Team:University of Lethbridge/Notebook/GeneralLabAugust
From 2008.igem.org
m (→August 22) |
(→August 22, 2008) |
||
| Line 165: | Line 165: | ||
| + | |||
| + | ===August 21, 2008=== | ||
| + | ====Roxanne==== | ||
| + | -Ran the PCR products on a 1% Agarose Gel at 100 V for 33 minutes. The gel was empty with the exception of primer dimers. | ||
| + | |||
| + | ====Nathan Puhl, Roxanne==== | ||
| + | -went over the SELEX protocol with HJ to determine the primers we will need to do this, and how exactly we plan on perfoming the evolution. | ||
| + | |||
| + | -setup a restriction digest for pSB1A7 using XbaI and SpeI, ran overnight. | ||
===August 22, 2008=== | ===August 22, 2008=== | ||
| Line 170: | Line 179: | ||
Objective: Run a gel to confirm that appropriate inserts were amplified from the PCRs and do a gel extraction of the inserts to prepare them for the biobrick format. | Objective: Run a gel to confirm that appropriate inserts were amplified from the PCRs and do a gel extraction of the inserts to prepare them for the biobrick format. | ||
| - | -Could not obtain a picture of the gel (1% agarose) of the digested pSB1A7 and the recently amplified | + | -Could not obtain a picture of the gel (1% agarose) of the half of the digested pSB1A7 (15 uL x 3 wells) and |
| + | the recently amplified | ||
CheZ gene (15 uL x 3 wells) because the camera would not turn on. | CheZ gene (15 uL x 3 wells) because the camera would not turn on. | ||
The CheZ gene appeared at the correct size (~700 bp). | The CheZ gene appeared at the correct size (~700 bp). | ||
| Line 179: | Line 189: | ||
and CheZ were estimated to be 25 ng/uL and 80 ng/uL, respectively. | and CheZ were estimated to be 25 ng/uL and 80 ng/uL, respectively. | ||
| - | Next step: Digest | + | Next step: Digest pSB1A7 and cheZ with phosphatase. A restriction digest will be performed at a later date on CheZ to prepare it for insertion. |
| + | |||
| + | ===August 23, 2008=== | ||
| + | ====Nathan Puhl, Roxanne==== | ||
| + | -Digested pSB1A7 with Antartic Phosphatase | ||
| + | |||
| + | -9 uL of cut pSB1A7 | ||
| + | -1.5 uL of 10x Antarctic Phosphatase Buffer | ||
| + | -1 uL of Antarctic Phosphatase Enzyme | ||
| + | -3.5 uL of water | ||
| + | |||
| + | Allowed the Reaction to take place for 30 minutes to remove the 5` Phosphates from the pSB1A7 plasmid to | ||
| + | prevent religation. | ||
| + | |||
| + | -Ran the remainder of the pSB1A7 plasmid from August 22nd on 1 1% Agarose Gel at 100 v for 27 minutes. | ||
| + | |||
| + | -Gel Extracted the plasmid DNA. | ||
| + | |||
| + | -Purified the Phosphatase reaction to isolate the pSB1A7 DNA. | ||
| + | |||
| + | -Ran a 1% gel to quantify the amount of plasmid DNA present. | ||
| + | |||
| + | -Ligated RS1 and RS2 into the dephosphorylated pSB1A7 using T4 DNA Ligase. | ||
| + | |||
| + | -1 uL of RS1 or RS2 | ||
| + | -4 uL of dephosphorylated pSB1A7 | ||
| + | -1 uL of 10X T4 DNA Ligase Buffer | ||
| + | -0.33 uL T4 DNA Ligase Enzyme | ||
| + | -3.67 uL water | ||
| + | |||
| + | Reaction was allowed to go overnight | ||
| + | |||
| + | ===August 24, 2008=== | ||
| + | ====Nathan Puhl==== | ||
| + | -Transformed DH5a cells with the RS1+pSB1A7 or RS2+pSB1A7 plasmid on semi-solid agar plates containing 100 ug/mLof amppicillin. | ||
Revision as of 22:42, 24 August 2008
(Aug. 1- 7 belong in other August notebooks)
Contents |
August 1, 2008
Nathan Puhl, Roxanne
Riboswitch 20 uL PCR. Set up 4 reactions (25 uL for each total volume). 1 uL or 1/100 pTopp and 1 uL or H1/100 PCR from July 28, 2008.
PCR conditions:
A. Initial denaturation: 98 C (3 min)
B. -Denaturation: 98 C (10 sec)
- Annealing: 55 C (30 sec)
-Extension: 72 C (15 sec)
-30 cycles
C. Final extension: 72 C (7 min)
August 2, 2008
Nathan Puhl, Roxanne
Ran riboswitch (Aug. 1, 2008) on 3% agarose gel. Results are in the hard copy lab notebook. Looks like 76 bp band will extract.
August 5, 2008
Nathan Puhl et al
Gel extracted two bands really closer to each other from PCR (August 1) at ~76 bp. Used Qiagen MiniElute kit. Results are in the hard copy lab notebook. Seems as if gel extraction was successful. ____ 2 color reporter system:
Set up digest of LacI and DT.
LacI - EcoRI and SpeI
DT - EcoRI and XbaI
Reaction mixture:
-5 uL template -5 uL NEB 2 Buffer -0.5 uL BSA? -1 uL RE #1 -1 uL RE #2 - 37.5 uL ddH20
Left at 37 C overnight. Heat deactivation (65 C) for 15 minutes.
August 6, 2008
Nathan Puhl, Roxanne
2 color reporter system:
Gel extracted LacI insert and DT vector. Didn't run gel yet.
August 7, 2008
Nathan Puhl, Roxanne
Riboswitch:
Set up PCR using purified riboswitch from Aug. 5, 2008 with platinum Taq (50 uL reaction). Made Master Mix for three reactions. Master Mix:
-10x Buffer (no Mg2+): 15 uL -10 mM dNTPs: 3 uL -50 mM Mg2+: 4.5 uL -10uM RF: 3 uL -10uM RR: 3 uL -Plat. poly: 0.6 iL -H20: 120.9 uL -template: 1 uL
Cycle conditions:
A. Initial denaturation: 94 C (2 min)
B. -Denaturation: 94 C (30 sec)
- Annealing: 55 C (30 sec)
-Extension: 72 C (30 sec)
-30 cycles
C. Final extension: 72 C (7 min)
Roxanne
-Ran the PCR Product on a 3% gel using only 1uL of DNA from the riboswitch.
August 13, 2008
Nathan Puhl and Sebastian
Verified and quantified the LacI and DT restriction digest products.
Set up a Ligase experiment for LacI and DT
- 1.3 uL vector(DT) - 8 uL insert(LacI) - 2 uL 10x Ligase buffer - 1.5 uL Ligase - 7.2 uL ddH2O - 20 uL total volume
August 14, 15
Nathan Puhl, Munima, Selina
Poured 61 LB + Amp plates. Stored in iGEM 4 C fridge.
August 15, 2008
Roxanne
Used Qiagen Plasmid MiniPrep Kit to Plasmid Prep Last Year's Biobrick Parts that were Incubated overnight.
Ran Plasmids in a 1% Agarose Gel at 100 V for 30 minutes.
August 16, 2008
Nathan Puhl, Roxanne, Munima
-Restriction Digested the purified riboswitch and pSB1A7 with XbaI and SpeI, let it run for 4 hours
Nathan Puhl, Roxanne
-Ran all of the restricted pSB1A7 plasmid through a 1% agarose gel at 100V for 25 minutes. -Ran a gel extraction on the pSB1A7 cut plasmid, and ran a PCR clean-up reaction on the digested RS1 and RS2 amplicons. -Ran 1 uL of each on a 1% to quantify the amount of DNA present. -Ligated RS1 + pSB1A7, and RS2 + pSB1A7, using T4 DNA Ligase.
-1 uL of RS1/RS2 -4 uL of pSB1A7 -1 uL of 10x T4 DNA Ligase Buffer -0.33 uL of T4 DNA Ligase -3.67 uL of water
allowed the reaction to go overnight
August 17, 2008
Nathan Puhl
-Transformed DH5a cells with the pSB1A7 + RS1, and pSB1A7 + RS2 plasmids.
-Plated on semi-solid agar plates containing 100 ug/mL of ampicillin.
August 18
Christa, Nathan Puhl, Munima
-Nathan checked the plates for growth. Colonies are present.
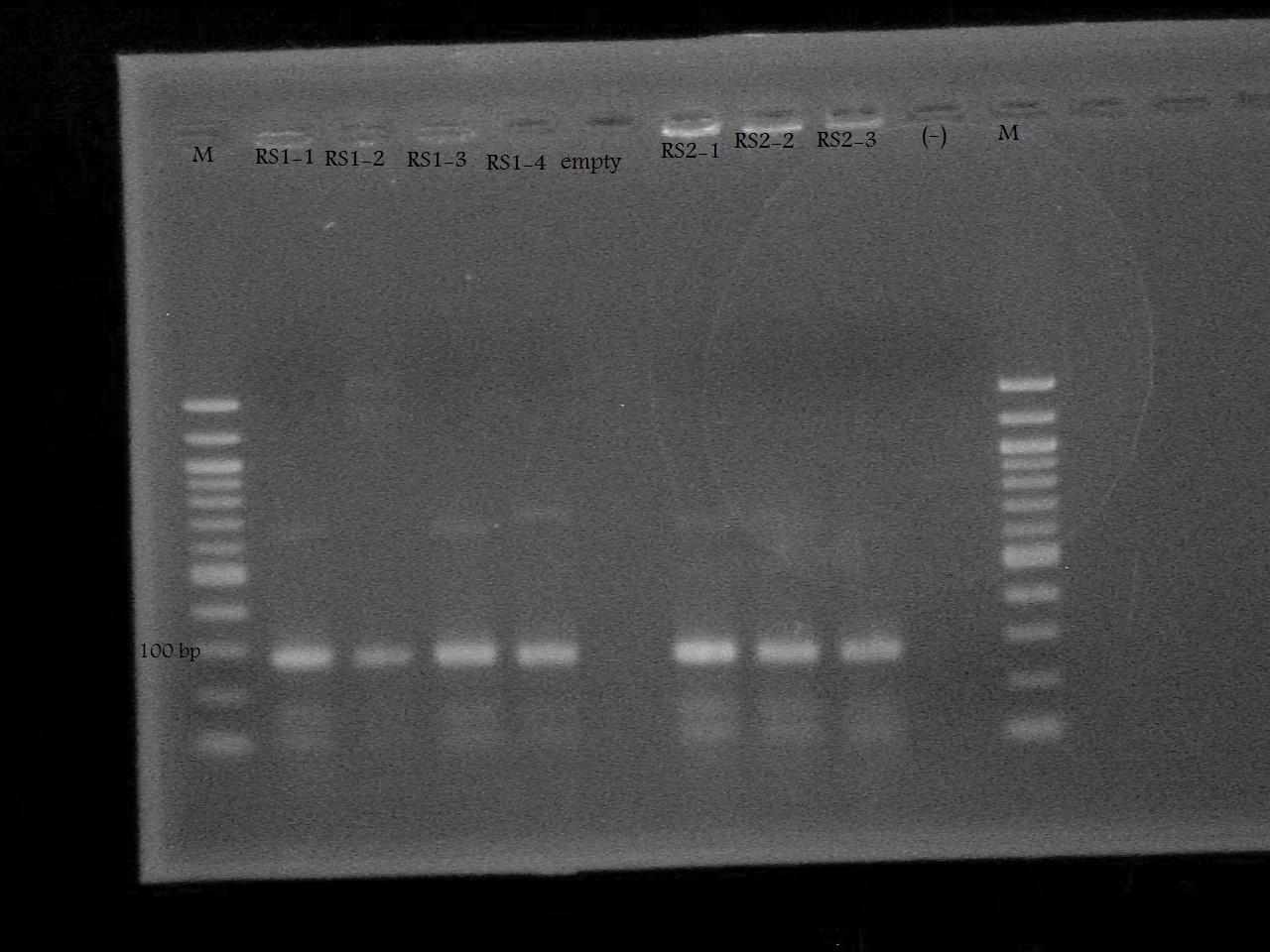
-Ran a colony PCR of the pSB1A7 + RS1, and pSB1A7 + RS2 recombinant cells transformed by Nathan and Roxanne on August 17.
-Inoculated the cells into tubes of liquid media + 100 ug/ml of ampicillin.
Roxanne will remove the PCR tube from the thermocycler in the morning.
August 19, 2008
Roxanne
-Ran the PCR Product on a 2% Agarose Gel at 100 V for 33 minutes.
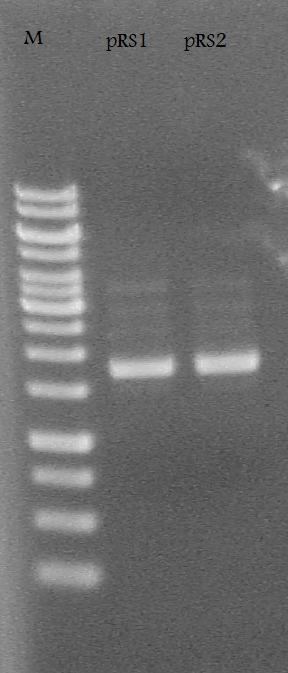
-Plasmid Prepped and made glycerol stocks from the RS1-1 and RS2-1 tubes of cells incubated in LB media + 100 ug/mL ampicillin.
Ran the pRS1 and pRS2 plasmids on a 1% Agarose Gel at 100 V for 30 minutes.
Christa, Munima, Sebastian
Did a plasmid prep on pSB1A7 using QIAprep Spin Miniprep Kit. Stocked 4- 50uL of pSB1A7 and stored in iGEM -20 C.
Sebastian, Nathan Puhl
Ran products from plasmid prep (pSB1A7 x 4 samples) on 1% agarose gel for 27 minutes.
-Lane 1 - 1 kb GeneRuler ladder (2 uL) -Lane 2 -6 pSB1A7 (3 uL) + 6x loading dye (2 uL); Mixed up what sample was in Lane 4, so Lanes 5 and 6 were run.
Conclusion: The bands appeared to be at the correct size for pSB1A7.
August 21
Nathan Puhl, Roxanne
-Screened the pSB1A7 + RS1, and pSB1A7 + RS2 by PCR using the VF2 and RS1/RS2 Reverse Primers determine whether the plasmids obtained from the recombinant cells contain the riboswitch, and if so, if it inserted in the correct orientation.
August 21, 2008
Roxanne
-Ran the PCR products on a 1% Agarose Gel at 100 V for 33 minutes. The gel was empty with the exception of primer dimers.
Nathan Puhl, Roxanne
-went over the SELEX protocol with HJ to determine the primers we will need to do this, and how exactly we plan on perfoming the evolution.
-setup a restriction digest for pSB1A7 using XbaI and SpeI, ran overnight.
August 22, 2008
Christa, Munima, Nathan Puhl, Roxanne
Objective: Run a gel to confirm that appropriate inserts were amplified from the PCRs and do a gel extraction of the inserts to prepare them for the biobrick format.
-Could not obtain a picture of the gel (1% agarose) of the half of the digested pSB1A7 (15 uL x 3 wells) and the recently amplified CheZ gene (15 uL x 3 wells) because the camera would not turn on. The CheZ gene appeared at the correct size (~700 bp). -Did a gel extraction of pSB1A7 and CheZ from that gel with the Qiagen MiniElute Gel Extraction kit. Final volume of each was 10 uL. -Ran another 1% agarose gel to confirm that the gel extraction was successful. Ran 1 uL of each sample. Bands appeared at appropriate sizes. The concentrations of pSB1A7 and CheZ were estimated to be 25 ng/uL and 80 ng/uL, respectively.
Next step: Digest pSB1A7 and cheZ with phosphatase. A restriction digest will be performed at a later date on CheZ to prepare it for insertion.
August 23, 2008
Nathan Puhl, Roxanne
-Digested pSB1A7 with Antartic Phosphatase
-9 uL of cut pSB1A7 -1.5 uL of 10x Antarctic Phosphatase Buffer -1 uL of Antarctic Phosphatase Enzyme -3.5 uL of water Allowed the Reaction to take place for 30 minutes to remove the 5` Phosphates from the pSB1A7 plasmid to prevent religation.
-Ran the remainder of the pSB1A7 plasmid from August 22nd on 1 1% Agarose Gel at 100 v for 27 minutes.
-Gel Extracted the plasmid DNA.
-Purified the Phosphatase reaction to isolate the pSB1A7 DNA.
-Ran a 1% gel to quantify the amount of plasmid DNA present.
-Ligated RS1 and RS2 into the dephosphorylated pSB1A7 using T4 DNA Ligase.
-1 uL of RS1 or RS2 -4 uL of dephosphorylated pSB1A7 -1 uL of 10X T4 DNA Ligase Buffer -0.33 uL T4 DNA Ligase Enzyme -3.67 uL water Reaction was allowed to go overnight
August 24, 2008
Nathan Puhl
-Transformed DH5a cells with the RS1+pSB1A7 or RS2+pSB1A7 plasmid on semi-solid agar plates containing 100 ug/mLof amppicillin.
 "
"