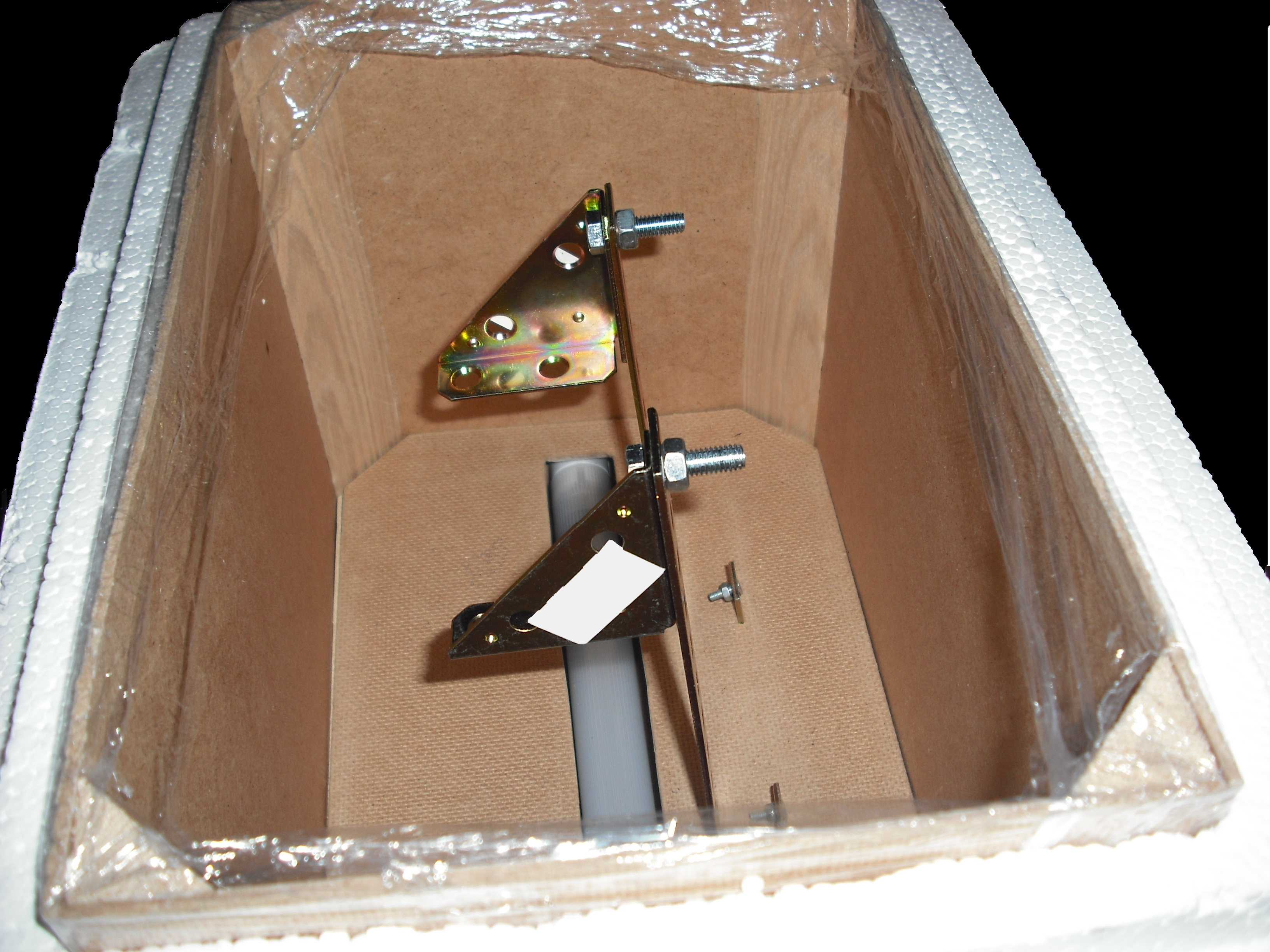
The UV source that we use is a GW6 Sylvania, a lamp that emitt UVC at 253,7nm near the absorption's peak of DNA with an optical output power of 1,6W; to use it safely we built a box of mdf(medium density-fibreboard) that surrounds the lamp and prevent the UVc leakeage, moreover we use a diaphragm that allows passage only to the light absorbing the remainder. We insert in that box two pierced brackets, that permits to choose the distance between the lamp and the sample and then in accord with the Lambert-Beer law's the exposure time. Finally we embedded this structure in a polistyrene box for its handling and greater safety (Figure 2-3).
Gel matrix
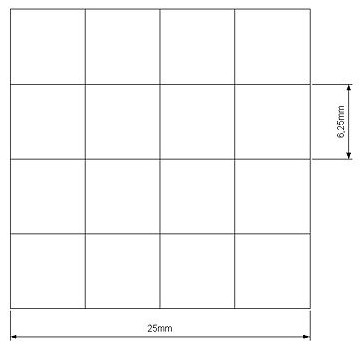
Our project use UV light for its space selectivity, that gives the possibility to irradiate a target zone without interfering with the other. To do that we build a mold to make a matrix of agarose gel;this give us a square of 25mm side's with 16 cells inside where we can locate bacteria.
Once do that is easy with an optical mask, like those used in photolithography for electronics
circuits, stimulate only the selected bacteria.To realize this device we use palsticard because it is
very easy to shape and clean.
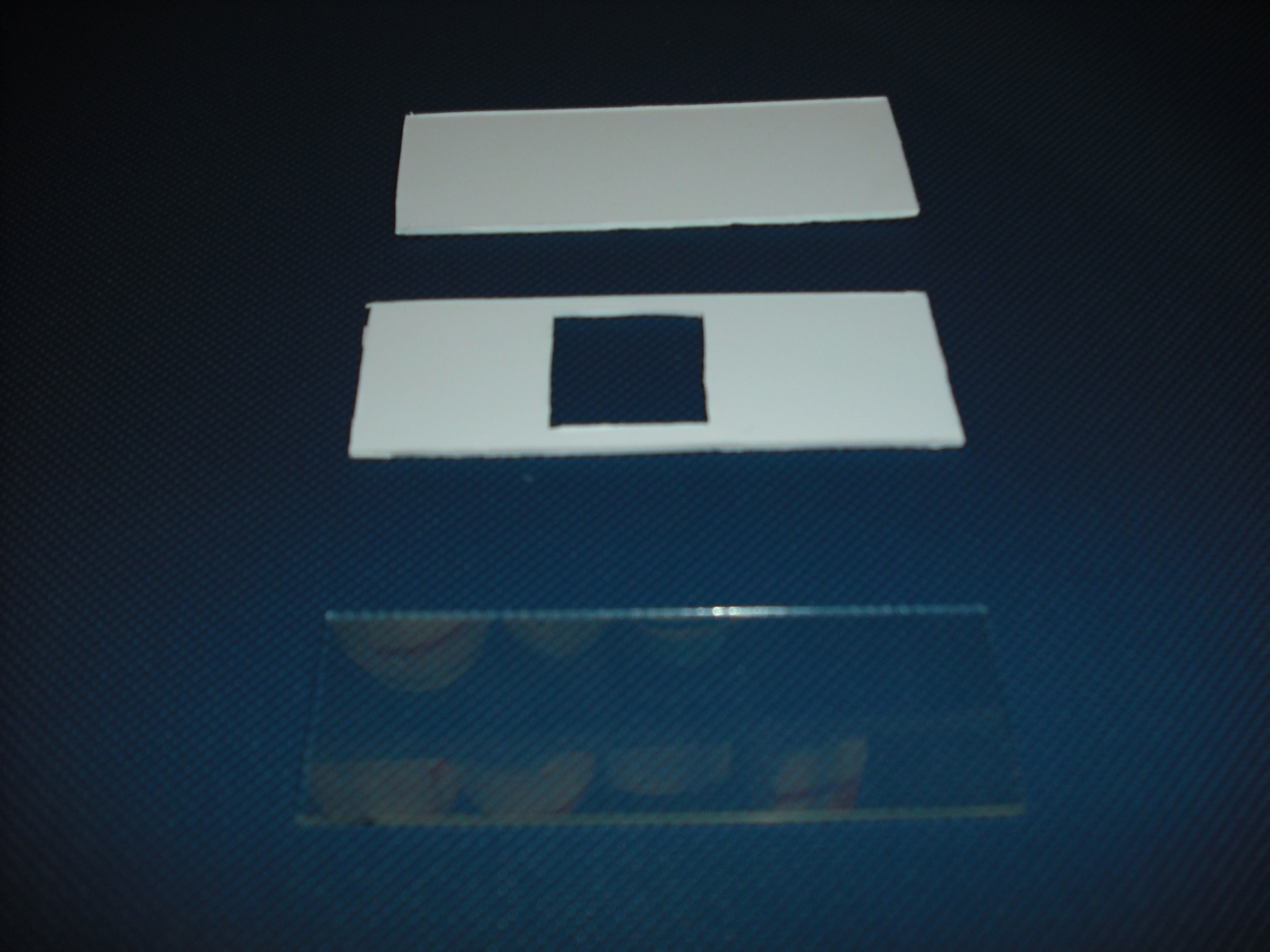
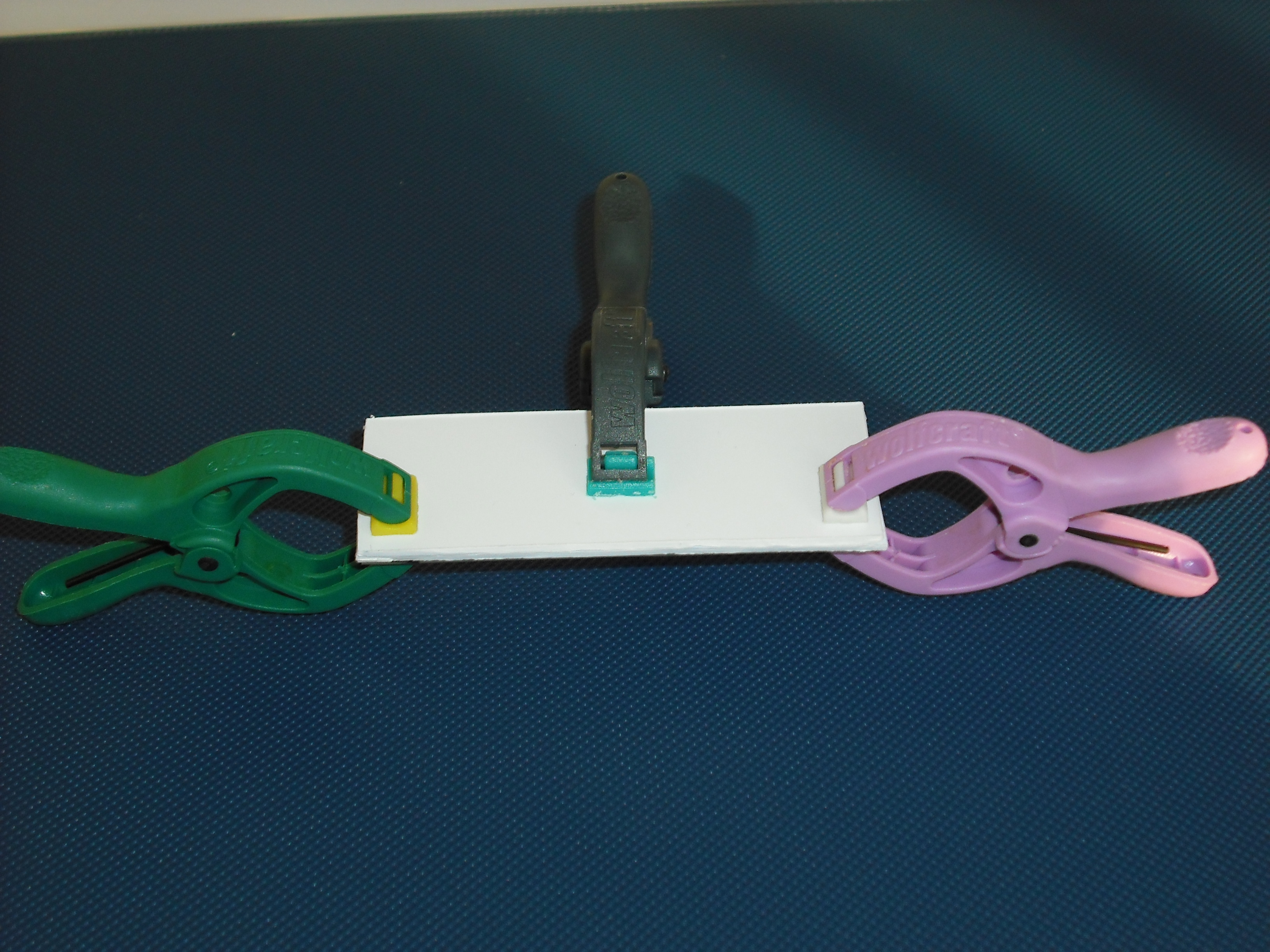
The mold is made of two parts that will form a sandwich with the slide: one will create the shape
of the gel matrix the other is a cover that that will give the picture into gel .
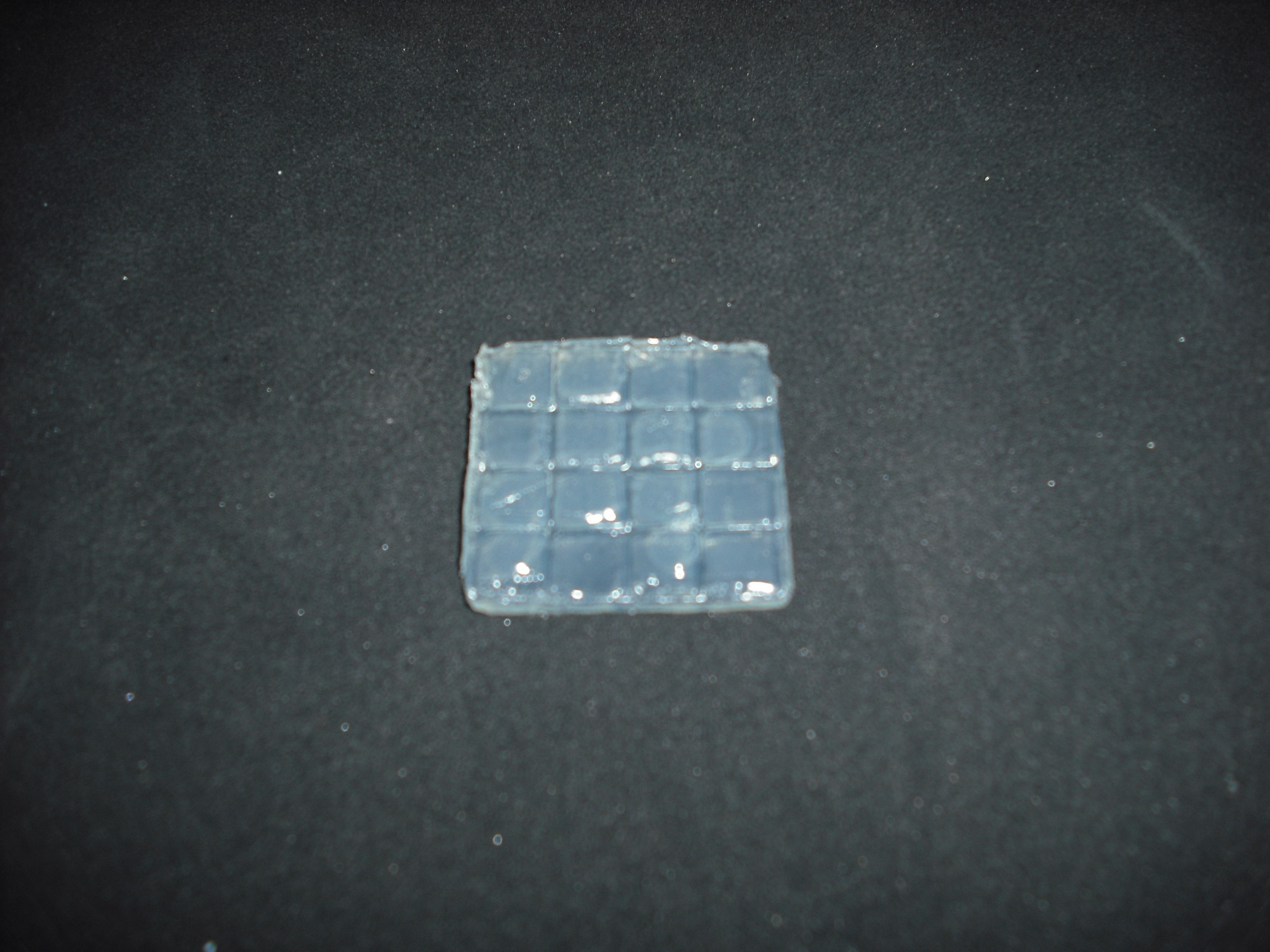
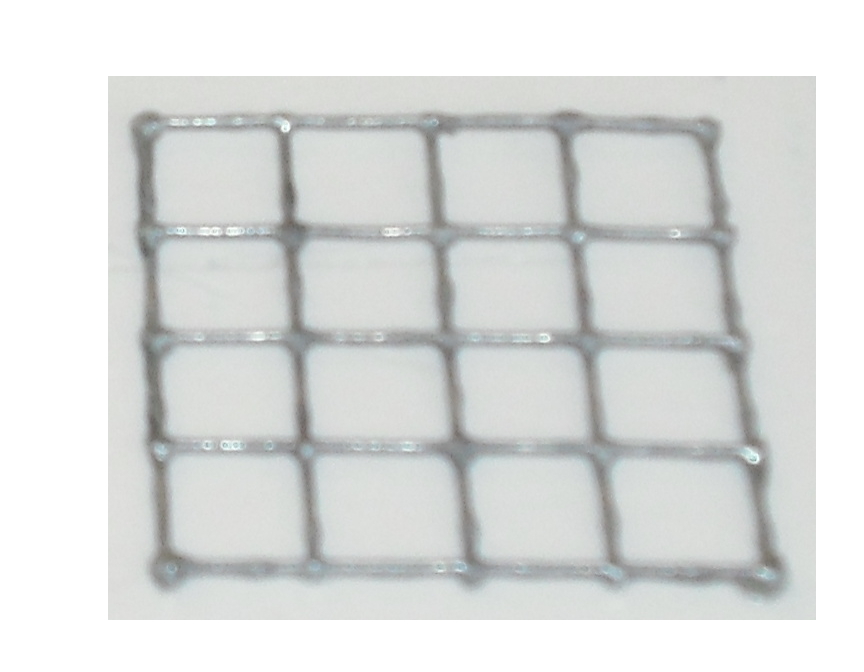
The matrix is obtained through the pressure of the cover part over the mold part
and that is the result:
Operator sites are dna sequences very small in length (15 to 20 bp) and a restriction digestion for the religation in standard plasmids is not possible with the existing purification kits. In fact, only Dna sequences down to at least 40 bp can be purified with the standard kits available. So, we decided to set a new protocol up for the isolation and cloning of single operator sites into standard BioBricks plasmids.
We started from the analysis of an existing promoter library into the Registry (link). As it can be seen in figure, these plasmids were meant for the construction of promoter basic parts and their derivatives. They can be used two ways: 1. Insertion of a promoter element between XbaI and SpeI sites results in a RFP reporter while retaining the ability to do BioBrick assembly. Part J61002 is the tet promoter variant of the plasmid. 2. Insertion of a protein generating device or RNA gene (cutting the part with XbaI/PstI, inserting into SpeI/PstI of J61002) results in a standard pSB1A2 plasmid containing an r0040.yourpart composite.
FOTO!!!!
Thus, we decided to isolate the RFP protein from one of the promoter family member with a SpeI- PstI enzymatic digestion. Then, this part could be assemble with an operator sequence digested SpeI- PstI, too. This could allow the isolation of the Operator- RFP sequence from the commercial plasmid and the subsequent religation into a standard plasmid. Then, a final extraction of the RFP gene with a SpeI- PstI digestion would leave the operator site inside the standard plasmid.
 "
"