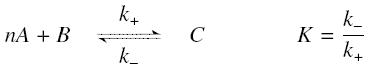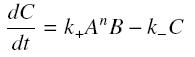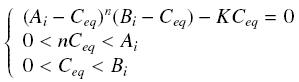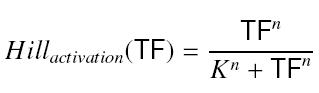Complexations Caracterisations
The first hypothesis is that a complexation reaction is fully determined by the following :
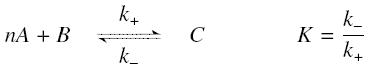
and that the rates k+ et k- stay constants under all conditions.
We then have the following kinetic : 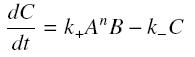
so, at steady-state : File:Sshill.jpf.
Then, since we guess that the only data we will first have are the quantites of A and B introduced, the only equations we will deal with is the following, entirely determining the concentration Ceq at steady-state :
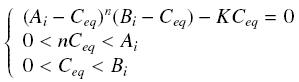
- Inducer TF on Py :
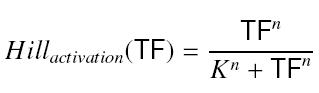
formation rate of y = β*Hillactivation(TF)
- Inhibitor TF on Py :

formation rate of y = β*Hillrepression(TF)
With following chemical interpretations :
- β = "quantity of DNA times the transduction rate"
- K = "activation constant : dissociation constant of the fixation of x on py"
- n = "stoechiometric coef. of x in the fixation of x on py"
Finding Parameters
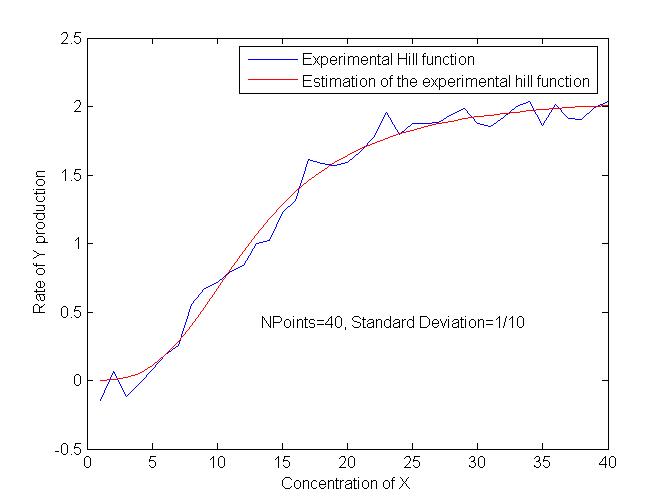
This program aims at fill this data bank, from experimental data obtained by our wet lab.
After some trials, we have obtained interesting results in terms of precision. We are now trying to test the robustness of the estimations in order to quantify the influence of biological variance in the results as well as the influence of the number of points available.
The corresponding code can be found there : Estimation of the parameters
|  "
"