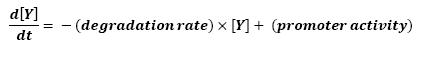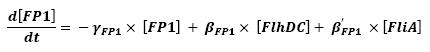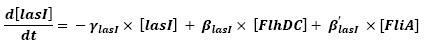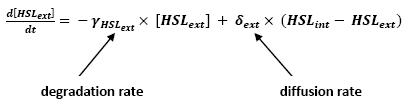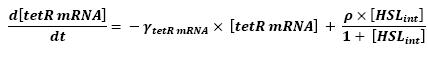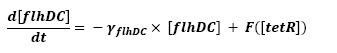Team:Paris/Modeling
From 2008.igem.org
(→An Oscillatory Biological Model) |
(→An Oscillatory Biological Model) |
||
| Line 45: | Line 45: | ||
Musch of our inspiration comes from four articles to which we shall refer in the next subsections : | Musch of our inspiration comes from four articles to which we shall refer in the next subsections : | ||
<br> | <br> | ||
| - | + | * [1] Shiraz Kalir, Uri Alon. Using quantitative blueprint to reprogram the dynamics of the flagella network. Cell, June 11, 2004, Vol.117, 713-720. | |
| - | + | *[2]Jordi Garcia-Ojalvo, Michael B. Elowitz, Steven H. Strogratz. Modeling a synthetic multicellular clock : repressilators coupled by quorum sensing. PNAS, July 27, 204, Vol. 101, no. 30. | |
| - | + | * [3]Nitzan Rosenfeld, Uri Alon. Response delays and the structure of transcription networks. JMB, 2003, 329, 645-654. | |
| - | + | * [4]Nitzan Rosenfeld, Michael B. Elowitz, Uri Alon. Negative autoregulation speeds the response times of transcription networks. JMB, 2003, 323, 785-793. | |
===Equations=== | ===Equations=== | ||
| Line 85: | Line 85: | ||
* Asumptions and Discussion | * Asumptions and Discussion | ||
| - | We can see that there is a retroaction from fliA | + | ** We can see that there is a retroaction from fliA over fliA. In a first approach, we decided not to take into account this interaction. This will be seen in a second step. |
| - | Another step consists in choosing the proper units, so as to have a homogenous set of parameters. | + | ** Another step consists in choosing the proper units, so as to have a homogenous set of parameters. |
<br> | <br> | ||
| Line 100: | Line 100: | ||
* Discution and Asumptions | * Discution and Asumptions | ||
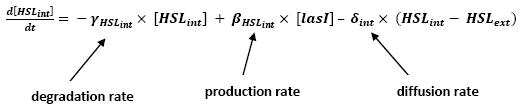
| - | Even though we considered a single cell, we decided to model both HSL inside and outside the cell. In a first approach, we assumed that HSL could be modelized in the same fashion as AHL. The process was well detailed in [2] and we adapted their model to our present situation. | + | ** Even though we considered a single cell, we decided to model both HSL inside and outside the cell. In a first approach, we assumed that HSL could be modelized in the same fashion as AHL. |
| + | <br>The process was well detailed in [2] and we adapted their model to our present situation. | ||
| + | <br>Furthermore, a biologically coherent set of parameters is also given in this article. We decided then to use them as well for our simulation. | ||
| + | ** As explained before, in this model, we only considered a single cell. The method to model the interactions with other cells is also described. | ||
<br> | <br> | ||
---- | ---- | ||
| - | + | * Steps | |
| - | * HSL<sub>int</sub> → tetR mRNA (12) | + | ** HSL<sub>int</sub> → tetR mRNA (12) |
| - | * tetR mRNA → tet R (13) | + | ** tetR mRNA → tet R (13) |
| - | + | * Equations | |
[[Image:tetR mRNA.jpg|center]] | [[Image:tetR mRNA.jpg|center]] | ||
[[Image:tetR.jpg|center]] | [[Image:tetR.jpg|center]] | ||
| + | |||
| + | * Discution and Asumptions | ||
| + | ** Here is the only case where we have not yet found out how we could manage to skip the mRNA step. Then, (12) represents the binding and transcription steps ; (13) represents the translation step. | ||
| + | |||
<br> | <br> | ||
Revision as of 14:45, 6 August 2008
|
 "
"