Team:NYMU-Taipei/Project/Urea
From 2008.igem.org
| Home | Project Overview: | pH Sensor | Attachment | Time Regulation | Waste Removal | Experiments and Parts | About Us |
Contents |
Why should urea be removed
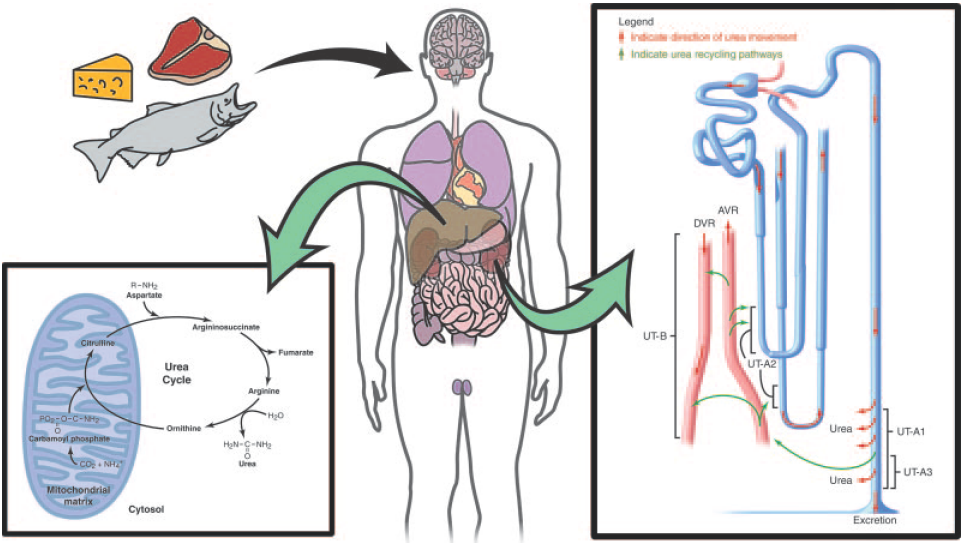
Physiologically, urea is synthesized from ammonia and carbon dioxide. The elimination of urea means getting rid of nitrogenous waste from the body. In healthy individuals, blood plasma are filtrated by kidney and the waste removed by urination; in patients with renal failure, such waste are accumulated in blood. These patients have to get renal dialysis treatment to remove the waste, or they would suffer from hepatic failure and encephalopathy, which is worse. For these reasons, urea should be removed by a healthy kidney or by medical treatment.
However, the renal dialysis treatment knocks down the quality of life because of the time spent being treated and equipment portage (or patient transportation). If urea can be removed by the enterobacteria (bacteria in intestines, such as E. coli or our BacToKidney), a capsule or an yogurt-like drug may help reduce the frequency of get renal dialyzed and increase the quality of life. If so, the next half of the patients' lifespan may be more fun.
How do we try to remove urea
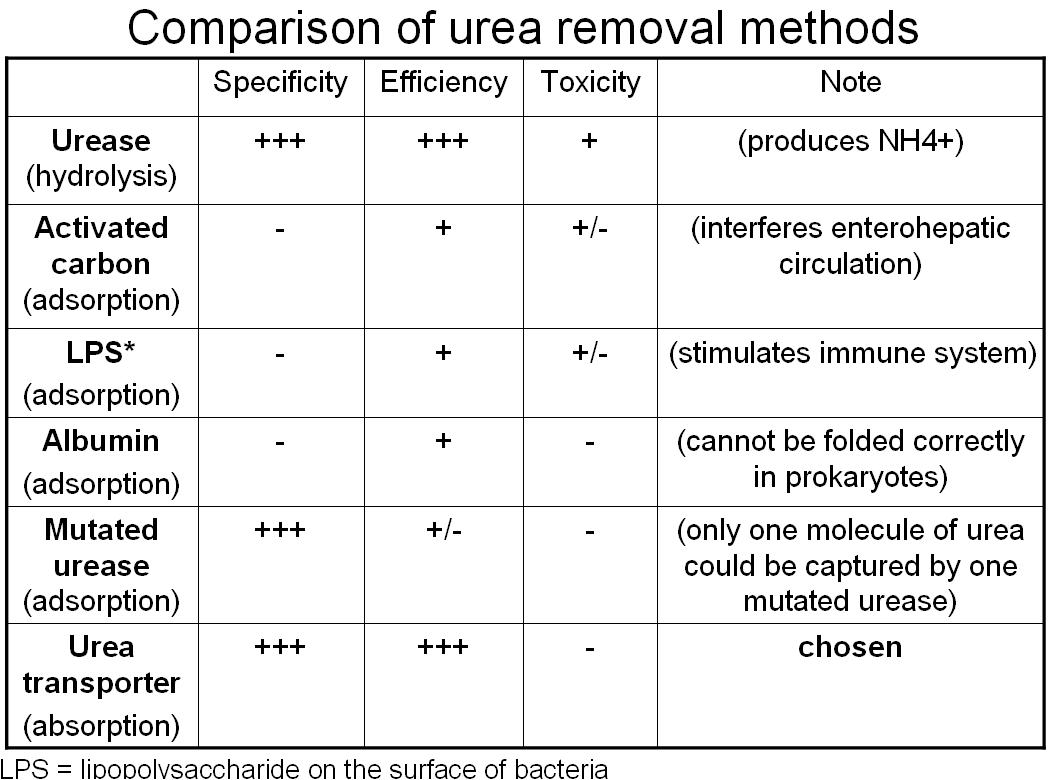
Absorption Method--Urea Transporter (chosen method)
- Urea may be absorbed by the micro dialysis machine (or be uptaken by the engineered bacteria), and this micro machine may be eliminated with stool passage.
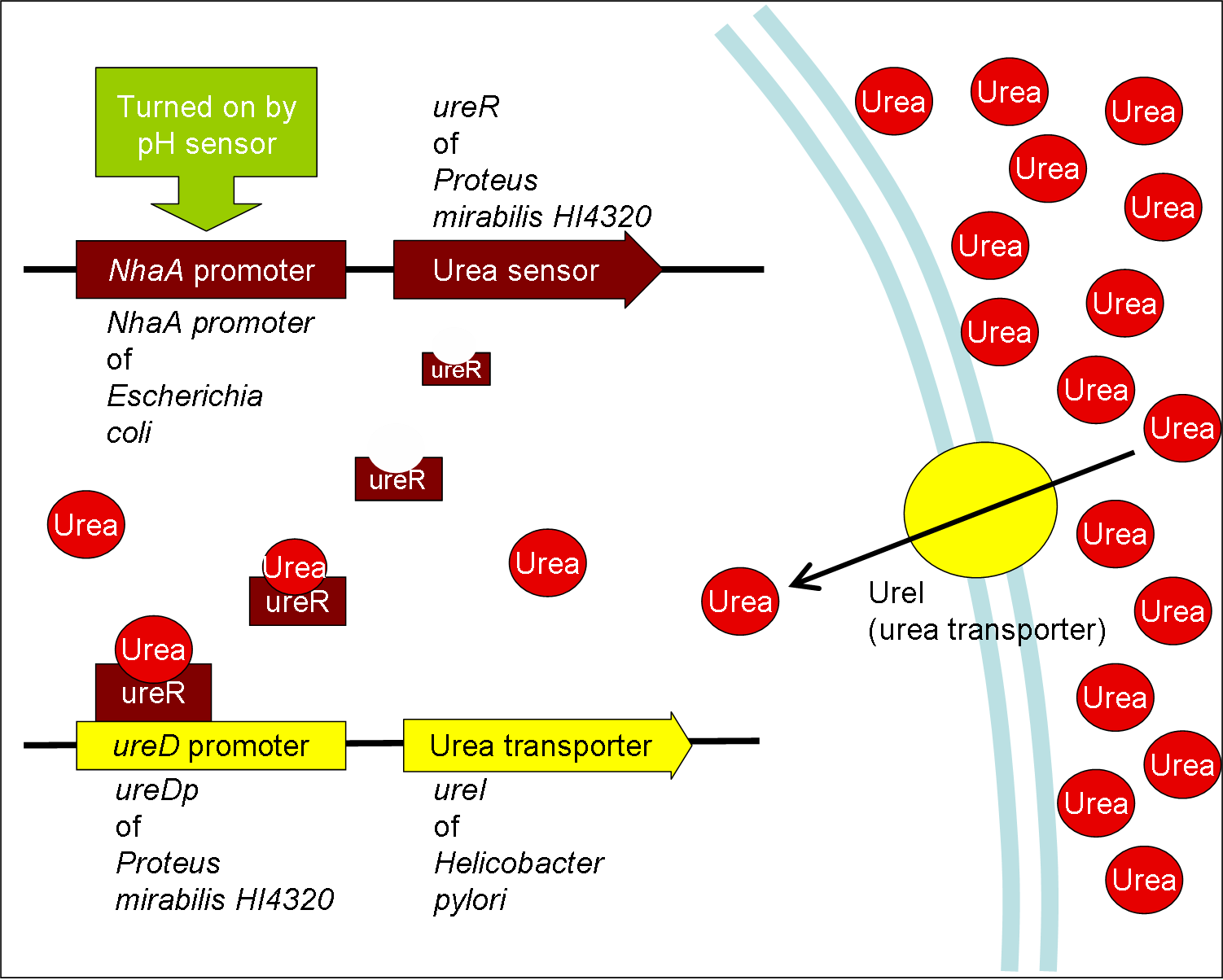
- Urea transporter would be expressed only if both pH sensor and urea sensor are turned on:
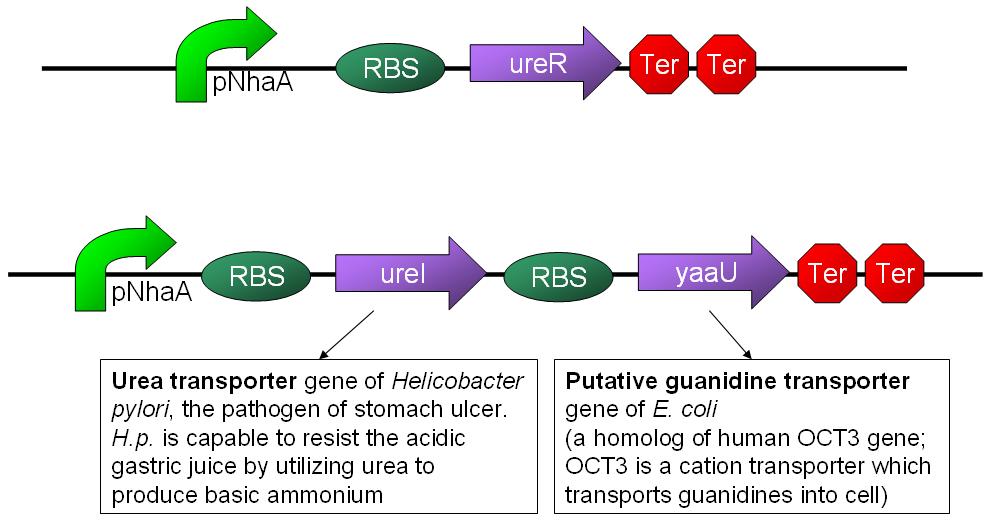
- When the device is turned on by pH sensor, ureR, the pH sensor is expressed. The ureR protein may bind with urea molecule to form a complex. This ureR-urea complex may activate a downstream promoter, ureDp, and then urea transporter will be expressed.
- In this circuit design, urea sensor turns not only urea transporter on but also guanidine transporter on at the same time. Guanidine transporter is integrated into this circuit because the concentratoin of guanidine in blood may be elevated after an rising of urea consentration.
Adsorption Method
- Urea remover 1: use albumin to adsorb urea on the surface of bacteria
- Urea remover 2: use urease to bind urea on the surface of bacteria
- Urea remover 3: "Lactobaccilize" the bacteria; produce more lactic acid and adsorb urea on the bacterial capsule in low pH level
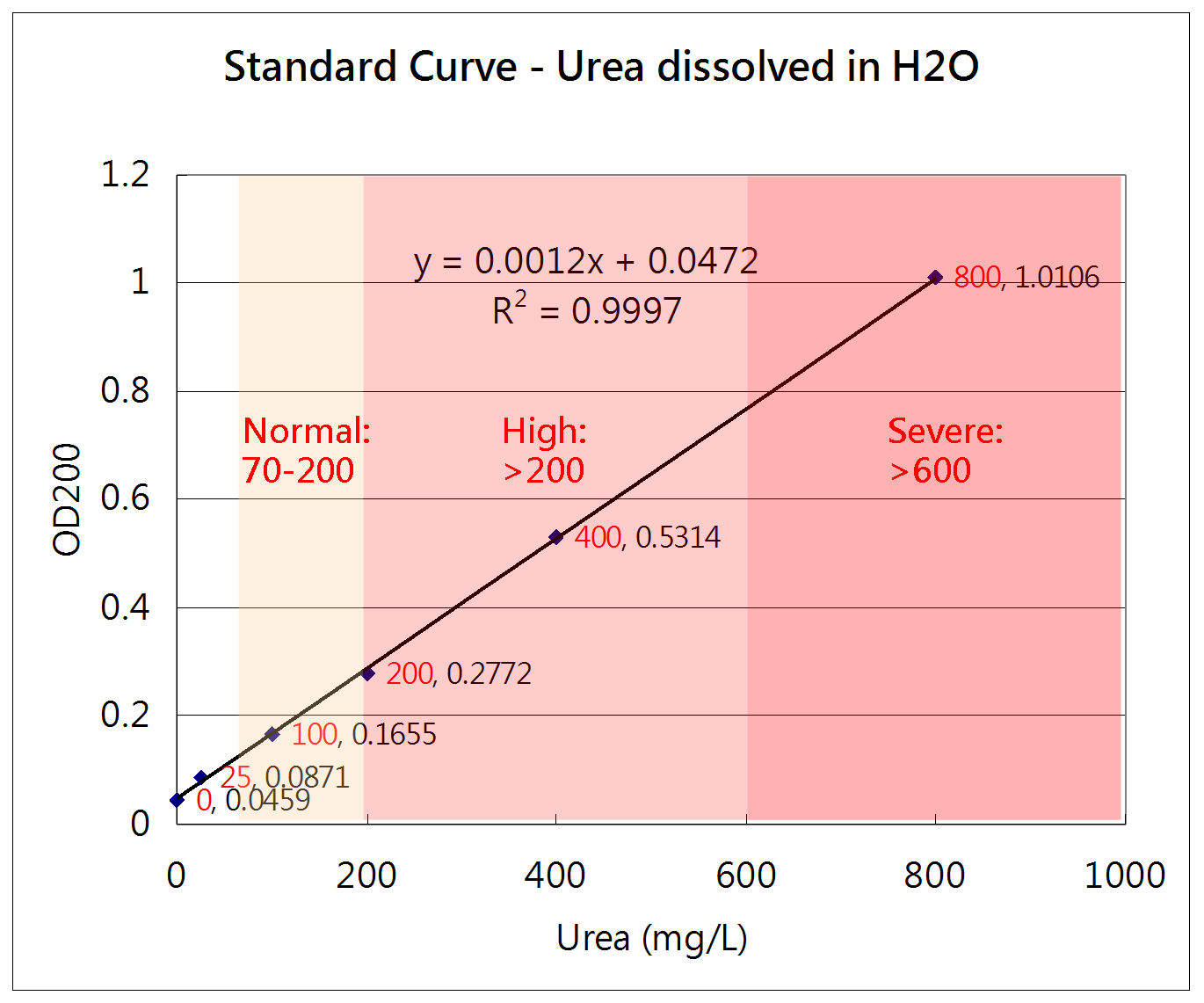
How to measure the concentration of urea
Urease method
Urea + H2O --(urease)--> 2NH4+ + CO2
NH4+ + alpha-KG + NADH --(GLDH)--> Glutamate + NAD+ + H2O
The change of NADH concentration reflects the urea concentration in the solution.
Absorbance of NADH = A340nm - A405nm
- use an in vitro diagnostic kit
- Formosa Biomedical Technology Blood Urea Nitrogen diagnostic kit
- or undertaken by clinical laboratories
- UCL (Union Clinical Laboratory) A test for urea nitrogen costs NTD$60 (as private patient without health insurance)
Diacetyl monoxime method:
Diacetyl Monoxime + Urea --> Diazine (yellow)
- WHO SOP
- The Automated Thiosemicarbazide-Diacetyl Monoxime Method for Plasma Urea, Clinical Chemistry 15: 393-396, 1969
 "
"