Team:Caltech/Project/Oxidative Burst
From 2008.igem.org
| Line 42: | Line 42: | ||
====Characterization of LuxR Inducer==== | ====Characterization of LuxR Inducer==== | ||
| + | [[Image:LuxR TetR response curves.png|thumb|center|400px|Figure X - Characterization of LuxR inducer and TetR inverter by flow cytometry. The figure shows the fluorescence of GFP in the constructs luxR-GFP and luxR-tetR-GFP. The left axis corresponds to the LuxR data and the right axis corresponds to the TetR data. At 100 nM AHL, cells with the luxR-GFP construct grew very poorly.]] | ||
| + | |||
| + | The TetR inverter was characterized by using a luxR-tetR-GFP construct. In this context, GFP provides a relative gauge of expression levels of KatG in the final construct. Figure 4 shows the tetracycline inverter behaved as expected. At high concentrations of AHL, the TetR inverter shows near full repression, as compared to the lower concentrations of AHL. Similar to the LuxR inducer, the TetR inverter showed a 50 fold difference in expression between the uninduced (low AHL) and induced (high AHL) states. However the absolute maximum level of expression from the TetR inverter was much lower than that from the LuxR inducer (compare the scale of the left TetR axis to the scale of the left LuxR axis). The low expression levels can be explained by the leakiness of the LuxR promoter, resulting in a basal level of repression from the TetR inverter. | ||
| + | The low expression level is advantageous for the engineered E. coli because catalase is a very efficient scavenger of hydrogen peroxide. The low expression level ensures there is enough KatG to buffer any leaky pyruvate oxidase expression, yet still clear quickly from the cytoplasm. Additionally, in the uninduced state there should be extremely low levels of KatG. The LAA degradation tag should reduce KatG levels even further. | ||
| + | |||
====Characterization of TetR Inverter==== | ====Characterization of TetR Inverter==== | ||
====Production of Hydrogen Peroxide==== | ====Production of Hydrogen Peroxide==== | ||
Revision as of 22:46, 16 October 2008
|
People
|
Pathogen Defense by Oxidative Burst
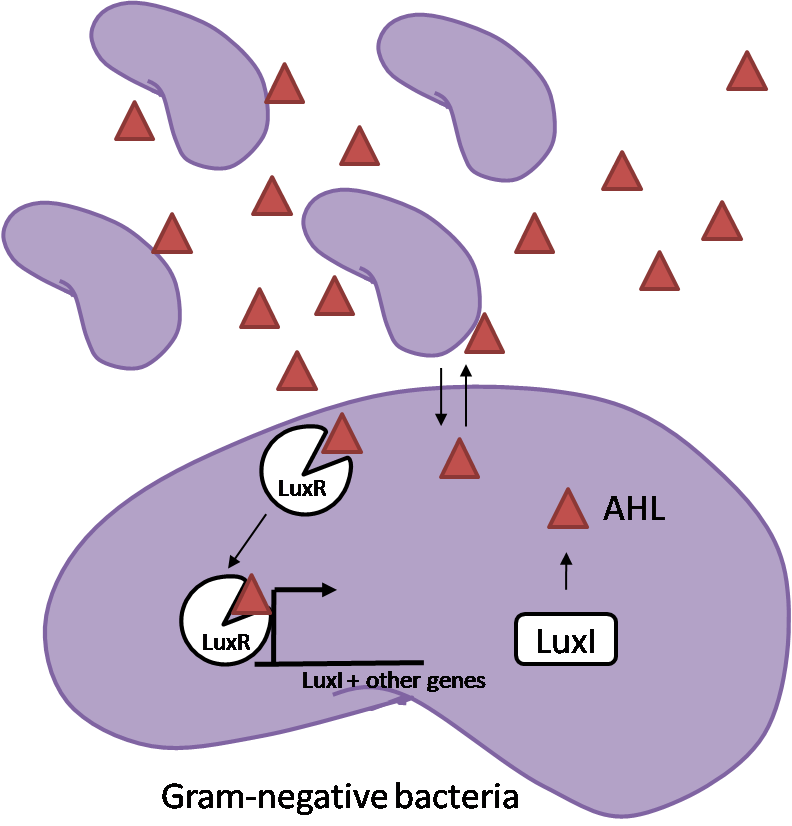
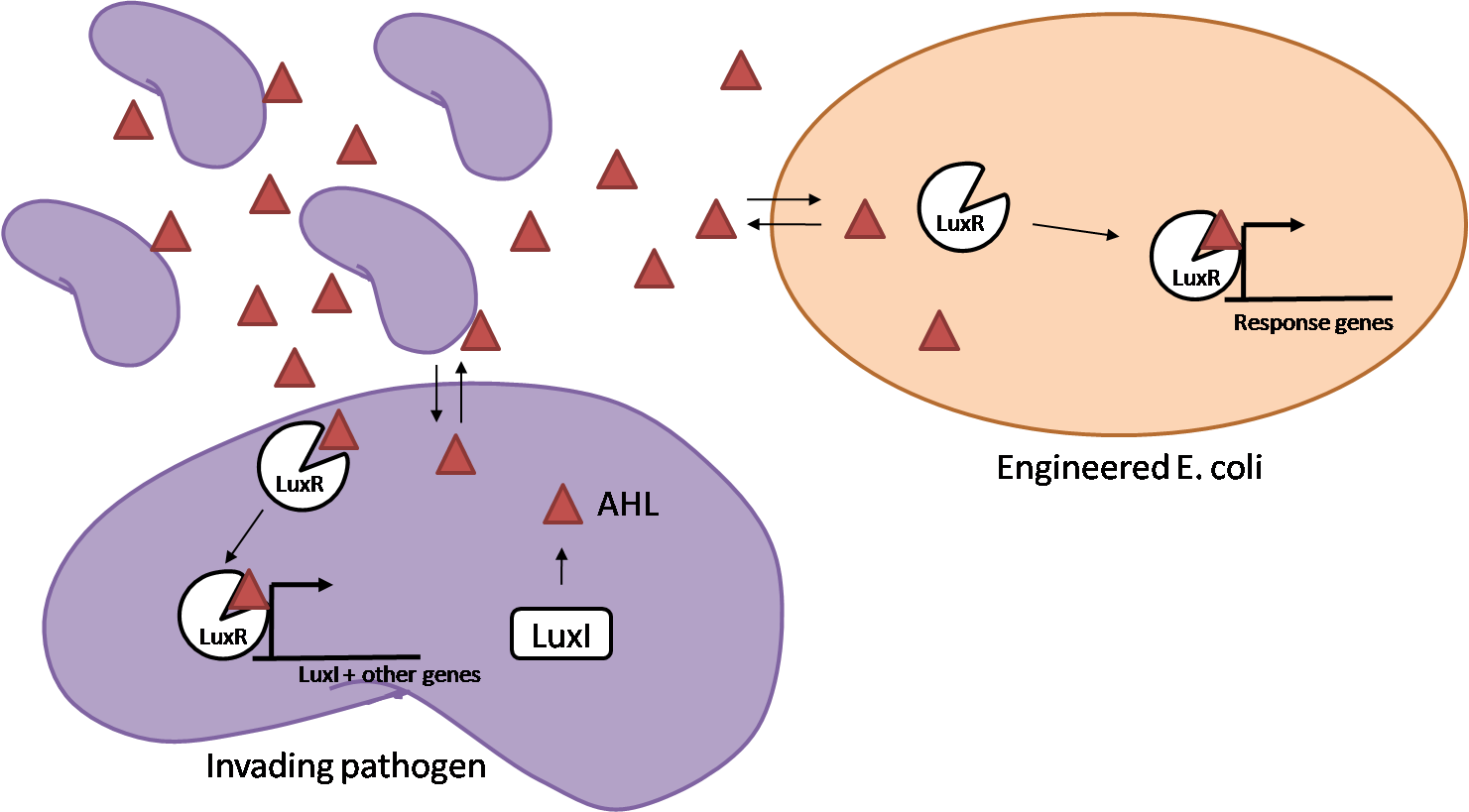
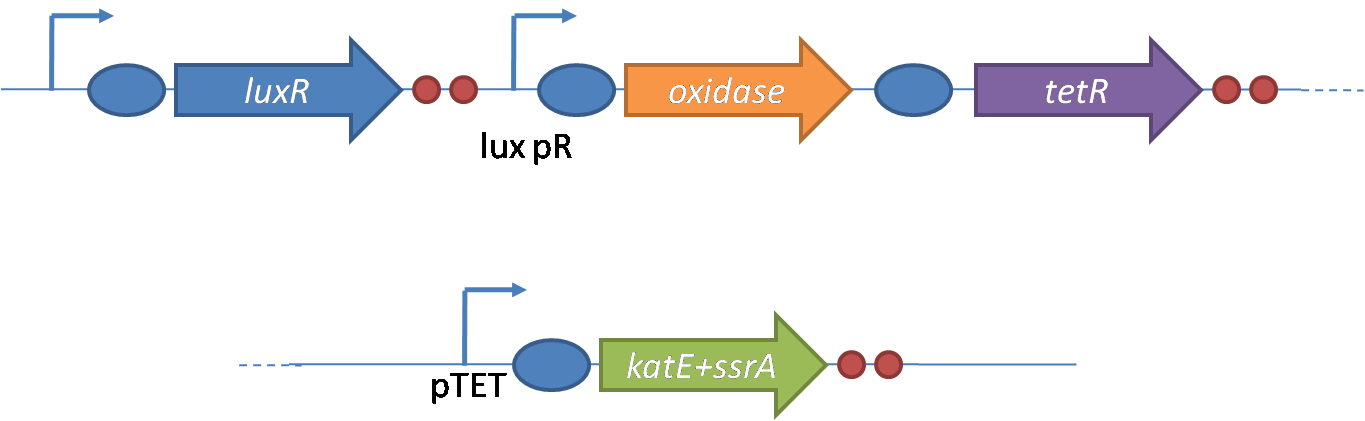
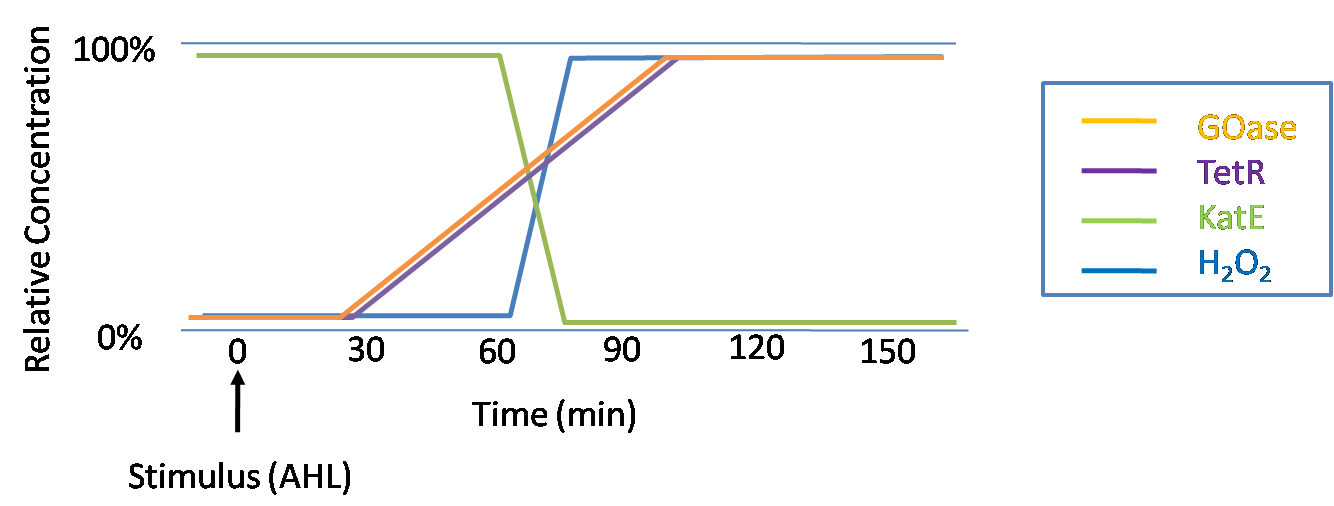
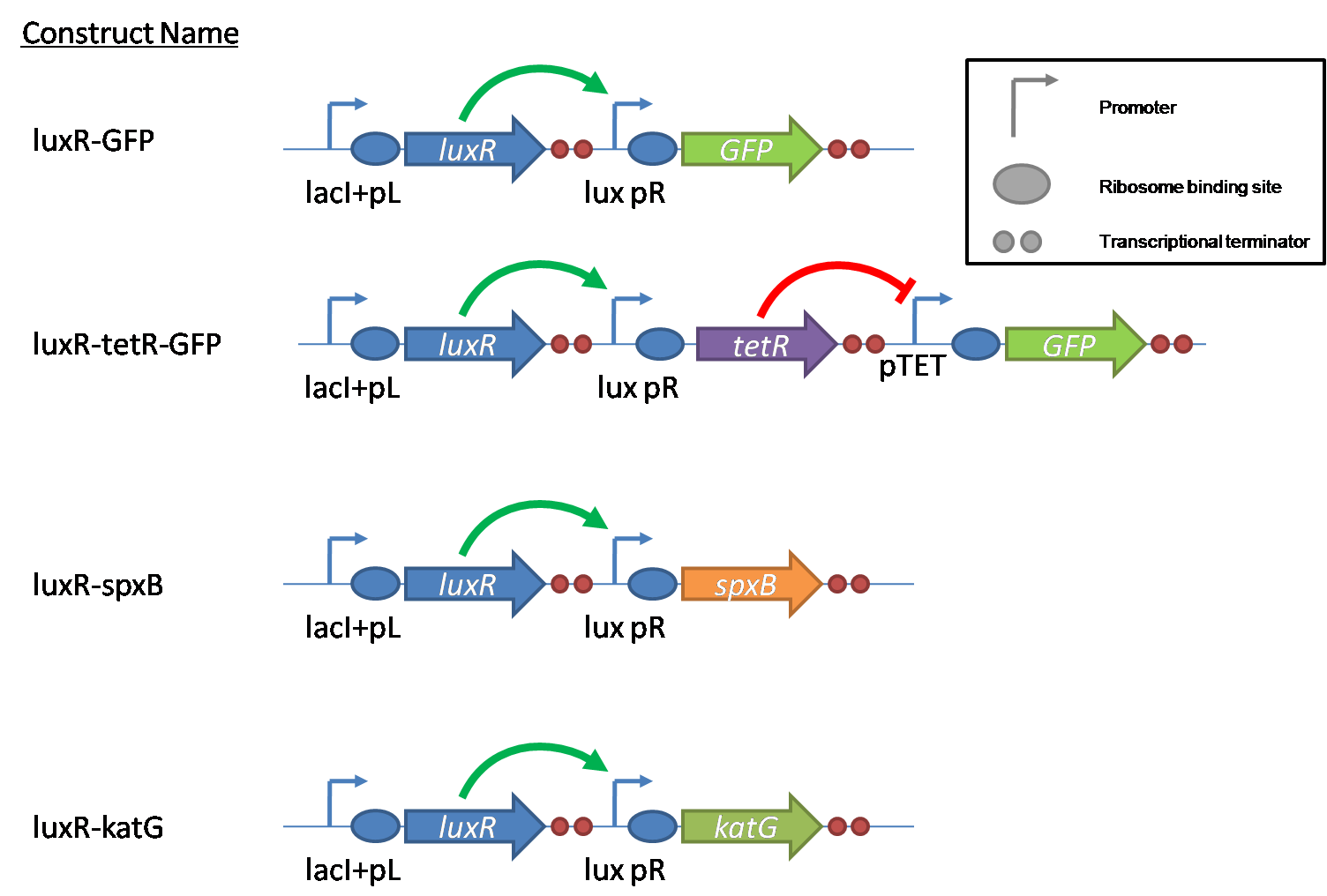
IntroductionIn order to help guard against infections of the gut, we wish to engineer a strain of E. coli capable of killing bacterial pathogens. White blood cells called neutrophils are already very efficient at killing bacteria. Neutrophils kill bacteria by bombardment it with a variety of reactive oxygen species including superoxide, hydrogen peroxide and hydrochlorous acid. The reactive oxygen species kill the bacteria by shredding biological molecules by way of their potent oxidizing properties. However neutrophils do not to migrate to the large intestinal lumen where pathogens can reside. Because bacteria are well adapted to live in the gut, this project’s goal is to engineer a strain of E. coli to detect and kill invading bacterial pathogens by means of a sudden burst of hydrogen peroxide. DetectionBacteria are able to communicate between individuals of the same species by way of quorum sensing. Small molecules serve as the signal between individual cells. Gram negative bacteria use acylhomoserine lactones (AHL), which can freely diffuse across the cell membrane. The quorum sensing machinery relies in two enzymes, LuxI, an AHL producer, and LuxR, an AHL-dependent transcriptional activator. Figure 1 illustrates how the quorum sensing system works. In isolation, each bacterium constitutively produces a small amount of AHL, which quickly diffuses into the surroundings. If other bacteria of the same species are also nearby, the AHL will diffuse across their membrane where it will bind to LuxR. LuxR activates transcription of several genes, including luxI. A positive feedback loop is created, in which more AHL induces more LuxI, which in turn produces more AHL. Each species of gram negative bacteria produces a unique AHL, requiring unique LuxI and LuxR proteins, and so avoids crosstalk between species. A group of bacteria can thus toggle between an “off” state and an “on” state by using quorum sensing. Our engineered strain will not participate directly in quorum sensing, but instead will eavesdrop on the conversation. It will be engineered to constitutively express a LuxR able to detect a species AHL. By taking advantage of the specificity of quorum sensing, our engineered strain will be able to be tuned to specifically respond to a variety of bacterial pathogens. Once the AHL is bound, LuxR will activate a set of genes which will lead to the overproduction of hydrogen peroxide, killing the invading cell. ResponseAfter sensing the presence of an invading pathogen, we want to engineer our E. coli to produce lethal amounts of hydrogen peroxide relatively quickly. We are not concerned with having the engineered E. coli survive either, as it is reasonable to assume that there are “unactivated” cells far away from the pathogen that could sustain the population. Even if all of the oxidative burst cells were wiped out, the population could be regenerated. In combination with the population variation project, the "master cell line" would differentiate into the oxidative burst state, thus reseeding the population. After LuxR binds AHL, it will activate transcription of pyruvate oxidase, which uses pyruvate to produce hydrogen peroxide in the following reaction: Pyruvate + phosphate + O2 --> H2O2 + CO2 + acetyl phosphate Streptococcus pneumoniae naturally uses pyruvate oxidase to kill off competing bacteria when infecting the lungs. Having already known its activity and antibacterial properties in vivo, pyruvate oxidase was an attract oxidase with which to work. Even though our activated engineered cells will eventually die from their oxidative burst, we want them to survive long enough to produce large amounts of the oxidase so they can produce large amounts of hydrogen peroxide. If the cells were left to produce hydrogen peroxide without any protection, they would produce just enough to be cytotoxic and then fissle, killing only themselves and little else. To avoid this problem, an E. coli catalase will be constitutively expressed, and then turned off shortly after pyruvate oxidase expression is triggered. We’re accomplishing this by putting katG (on of two E. coli catalase genes) behind the tetR sensitive promoter (tetR P) and having tetR co-transciptionally expressed with the oxidase. The time it takes for tetR to accumulate in the cell provides the delay in repressing katG expression. To ensure katG is rapidly cleared from the cells, it has a C-terminal ssrA degradation tag, which should reduce the protein’s half life to the order of minutes. In this way, the cell can be temporarily protected from hydrogen peroxide, but large amounts can accumulate before the substrate is exhausted.The final strain will have deletions of both catalases, ensuring no complementation. System DesignThe entire system will be engineered onto a high copy plasmid in E. coli as seen in Figure 3. Promoters are shown as bent, thin arrows, genes as thick arrows, ribosome binding sites as blue ovals, and a double transcriptional terminator as two red circles. Initially, the system will function independently, with luxR under a constitutive promoter. Later, luxR will be put under the control of a recombinase system that will only turn on the pathway randomly in a subset of cells. This will be one of three fates the master strain will be able to differenciate into. For more information, visit the population variation page. An ideal system would behave as outlined in Figure 4. It shows the relative abundances of the proteins and molecules used in the system. It is only meant to show if a particular species is relatively high or low, not its absolute concentration. ResultsConstructsCharacterization of LuxR Inducer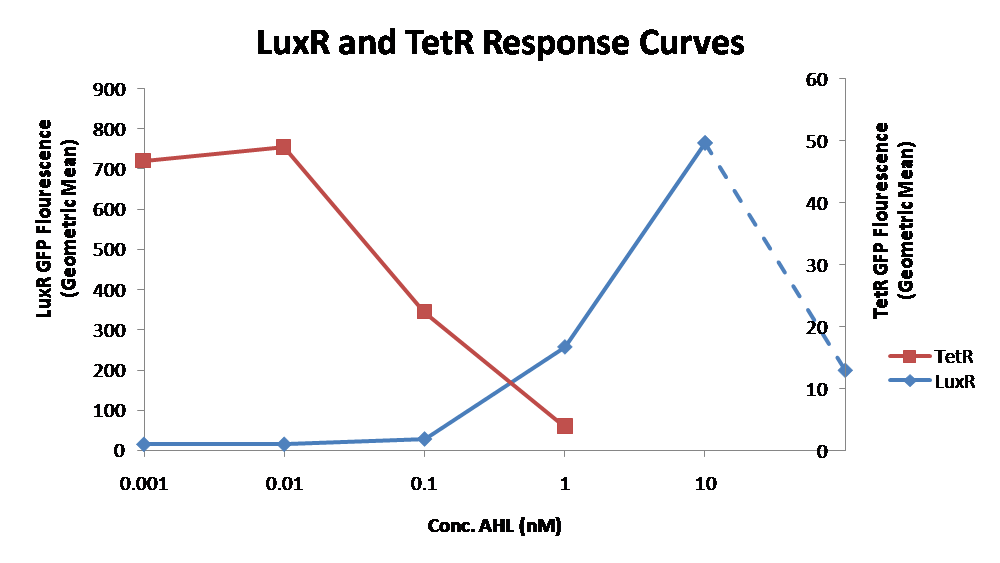 Figure X - Characterization of LuxR inducer and TetR inverter by flow cytometry. The figure shows the fluorescence of GFP in the constructs luxR-GFP and luxR-tetR-GFP. The left axis corresponds to the LuxR data and the right axis corresponds to the TetR data. At 100 nM AHL, cells with the luxR-GFP construct grew very poorly. The TetR inverter was characterized by using a luxR-tetR-GFP construct. In this context, GFP provides a relative gauge of expression levels of KatG in the final construct. Figure 4 shows the tetracycline inverter behaved as expected. At high concentrations of AHL, the TetR inverter shows near full repression, as compared to the lower concentrations of AHL. Similar to the LuxR inducer, the TetR inverter showed a 50 fold difference in expression between the uninduced (low AHL) and induced (high AHL) states. However the absolute maximum level of expression from the TetR inverter was much lower than that from the LuxR inducer (compare the scale of the left TetR axis to the scale of the left LuxR axis). The low expression levels can be explained by the leakiness of the LuxR promoter, resulting in a basal level of repression from the TetR inverter. The low expression level is advantageous for the engineered E. coli because catalase is a very efficient scavenger of hydrogen peroxide. The low expression level ensures there is enough KatG to buffer any leaky pyruvate oxidase expression, yet still clear quickly from the cytoplasm. Additionally, in the uninduced state there should be extremely low levels of KatG. The LAA degradation tag should reduce KatG levels even further. Characterization of TetR InverterProduction of Hydrogen PeroxideFunctionality of KatGCoculture AssayCurrent Progress
References |
 "
"