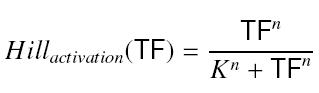Team:Paris/Modeling/Programs
From 2008.igem.org
(Difference between revisions)
(→Finding Parameters) |
(→Finding Parameters) |
||
| Line 3: | Line 3: | ||
|-valign="top" | |-valign="top" | ||
|style="background:#ffffff"| | |style="background:#ffffff"| | ||
| - | |||
| - | |||
We are looking for constants involved in Hill functions : <br> | We are looking for constants involved in Hill functions : <br> | ||
| Line 21: | Line 19: | ||
* K = "activation constant : dissociation constant of the fixation of x on py" <br> | * K = "activation constant : dissociation constant of the fixation of x on py" <br> | ||
* n = "stoechiometric coef. of x in the fixation of x on py" <br> | * n = "stoechiometric coef. of x in the fixation of x on py" <br> | ||
| + | |||
| + | ==Finding Parameters== | ||
| + | This program aims at fill [[Team:Paris/Modeling/Parameters|this data bank]], from experimental data obtained by our wet lab. | ||
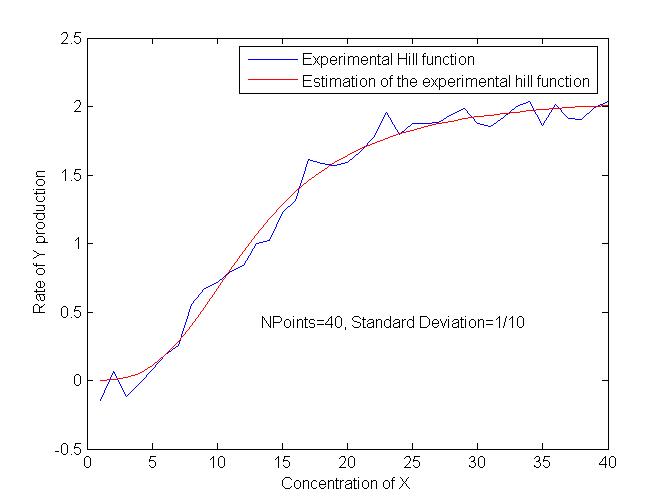
After some trials, we have obtained interesting results in terms of precision. We are now trying to test the robustness of the estimations in order to quantify the influence of biological variance in the results as well as the influence of the number of points available. | After some trials, we have obtained interesting results in terms of precision. We are now trying to test the robustness of the estimations in order to quantify the influence of biological variance in the results as well as the influence of the number of points available. | ||
Revision as of 11:48, 15 September 2008
|
 "
"