Team:Edinburgh/Results
From 2008.igem.org
(Difference between revisions)
| Line 1: | Line 1: | ||
<div id="header">{{Template:Team:Edinburgh/Templates/Header}}</div> | <div id="header">{{Template:Team:Edinburgh/Templates/Header}}</div> | ||
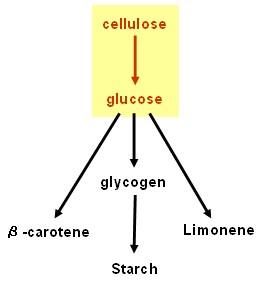
| - | == | + | ==Cellulolysis device== |
| - | + | [[Image:Edinburgh Map1 Cellulolysis.jpg|150px]] | |
| - | + | ||
* '''''cenA''''' (endoglucanase) and '''''cex''''' (exoglucanase): Biobricks made, but encountered problems when adding ribosome binding site. | * '''''cenA''''' (endoglucanase) and '''''cex''''' (exoglucanase): Biobricks made, but encountered problems when adding ribosome binding site. | ||
* '''''bglX''''' (beta glucosidase): Biobrick made. | * '''''bglX''''' (beta glucosidase): Biobrick made. | ||
| Line 9: | Line 8: | ||
** [[Team:Edinbrugh/Results/PcstA-xylE|''P<sub>cstA</sub>'' promoter assay with ''xylE'' as repoter]] | ** [[Team:Edinbrugh/Results/PcstA-xylE|''P<sub>cstA</sub>'' promoter assay with ''xylE'' as repoter]] | ||
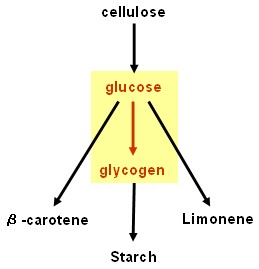
| - | + | == Glycogenesis device == | |
| + | [[Image:Edinburgh Map2 Glycogen.jpg|150px]] | ||
* '''''glgC''''' and '''''glgC16''''' (ADP-glucose pyrophosphorylase, responsible for rate limiting step in glycogenesis): Biobricks made and tested - successful. | * '''''glgC''''' and '''''glgC16''''' (ADP-glucose pyrophosphorylase, responsible for rate limiting step in glycogenesis): Biobricks made and tested - successful. | ||
** [[Team:Edinburgh/Results/Glycogen1|Glycogen Assay 1 (Qualitative)]] | ** [[Team:Edinburgh/Results/Glycogen1|Glycogen Assay 1 (Qualitative)]] | ||
| Line 15: | Line 15: | ||
** [[Team:Edinburgh/Results/Glycogen3|Glycogen Assay 3 (Quantitative)]] | ** [[Team:Edinburgh/Results/Glycogen3|Glycogen Assay 3 (Quantitative)]] | ||
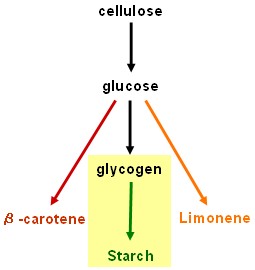
| - | + | == Starch synthesis device == | |
| + | [[Image:Edinburgh Map3 Starch.jpg|150px]] | ||
* '''''su1''''' and '''''iso2''''' (isoamylases): Work in progress - to be continued in an undergraduate honours project. | * '''''su1''''' and '''''iso2''''' (isoamylases): Work in progress - to be continued in an undergraduate honours project. | ||
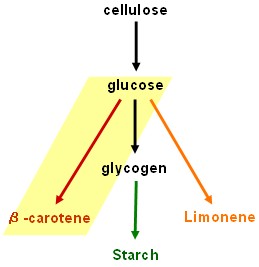
| - | + | ==Lycopene generator == | |
| + | [[Image:Edinburgh Map4 BCarotene.jpg|150px]] | ||
* '''''dxs+crtE+crtB+crtI+crtY''''': Work in progress - encountered problem at final step. | * '''''dxs+crtE+crtB+crtI+crtY''''': Work in progress - encountered problem at final step. | ||
| - | + | ==Limonene synthase device== | |
| + | [[Image:Edinburgh Map5 Limonene.jpg|150px]] | ||
* '''''dxs+LIMS+appY''''': Biobrick made - still being tested (last test failed to detect limonene synthesis). | * '''''dxs+LIMS+appY''''': Biobrick made - still being tested (last test failed to detect limonene synthesis). | ||
Revision as of 20:52, 27 October 2008
Contents |
Cellulolysis device
- cenA (endoglucanase) and cex (exoglucanase): Biobricks made, but encountered problems when adding ribosome binding site.
- bglX (beta glucosidase): Biobrick made.
- PcstA (promoter): Biobrick made and tested - successful.
Glycogenesis device
- glgC and glgC16 (ADP-glucose pyrophosphorylase, responsible for rate limiting step in glycogenesis): Biobricks made and tested - successful.
Starch synthesis device
- su1 and iso2 (isoamylases): Work in progress - to be continued in an undergraduate honours project.
Lycopene generator
- dxs+crtE+crtB+crtI+crtY: Work in progress - encountered problem at final step.
Limonene synthase device
- dxs+LIMS+appY: Biobrick made - still being tested (last test failed to detect limonene synthesis).
 "
"