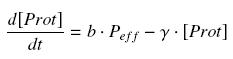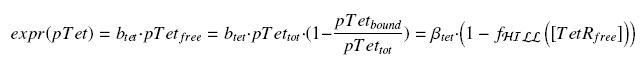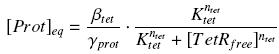Team:Paris/Modeling/estimation
From 2008.igem.org
(→first hypothesis) |
|||
| Line 44: | Line 44: | ||
===how to control the concentration of the transcription factor ?=== | ===how to control the concentration of the transcription factor ?=== | ||
| - | ====<center>By aTc / TetR / | + | ====<center>By aTc / TetR / pTet </center>==== |
| - | Now, we must use as a variable of reference an element that could be introduced in the bacteria, well-controlled, and from which all the concentrations of our transcription factor will depend. We propose a construction in which our transcription factor is put after the promoter | + | Now, we must use as a variable of reference an element that could be introduced in the bacteria, well-controlled, and from which all the concentrations of our transcription factor will depend. We propose a construction in which our transcription factor is put after the promoter pTet, which is under the repression of TetR. Since aTc is a small diffusive molecule that binds to TetR and inhibits this way the repression of pTet, we can use it as an 'inducer'. To do so, we must place in the bacterium the gene ''tetR'' after a constitutive promoter (like J23101). According to previous hypothesis, this will provide at steady-state a 'constant concentration' of TetR (we note [TetR]<sub>tot</sub>, and it is supposed to be the TOTAL concentration of TetR, under every form) in the bacterium. If we consider the binding reaction this way (where aTc_-_TetR denotes the complex) |
<center> [[Image:aTcTetRn.jpg]]</center> | <center> [[Image:aTcTetRn.jpg]]</center> | ||
| Line 56: | Line 56: | ||
where [aTc] denotes the concentration of aTc we introduced in the medium, that will stay constant in all the bacteria along time, assuming that its degradation is near 0, and that the diffusion is quick. | where [aTc] denotes the concentration of aTc we introduced in the medium, that will stay constant in all the bacteria along time, assuming that its degradation is near 0, and that the diffusion is quick. | ||
| - | According to the [[Team:Paris/Modeling#first_Hypothesis|hypothesis '''(3)''']], the activity of | + | According to the [[Team:Paris/Modeling#first_Hypothesis|hypothesis '''(3)''']], the activity of pTet would verify (keeping the same notations) : |
<center> [[Image:exprTetR.jpg]]</center> | <center> [[Image:exprTetR.jpg]]</center> | ||
| Line 68: | Line 68: | ||
Then, we intend to program a new algorithm, based on the same principles of 'findparam' for a classic ''hill function'', but which is seeking more parameters. | Then, we intend to program a new algorithm, based on the same principles of 'findparam' for a classic ''hill function'', but which is seeking more parameters. | ||
| - | ====<center>By AraC / Arabinose / | + | ====<center>By AraC / Arabinose / pBad </center>==== |
| - | An other well-known promoter called | + | An other well-known promoter called pBad, is induced by the complex AraC_-_Arabinose, where AraC appears to be a protein ''constitutively produced'' by an operon attached to pBad, and Arabinose is a sugar, that we can add at will in the medium, and diffuses through cells. |
Then, our "inducer" will be the Arabinose, that we will treat as an ''abstract variable'' without intending to know the concentrations of AraC, or of the complex. | Then, our "inducer" will be the Arabinose, that we will treat as an ''abstract variable'' without intending to know the concentrations of AraC, or of the complex. | ||
| Line 88: | Line 88: | ||
*[[Team:Paris/Modeling/f0|[expr.(''J23101'')] = ƒ0]] | *[[Team:Paris/Modeling/f0|[expr.(''J23101'')] = ƒ0]] | ||
| - | *[[Team:Paris/Modeling/f1|[expr.( | + | *[[Team:Paris/Modeling/f1|[expr.(pTet)] = ƒ1([TetR],[aTc])]] |
| - | *[[Team:Paris/Modeling/f2|[expr.( | + | *[[Team:Paris/Modeling/f2|[expr.(pBad)] = ƒ2([Arabinose])]] |
| - | According to the [[Team:Paris/Modeling#first_hypothesis|hypothesis '''(1)''' and '''(2)''']], we assume this will directly give us [Protein] = ƒ0, [Protein] = ƒ1([aTc]) or [Protein] = ƒ2([Arabinose]) for a given Protein coded by a gene put behind the ''J23101'', | + | According to the [[Team:Paris/Modeling#first_hypothesis|hypothesis '''(1)''' and '''(2)''']], we assume this will directly give us [Protein] = ƒ0, [Protein] = ƒ1([aTc]) or [Protein] = ƒ2([Arabinose]) for a given Protein coded by a gene put behind the ''J23101'', pBad or pTet promoter. |
| - | *[[Team:Paris/Modeling/f3|[expr.( | + | *[[Team:Paris/Modeling/f3|[expr.(pFlhDC)] = ƒ3([OmpR*],[FliA])]] |
| - | *[[Team:Paris/Modeling/f3|[expr.( | + | *[[Team:Paris/Modeling/f3|[expr.(pFlhDC)] = ƒ3bis([EnvZ*],[FliA])]] |
| - | *[[Team:Paris/Modeling/f4|[expr.( | + | *[[Team:Paris/Modeling/f4|[expr.(pFlhDC)] = ƒ4([FlhDC],[FliA])]] |
| - | *[[Team:Paris/Modeling/f5|[expr.( | + | *[[Team:Paris/Modeling/f5|[expr.(pFliL)] = ƒ5([FlhDC],[FliA])]] |
| - | *[[Team:Paris/Modeling/f6|[expr.( | + | *[[Team:Paris/Modeling/f6|[expr.(pFlgA)] = ƒ6([FlhDC],[FliA])]] |
| - | *[[Team:Paris/Modeling/f7|[expr.( | + | *[[Team:Paris/Modeling/f7|[expr.(pFlgB)] = ƒ7([FlhDC],[FliA])]] |
| - | *[[Team:Paris/Modeling/f8|[expr.( | + | *[[Team:Paris/Modeling/f8|[expr.(pFlhB)] = ƒ8([FlhDC],[FliA])]] |
|}<br style="clear:both" /> | |}<br style="clear:both" /> | ||
Revision as of 14:16, 19 August 2008
|
 "
"