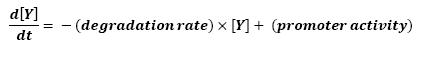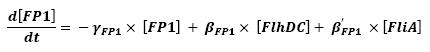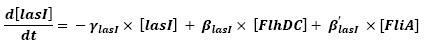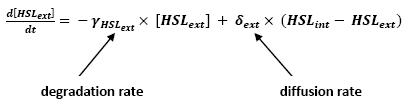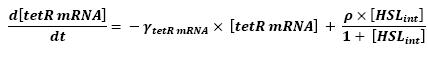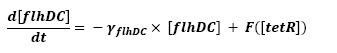Team:Paris/Modeling
From 2008.igem.org
(→An Oscillatory Biological Model) |
(→An Oscillatory Biological Model) |
||
| Line 51: | Line 51: | ||
===Equations=== | ===Equations=== | ||
| - | * Steps | + | * <b>Steps</b> |
** flhDC → fliA (1) | ** flhDC → fliA (1) | ||
** flhDC → fliL → Fluorescent Protein 1 (FP1) (2) | ** flhDC → fliL → Fluorescent Protein 1 (FP1) (2) | ||
| Line 65: | Line 65: | ||
<br> | <br> | ||
| - | * Model | + | * <b>Model</b> |
The transcription hierarchy of the flagella genes have been studied in [1]. It resuted that the promoter activity of the seven class 2 operons, among which fLiL, flgA, flhB, may be mathematically described in that way : | The transcription hierarchy of the flagella genes have been studied in [1]. It resuted that the promoter activity of the seven class 2 operons, among which fLiL, flgA, flhB, may be mathematically described in that way : | ||
<br> | <br> | ||
| Line 84: | Line 84: | ||
[[Image:lasI.jpg|center]] | [[Image:lasI.jpg|center]] | ||
| - | * | + | * <b>Discution and Asumptions</b> |
| - | ** | + | ** As presented in the first equation below, there is a retroaction from fliA over fliA. In a first approach, we decided not to take into account this interaction, that is setting <Beta>. This will be seen in a second step. |
** Another step consists in choosing the proper units, so as to have a homogenous set of parameters. | ** Another step consists in choosing the proper units, so as to have a homogenous set of parameters. | ||
| Line 91: | Line 91: | ||
---- | ---- | ||
| - | * Steps | + | * <b>Steps</b> |
** lasI → HSL<sub>ext</sub> (10) | ** lasI → HSL<sub>ext</sub> (10) | ||
** lasI → HSL<sub>int</sub> (11) | ** lasI → HSL<sub>int</sub> (11) | ||
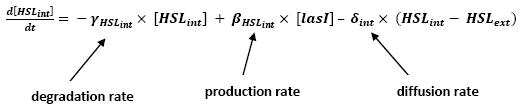
| - | * | + | * <b>Model</b> |
[[Image:HSLint.jpg|center]] | [[Image:HSLint.jpg|center]] | ||
[[Image:HSLext.jpg|center]] | [[Image:HSLext.jpg|center]] | ||
| - | * Discution and Asumptions | + | * <b>Discution and Asumptions</b> |
** Even though we considered a single cell, we decided to model both HSL inside and outside the cell. In a first approach, we assumed that HSL could be modelized in the same fashion as AHL. | ** Even though we considered a single cell, we decided to model both HSL inside and outside the cell. In a first approach, we assumed that HSL could be modelized in the same fashion as AHL. | ||
<br>The process was well detailed in [2] and we adapted their model to our present situation. | <br>The process was well detailed in [2] and we adapted their model to our present situation. | ||
| Line 107: | Line 107: | ||
<br> | <br> | ||
---- | ---- | ||
| - | * Steps | + | * <b>Steps</b> |
** HSL<sub>int</sub> → tetR mRNA (12) | ** HSL<sub>int</sub> → tetR mRNA (12) | ||
** tetR mRNA → tet R (13) | ** tetR mRNA → tet R (13) | ||
| - | * | + | * <b>Model</b> |
[[Image:tetR mRNA.jpg|center]] | [[Image:tetR mRNA.jpg|center]] | ||
[[Image:tetR.jpg|center]] | [[Image:tetR.jpg|center]] | ||
| - | * Discution and Asumptions | + | * <b>Discution and Asumptions</b> |
** Here is the only case where we have not yet found out how we could manage to skip the mRNA step. Then, (12) represents the binding and transcription steps ; (13) represents the translation step. | ** Here is the only case where we have not yet found out how we could manage to skip the mRNA step. Then, (12) represents the binding and transcription steps ; (13) represents the translation step. | ||
Revision as of 14:49, 6 August 2008
|
 "
"