Team:University of Lethbridge/Notebook/GeneralLabAugust
From 2008.igem.org
Munima.alam (Talk | contribs) m (link to go back to the Main notebook) |
m (→August 2, 2008) |
||
| Line 15: | Line 15: | ||
| - | |||
| - | |||
| - | |||
Revision as of 04:20, 28 August 2008
Back to The University of Lethbridge Main Notebook
(Aug. 1- 7 belong in other August notebooks)
Contents |
August 1, 2008
Nathan Puhl, Roxanne
Riboswitch 20 uL PCR. Set up 4 reactions (25 uL for each total volume). 1 uL or 1/100 pTopp and 1 uL or H1/100 PCR from July 28, 2008.
PCR conditions:
A. Initial denaturation: 98 C (3 min)
B. -Denaturation: 98 C (10 sec)
- Annealing: 55 C (30 sec)
-Extension: 72 C (15 sec)
-30 cycles
C. Final extension: 72 C (7 min)
August 5, 2008
Nathan Puhl et al
Gel extracted two bands really closer to each other from PCR (August 1) at ~76 bp. Used Qiagen MiniElute kit. Results are in the hard copy lab notebook. Seems as if gel extraction was successful. ____ 2 color reporter system:
Set up digest of LacI and DT.
LacI - EcoRI and SpeI
DT - EcoRI and XbaI
Reaction mixture:
-5 uL template -5 uL NEB 2 Buffer -0.5 uL BSA? -1 uL RE #1 -1 uL RE #2 - 37.5 uL ddH20
Left at 37 C overnight. Heat deactivation (65 C) for 15 minutes.
August 6, 2008
Nathan Puhl, Roxanne
2 color reporter system:
Gel extracted LacI insert and DT vector. Didn't run gel yet.
August 7, 2008
Nathan Puhl, Roxanne
Riboswitch:
Set up PCR using purified riboswitch from Aug. 5, 2008 with platinum Taq (50 uL reaction). Made Master Mix for three reactions. Master Mix:
-10x Buffer (no Mg2+): 15 uL -10 mM dNTPs: 3 uL -50 mM Mg2+: 4.5 uL -10uM RF: 3 uL -10uM RR: 3 uL -Plat. poly: 0.6 iL -H20: 120.9 uL -template: 1 uL
Cycle conditions:
A. Initial denaturation: 94 C (2 min)
B. -Denaturation: 94 C (30 sec)
- Annealing: 55 C (30 sec)
-Extension: 72 C (30 sec)
-30 cycles
C. Final extension: 72 C (7 min)
Roxanne
-Ran the PCR Product on a 3% gel using only 1uL of DNA from the riboswitch.
August 13, 2008
Nathan Puhl and Sebastian
Verified and quantified the LacI and DT restriction digest products.
Set up a Ligase experiment for LacI and DT
- 1.3 uL vector(DT) - 8 uL insert(LacI) - 2 uL 10x Ligase buffer - 1.5 uL Ligase - 7.2 uL ddH2O - 20 uL total volume
August 14, 15
Nathan Puhl, Munima, Selina
Poured 61 LB + Amp plates. Stored in iGEM 4 C fridge.
August 15, 2008
Roxanne
Used Qiagen Plasmid MiniPrep Kit to Plasmid Prep Last Year's Biobrick Parts that were Incubated overnight.
Ran Plasmids in a 1% Agarose Gel at 100 V for 30 minutes.
August 16, 2008
Nathan Puhl, Roxanne, Munima
-Restriction Digested the purified riboswitch and pSB1A7 with XbaI and SpeI, let it run for 4 hours
Nathan Puhl, Roxanne
-Ran all of the restricted pSB1A7 plasmid through a 1% agarose gel at 100V for 25 minutes. -Ran a gel extraction on the pSB1A7 cut plasmid, and ran a PCR clean-up reaction on the digested RS1 and RS2 amplicons. -Ran 1 uL of each on a 1% to quantify the amount of DNA present. -Ligated RS1 + pSB1A7, and RS2 + pSB1A7, using T4 DNA Ligase.
-1 uL of RS1/RS2 -4 uL of pSB1A7 -1 uL of 10x T4 DNA Ligase Buffer -0.33 uL of T4 DNA Ligase -3.67 uL of water
allowed the reaction to go overnight
August 17, 2008
Nathan Puhl
-Transformed DH5a cells with the pSB1A7 + RS1, and pSB1A7 + RS2 plasmids.
-Plated on semi-solid agar plates containing 100 ug/mL of ampicillin.
August 18
Christa, Nathan Puhl, Munima
-Nathan checked the plates for growth. Colonies are present.
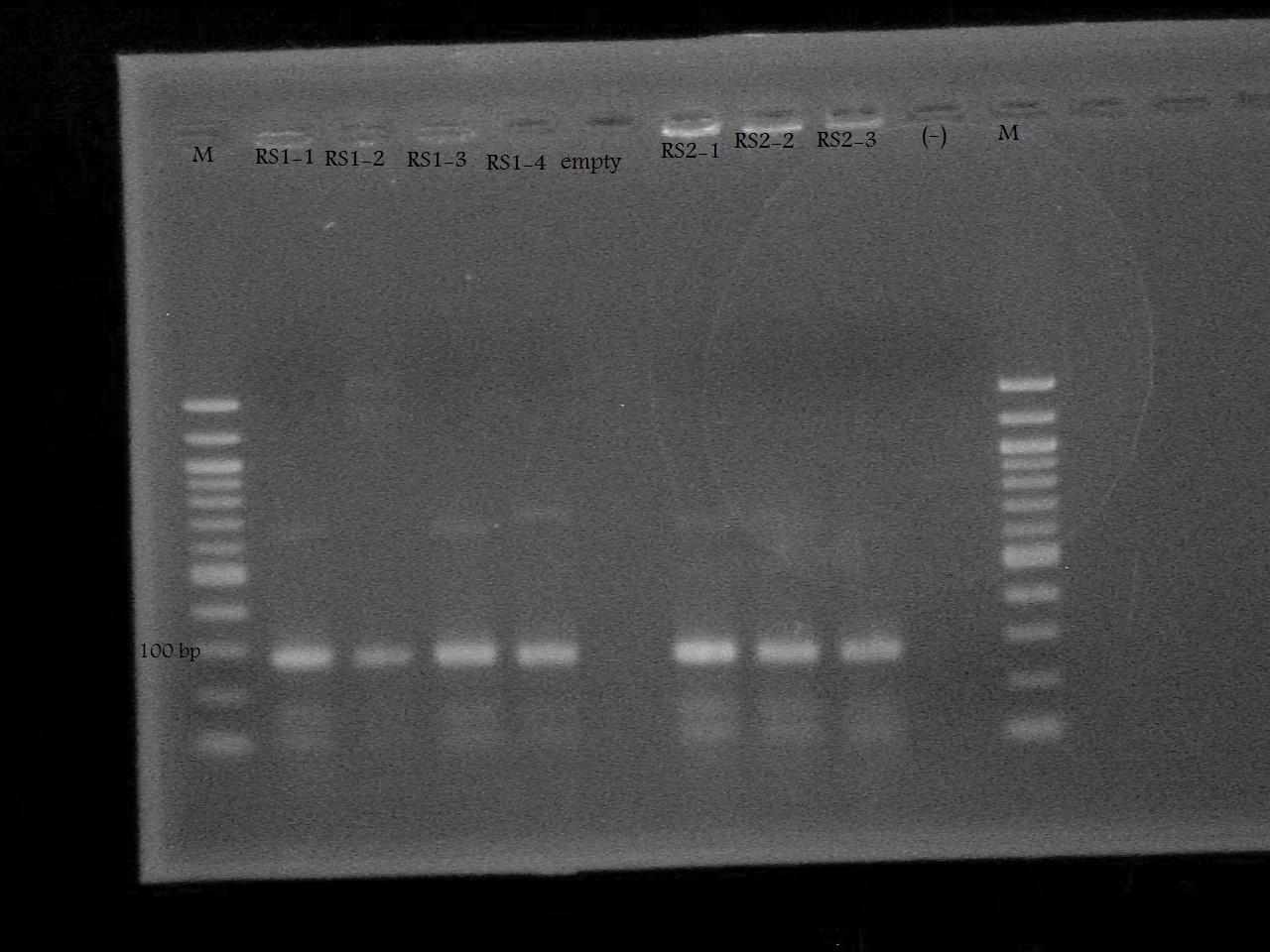
-Ran a colony PCR of the pSB1A7 + RS1, and pSB1A7 + RS2 recombinant cells transformed by Nathan and Roxanne on August 17.
-Inoculated the cells into tubes of liquid media + 100 ug/ml of ampicillin.
Roxanne will remove the PCR tube from the thermocycler in the morning.
August 19, 2008
Roxanne
-Ran the PCR Product on a 2% Agarose Gel at 100 V for 33 minutes.
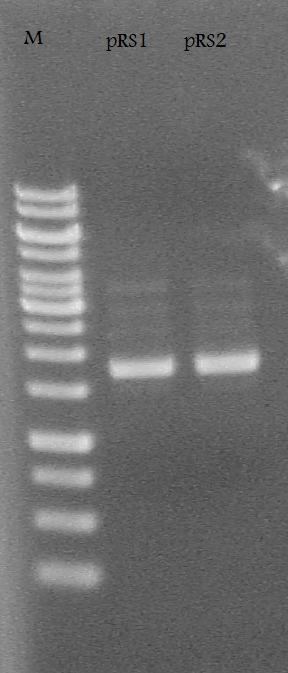
-Plasmid Prepped and made glycerol stocks from the RS1-1 and RS2-1 tubes of cells incubated in LB media + 100 ug/mL ampicillin.
Ran the pRS1 and pRS2 plasmids on a 1% Agarose Gel at 100 V for 30 minutes.
Christa, Munima, Sebastian
Did a plasmid prep on pSB1A7 using QIAprep Spin Miniprep Kit. Stocked 4- 50uL of pSB1A7 and stored in iGEM -20 C.
Sebastian, Nathan Puhl
Ran products from plasmid prep (pSB1A7 x 4 samples) on 1% agarose gel for 27 minutes.
-Lane 1 - 1 kb GeneRuler ladder (2 uL) -Lane 2 -6 pSB1A7 (3 uL) + 6x loading dye (2 uL); Mixed up what sample was in Lane 4, so Lanes 5 and 6 were run.
Conclusion: The bands appeared to be at the correct size for pSB1A7.
August 21
Nathan Puhl, Roxanne
-Screened the pSB1A7 + RS1, and pSB1A7 + RS2 by PCR using the VF2 and RS1/RS2 Reverse Primers determine whether the plasmids obtained from the recombinant cells contain the riboswitch, and if so, if it inserted in the correct orientation.
August 21, 2008
Roxanne
-Ran the PCR products on a 1% Agarose Gel at 100 V for 33 minutes. The gel was empty with the exception of primer dimers.
Nathan Puhl, Roxanne
-went over the SELEX protocol with HJ to determine the primers we will need to do this, and how exactly we plan on perfoming the evolution.
-setup a restriction digest for pSB1A7 using XbaI and SpeI, ran overnight.
August 22, 2008
Christa, Munima, Nathan Puhl, Roxanne
Objective: Run a gel to confirm that appropriate inserts were amplified from the PCRs and do a gel extraction of the inserts to prepare them for the biobrick format.
-Could not obtain a picture of the gel (1% agarose) of the half of the digested pSB1A7 (15 uL x 3 wells) and the recently amplified CheZ gene (15 uL x 3 wells) because the camera would not turn on. The CheZ gene appeared at the correct size (~700 bp). -Did a gel extraction of pSB1A7 and CheZ from that gel with the Qiagen MiniElute Gel Extraction kit. Final volume of each was 10 uL. -Ran another 1% agarose gel to confirm that the gel extraction was successful. Ran 1 uL of each sample. Bands appeared at appropriate sizes. The concentrations of pSB1A7 and CheZ were estimated to be 25 ng/uL and 80 ng/uL, respectively.
Next step: Digest pSB1A7 with antarctic phosphatase. A restriction digest will be performed at a later date on CheZ to prepare it for insertion.
August 23, 2008
Nathan Puhl, Roxanne
-Digested pSB1A7 with Antarctic Phosphatase
-9 uL of cut pSB1A7 -1.5 uL of 10x Antarctic Phosphatase Buffer -1 uL of Antarctic Phosphatase Enzyme -3.5 uL of water Allowed the Reaction to take place for 30 minutes to remove the 5` Phosphates from the pSB1A7 plasmid to prevent religation.
-Ran the remainder of the pSB1A7 plasmid from August 22nd on 1 1% Agarose Gel at 100 v for 27 minutes.
-Gel Extracted the plasmid DNA.
-Purified the Phosphatase reaction to isolate the pSB1A7 DNA.
-Ran a 1% gel to quantify the amount of plasmid DNA present.
-Ligated RS1 and RS2 into the dephosphorylated pSB1A7 using T4 DNA Ligase.
-1 uL of RS1 or RS2 -4 uL of dephosphorylated pSB1A7 -1 uL of 10X T4 DNA Ligase Buffer -0.33 uL T4 DNA Ligase Enzyme -3.67 uL water Reaction was allowed to go overnight
August 24, 2008
Nathan Puhl
-Transformed DH5a cells with the RS1+pSB1A7 or RS2+pSB1A7 plasmid on semi-solid agar plates containing 100 ug/mLof ampicillin.
 "
"