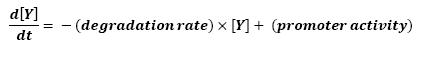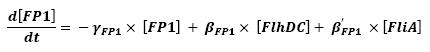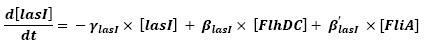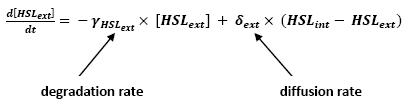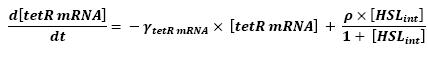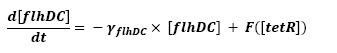Team:Paris/Modeling
From 2008.igem.org
(→Parameters summary) |
(→Parameters summary) |
||
| Line 154: | Line 154: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |γ<sub>FliA</sub> | | style="background: #D4E2EF;" |γ<sub>FliA</sub> | ||
| - | | | + | | Degradation rate |
| GTTTCTTCGAATTCGCGGCCGCTTCTAGAGAGCATATCTCCTCCGCAGGTATCAAAAT | | GTTTCTTCGAATTCGCGGCCGCTTCTAGAGAGCATATCTCCTCCGCAGGTATCAAAAT | ||
| 58 | | 58 | ||
| Line 161: | Line 161: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |β<sub>FliA</sub> | | style="background: #D4E2EF;" |β<sub>FliA</sub> | ||
| - | | | + | | FlhDC activation coefficient |
| GTTTCTTCCTGCAGCGGCCGCTACTAGTAACAGTATCGCGATGATCGCCACGCTACGT | | GTTTCTTCCTGCAGCGGCCGCTACTAGTAACAGTATCGCGATGATCGCCACGCTACGT | ||
| 58 | | 58 | ||
| Line 168: | Line 168: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |β'<sub>FliA</sub> | | style="background: #D4E2EF;" |β'<sub>FliA</sub> | ||
| - | | | + | | FliA activation coefficient |
| GTTTCTTCGAATTCGCGGCCGCTTCTAGAGACAGTATCGCGATGATCGCCACGCTACG | | GTTTCTTCGAATTCGCGGCCGCTTCTAGAGACAGTATCGCGATGATCGCCACGCTACG | ||
| 58 | | 58 | ||
| Line 175: | Line 175: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |γ<sub>FP1</sub> | | style="background: #D4E2EF;" |γ<sub>FP1</sub> | ||
| - | | | + | | Degradation rate |
| GTTTCTTCCTGCAGCGGCCGCTACTAGTAAGCATATCTCCTCCGCAGGTATCAAAATT | | GTTTCTTCCTGCAGCGGCCGCTACTAGTAAGCATATCTCCTCCGCAGGTATCAAAATT | ||
| 58 | | 58 | ||
| Line 182: | Line 182: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |β<sub>FP1</sub> | | style="background: #D4E2EF;" |β<sub>FP1</sub> | ||
| - | | | + | | FlhDC activation coefficient |
| GTTTCTTCGAATTCGCGGCCGCTTCTAGAGCAGCAGATGAAATCGATAAGCGACAAGG | | GTTTCTTCGAATTCGCGGCCGCTTCTAGAGCAGCAGATGAAATCGATAAGCGACAAGG | ||
| 58 | | 58 | ||
| Line 189: | Line 189: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |β'<sub>FP1</sub> | | style="background: #D4E2EF;" |β'<sub>FP1</sub> | ||
| - | | | + | | FliA activation coefficient |
| GTTTCTTCCTGCAGCGGCCGCTACTAGTAAACGCCGCCTGGGCCGCGTTCAGTCCGCT | | GTTTCTTCCTGCAGCGGCCGCTACTAGTAAACGCCGCCTGGGCCGCGTTCAGTCCGCT | ||
| 58 | | 58 | ||
| Line 196: | Line 196: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |γ<sub>FP2</sub> | | style="background: #D4E2EF;" |γ<sub>FP2</sub> | ||
| - | | | + | | Degradation rate |
| GTTTCTTCGAATTCGCGGCCGCTTCTAGAGAGCGGCGTTGTTGATGCAGATGGGAATA | | GTTTCTTCGAATTCGCGGCCGCTTCTAGAGAGCGGCGTTGTTGATGCAGATGGGAATA | ||
| 58 | | 58 | ||
| Line 203: | Line 203: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |β<sub>FP2</sub> | | style="background: #D4E2EF;" |β<sub>FP2</sub> | ||
| - | | | + | | FlhDC activation coefficient |
| GTTTCTTCCTGCAGCGGCCGCTACTAGTACGAAGTGCGATCAATACTCATGGTTTATT | | GTTTCTTCCTGCAGCGGCCGCTACTAGTACGAAGTGCGATCAATACTCATGGTTTATT | ||
| 58 | | 58 | ||
| Line 210: | Line 210: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |β'<sub>FP2</sub> | | style="background: #D4E2EF;" |β'<sub>FP2</sub> | ||
| - | | | + | | FliA activation coefficient |
| GTTTCTTCGAATTCGCGGCCGCTTCTAGAGCCACGTCATATCAGGCGGTCTGATAAGG | | GTTTCTTCGAATTCGCGGCCGCTTCTAGAGCCACGTCATATCAGGCGGTCTGATAAGG | ||
| 58 | | 58 | ||
| Line 217: | Line 217: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |γ<sub>FP3</sub> | | style="background: #D4E2EF;" |γ<sub>FP3</sub> | ||
| - | | | + | | Degradation rate |
| GTTTCTTCCTGCAGCGGCCGCTACTAGTAGTTTTGTCGTCGCTCTCGTCAGACACGTC | | GTTTCTTCCTGCAGCGGCCGCTACTAGTAGTTTTGTCGTCGCTCTCGTCAGACACGTC | ||
| 58 | | 58 | ||
| Line 224: | Line 224: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |β<sub>FP3</sub> | | style="background: #D4E2EF;" |β<sub>FP3</sub> | ||
| - | | FlhDC | + | | FlhDC activation coefficient |
| GTTTCTTCGAATTCGCGGCCGCTTCTAGAGTTGTATGTGCGTGTAGTGACGAGTACAG | | GTTTCTTCGAATTCGCGGCCGCTTCTAGAGTTGTATGTGCGTGTAGTGACGAGTACAG | ||
| 58 | | 58 | ||
| Line 230: | Line 230: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |β'<sub>FP3</sub> | | style="background: #D4E2EF;" |β'<sub>FP3</sub> | ||
| - | | | + | | FliA activation coefficient |
| GTTTCTTCGAATTCGCGGCCGCTTCTAGAGTCATTTTTGCTTGCTAGCGTACGGAAAA | | GTTTCTTCGAATTCGCGGCCGCTTCTAGAGTCATTTTTGCTTGCTAGCGTACGGAAAA | ||
| 58 | | 58 | ||
| Line 236: | Line 236: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |γ<sub>lasI</sub> | | style="background: #D4E2EF;" |γ<sub>lasI</sub> | ||
| - | | | + | | Degradation rate |
| GTTTCTTCCTGCAGCGGCCGCTACTAGTATCCCACCCAGAATAACCAACTTTAT | | GTTTCTTCCTGCAGCGGCCGCTACTAGTATCCCACCCAGAATAACCAACTTTAT | ||
| 54 | | 54 | ||
| | | | ||
| - | |||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |β<sub>lasI</sub> | | style="background: #D4E2EF;" |β<sub>lasI</sub> | ||
| - | | FlhDC | + | | FlhDC activation coefficient |
| GTTTCTTCCTGCAGCGGCCGCTACTAGTACAGAATAACCAACTTTATTTTTATG | | GTTTCTTCCTGCAGCGGCCGCTACTAGTACAGAATAACCAACTTTATTTTTATG | ||
| 54 | | 54 | ||
| | | | ||
| - | |||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |β'<sub>lasI</sub> | | style="background: #D4E2EF;" |β'<sub>lasI</sub> | ||
| - | | FliA | + | | FliA activation coefficient |
| GTTTCTTCGAATTCGCGGCCGCTTCTAGAGCCTGATTAACTGAGACTGACGGCAACGC | | GTTTCTTCGAATTCGCGGCCGCTTCTAGAGCCTGATTAACTGAGACTGACGGCAACGC | ||
| 58 | | 58 | ||
| Line 257: | Line 255: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |γ<sub>HSL<sub>int</sub></sub> | | style="background: #D4E2EF;" |γ<sub>HSL<sub>int</sub></sub> | ||
| - | | | + | | Degradation rate |
| CAGCCCTGCGTTATATGAGTTATCGGCATGATTATCCGTTTCCGCAGGGTTTTTAATCG | | CAGCCCTGCGTTATATGAGTTATCGGCATGATTATCCGTTTCCGCAGGGTTTTTAATCG | ||
| 59 | | 59 | ||
| Line 264: | Line 262: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |β<sub>HSL<sub>int</sub></sub> | | style="background: #D4E2EF;" |β<sub>HSL<sub>int</sub></sub> | ||
| - | | | + | | Degradation rate |
| GAAACGGATAATCATGCCGATAACTCATATAACGCAGGGCTG | | GAAACGGATAATCATGCCGATAACTCATATAACGCAGGGCTG | ||
| 42 | | 42 | ||
| Line 279: | Line 277: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |γ<sub>HSL<sub>ext</sub></sub> | | style="background: #D4E2EF;" |γ<sub>HSL<sub>ext</sub></sub> | ||
| - | | | + | | Degradation rate |
| GTTTCTTCGAATTCGCGGCCGCTTCTAGAGCCAGCGATGAAATACTTGCCATGCGATT | | GTTTCTTCGAATTCGCGGCCGCTTCTAGAGCCAGCGATGAAATACTTGCCATGCGATT | ||
| 58 | | 58 | ||
| Line 293: | Line 291: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |γ<sub>tetR mRNA</sub> | | style="background: #D4E2EF;" |γ<sub>tetR mRNA</sub> | ||
| - | | | + | | Degradation rate |
| GTTTCTTCGAATTCGCGGCCGCTTCTAGAGGTCGCATTCATCGCGCACCTCGTGGCTG | | GTTTCTTCGAATTCGCGGCCGCTTCTAGAGGTCGCATTCATCGCGCACCTCGTGGCTG | ||
| 58 | | 58 | ||
| Line 314: | Line 312: | ||
|- style="background: #dddddd;" | |- style="background: #dddddd;" | ||
| style="background: #D4E2EF;" |γ<sub>flhDC</sub> | | style="background: #D4E2EF;" |γ<sub>flhDC</sub> | ||
| - | | | + | | Degradation rate |
| GTTTCTTCCTGCAGCGGCCGCTACTAGTAGTCGCATTCATCGCGCACCTCGTGGCTGA | | GTTTCTTCCTGCAGCGGCCGCTACTAGTAGTCGCATTCATCGCGCACCTCGTGGCTGA | ||
| 58 | | 58 | ||
Revision as of 17:10, 6 August 2008
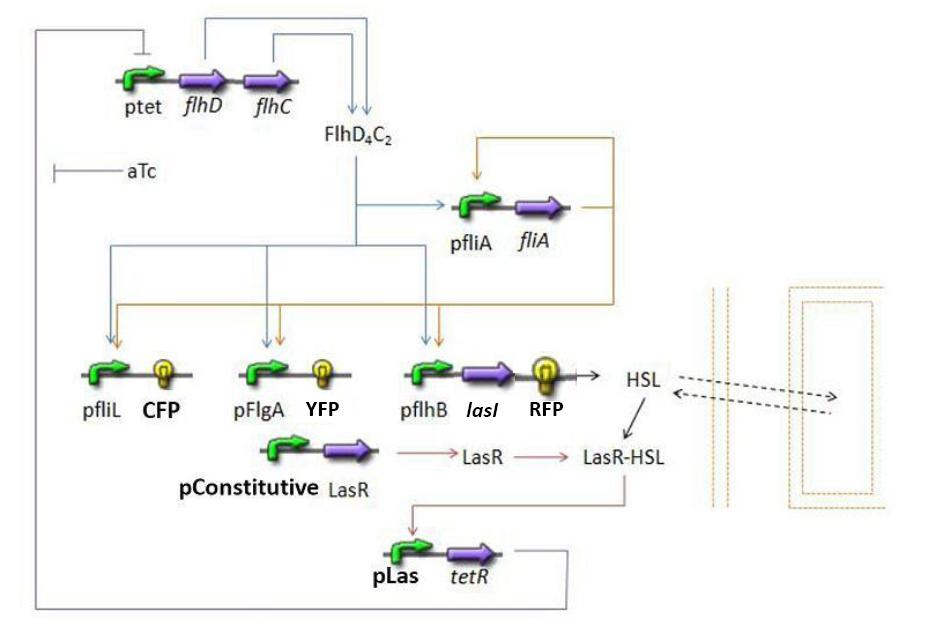
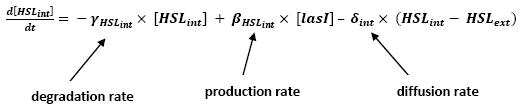
Graph screenshotsFirst Mathematical ApproachIntroductionAs a first approach, we had decided to take into account the binding to the promoters steps. Moreover, the transcription rates were expected to be Hill functions. Obvisouly, this modeling requires a huge number of parameters. To obtain them, we had planed to devise specific experiments (described below). Nonetheless, after reading some more articles, we have decided to change several asumptions of the modeling choice. Therefore, we have devised a perhaps more biologically relevant framework (see above). This part describes in detail the first approach and the codes that have been produced. First ApproachAs a first approximation, we have proposed a set of 5 ordinary differential equations, without taking into account the transcription step. Besides, we have had not introduced yet a synchronizaton module. Therefore, the repression of FlhDC is directly modeled by the presence of the 'Z3' gene (that is the last that is turned on). In this framework, we have found parameters that have provided oscillations as well as a function that automatically detects whether the output of the ode system is oscillating. This has allowed to screen a little the parameters used, in order to evaluate the robustness of the system. The methods employed are described there : First Approach. More precise Bio-Mathematical DescriptionAfter trying to obtain oscillations from a simple model, we have tried to described more precisely the studied system. Therefore, we have obtained the following formalism : Bio-Mathematical Description. BibliographyIn order to choose a proper modeling approach for our system, we have decided to list all the chemical reactions we will take into account. Afterwards, we will find the needed parameters reading articles or devising the required experiments. An overview of the work that has to be done can be found here : Parameters we have to use. Estimation of parametersThen, we will need many parameters to fully desribe the system according to the asumptions of the previous section. A natural way to have access to their value, after searching in the litterature, is to devise specific experiments. As a consequence of the characterization of the promoters activity, some Hill functions could be obtained. Thus, we have described the experimental approach required : Estimation of the parameters. Nonetheless, as mentioned above, we have changed the way to model the biological reactions. As a result have stopped investigating in this way to focus on the An Oscillatory Biological Model.
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"