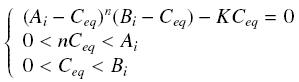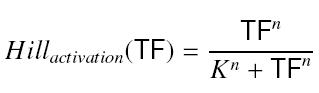Team:Paris/Modeling/Programs
From 2008.igem.org
(→Complexations Caracterisations) |
(→Complexations Caracterisations) |
||
| Line 15: | Line 15: | ||
and at steady-state : <center> [[Image:sshill.jpf]] </center> | and at steady-state : <center> [[Image:sshill.jpf]] </center> | ||
| - | Then, since we guess that the only data we will first have are the quantites of ''A'' and ''B'' introduced, the only equations we will deal with is the following, entirely determining the concentration ''C<sub>eq</sub>'' at steady-state : | + | Then, since we guess that the only data we will first have are the quantites of ''A'' and ''B'' introduced, the only equations we will deal with is the following, entirely determining the concentration ''C<sub>eq</sub>'' at steady-state (at least if we take the smallest real root of the equation, it is useless to demonstrate the unicity, or even the existence, of such a solution) : |
<center> [[Image:Eqnss.jpg]] </center> | <center> [[Image:Eqnss.jpg]] </center> | ||
| + | |||
Revision as of 15:48, 15 September 2008
|
 "
"