Team:Paris/August 27
From 2008.igem.org
(Difference between revisions)
(→Cloning of EnvZ* in pSB1A2) |
|||
| (40 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{Paris/Calendar_Links|August 26|August 28}} | {{Paris/Calendar_Links|August 26|August 28}} | ||
| - | + | =Construction of pFlhB - mRFP Tripart (LVA+)= | |
| - | = | + | |
Aim : Construction of ''' "pFlhB-RBS-mRFP-dbl ter" (pFlhB-I732078)''' [[Image:Part_icon_regulatory.png]][[Image:Part_icon_rbs.png]][[Image:icon_reporter.png]][[Image:Part_icon_terminator.png]][[Image:Part_icon_terminator.png]] | Aim : Construction of ''' "pFlhB-RBS-mRFP-dbl ter" (pFlhB-I732078)''' [[Image:Part_icon_regulatory.png]][[Image:Part_icon_rbs.png]][[Image:icon_reporter.png]][[Image:Part_icon_terminator.png]][[Image:Part_icon_terminator.png]] | ||
| - | We can only do the construction with mRFP Tripart (LVA+) because the stable strain with the Biobricks I732011 (mRFP Tripart LVA-) don't to growth. | + | <br>We can only do the construction with mRFP Tripart (LVA+) because the stable strain with the Biobricks I732011 (mRFP Tripart LVA-) don't to growth. |
| - | + | ||
==Digestion== | ==Digestion== | ||
| Line 13: | Line 11: | ||
[[Team:Paris/Notebook/Protocols#Concentration_of_the_Miniprep | Protocol (it's same that for Miniprep)]] | [[Team:Paris/Notebook/Protocols#Concentration_of_the_Miniprep | Protocol (it's same that for Miniprep)]] | ||
| - | {|border="1" style="text-align: center" | + | => the experiments of Gel Extraction have failed, so we need to repeat the step of digestion. |
| - | |''' | + | |
| - | |''' | + | ===Digestion=== |
| - | |''' | + | [[Team:Paris/Notebook/Protocols#Digestion|Protocol Digestion]] |
| + | |||
| + | {| border="1" style="text-align: center" | ||
| + | |'''Name''' | ||
| + | |'''Template DNA''' | ||
| + | |'''Description''' | ||
| + | |'''Vol MP (µl)''' | ||
| + | |'''Vol H2O (µl)''' | ||
| + | |'''Enzymes''' | ||
|- | |- | ||
|D187 | |D187 | ||
| - | | | + | |MP168.1 |
| - | | | + | | RBS - mRFP - term (FV) |
| + | |9.00 | ||
| + | |15.7 | ||
| + | |EcoRI and XbaI | ||
|} | |} | ||
| - | == | + | ==='''Gel Verification'''=== |
| - | [[Team:Paris/Notebook/Protocols# | + | [[Team:Paris/Notebook/Protocols#Migration after extraction |Protocol]] |
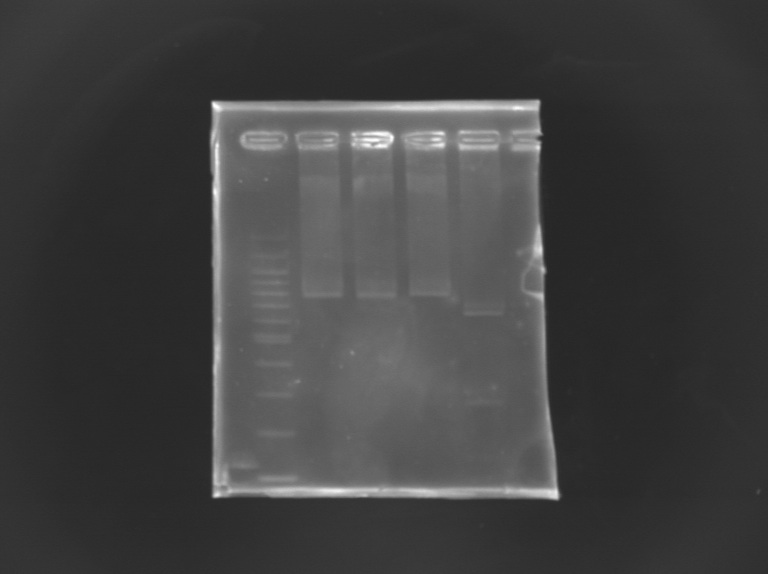
| + | [[Image:KR000246.JPG |thumb |Gel Verification of D187 digestion]] | ||
{|border="1" style="text-align: center" | {|border="1" style="text-align: center" | ||
| - | |''' | + | |'''Well''' |
| - | | | + | |1 |
| - | | | + | |2 |
| - | | | + | |3 |
| - | | | + | |4 |
| + | |5 | ||
| + | |6 | ||
|- | |- | ||
| - | | | + | |'''Sample''' |
| + | |100 pb ladder | ||
| + | |MP168.1 | ||
| + | |no sample | ||
|D187 | |D187 | ||
| + | |colspan="2"|no sample | ||
| + | |- | ||
| + | |'''Expected size (pb)''' | ||
| | | | ||
| - | | | + | |2 955 |
| | | | ||
| + | |2 940 | ||
| + | | colspan="2"| | ||
|- | |- | ||
| - | | | + | |'''Measured size (pb)''' |
| - | + | ||
| | | | ||
| - | | | + | |style="background: #cbff7B"|2 900 |
| - | | | + | | |
| + | |style="background: #cbff7B"|2 900 | ||
| + | |colspan="2"| | ||
|} | |} | ||
| - | |||
| + | =Cloning of EnvZ* in pSB1A2= | ||
==Transformation results== | ==Transformation results== | ||
| Line 66: | Line 87: | ||
|0 | |0 | ||
|0 | |0 | ||
| - | | | + | |9 |
| - | | | + | |6 |
|2 | |2 | ||
|- | |- | ||
| Line 75: | Line 96: | ||
|2 | |2 | ||
|2 | |2 | ||
| + | |- | ||
| + | |'''number of colonies picked up for screening bis''' | ||
| + | |colspan="3"| | ||
| + | |4 | ||
| + | |4 | ||
| + | | | ||
|} | |} | ||
| Line 86: | Line 113: | ||
*positive control: S158 (pSB3K3) | *positive control: S158 (pSB3K3) | ||
*negative control: no template | *negative control: no template | ||
| + | |||
| + | ==Electrophoresis== | ||
| + | |||
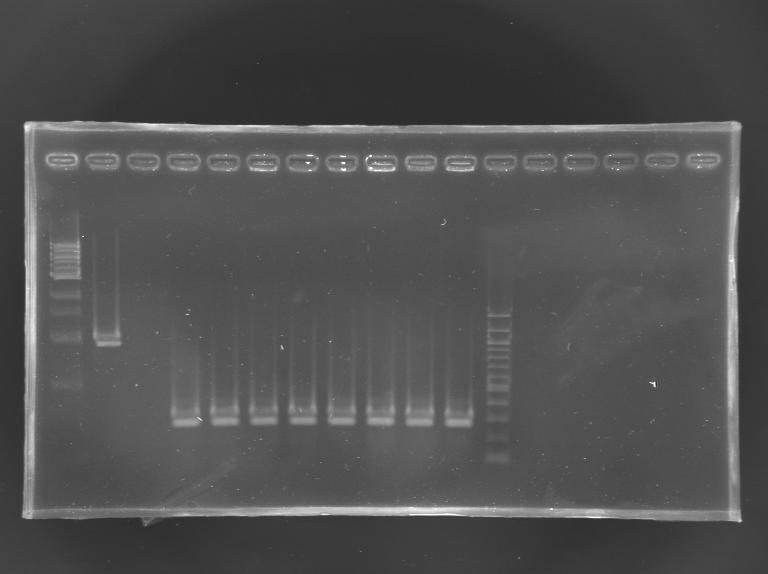
| + | [[Image:Screening EnvZ.JPG|thumb|Screening of the cloning of EnvZ* - clones 1 to 8]] | ||
| + | |||
| + | {|border="1" style="text-align:center" | ||
| + | |'''well n°''' | ||
| + | |1 | ||
| + | |2 | ||
| + | |3 | ||
| + | |4 | ||
| + | |5 | ||
| + | |6 | ||
| + | |7 | ||
| + | |8 | ||
| + | |9 | ||
| + | |10 | ||
| + | |11 | ||
| + | |12 | ||
| + | |- | ||
| + | |'''sample''' | ||
| + | |1 kb DNA ladder | ||
| + | |positive<br>control | ||
| + | |negative<br>control | ||
| + | |L165.1 | ||
| + | |L165.2 | ||
| + | |L165.3 | ||
| + | |L165.4 | ||
| + | |L165.5 | ||
| + | |L165.6 | ||
| + | |L165.7 | ||
| + | |L165.8 | ||
| + | |100 bp DNA ladder | ||
| + | |- | ||
| + | |'''expected size''' | ||
| + | | | ||
| + | | | ||
| + | | | ||
| + | |colspan="8"|1659 bp | ||
| + | | | ||
| + | |- | ||
| + | |style="background: #ff6d73" |'''measured size''' | ||
| + | | | ||
| + | | | ||
| + | | | ||
| + | |colspan="8"|0,3 kb | ||
| + | | | ||
| + | |- | ||
| + | |} | ||
| + | <br> | ||
| + | No correct clone | ||
| + | <br>The 8 other clones were also screened. | ||
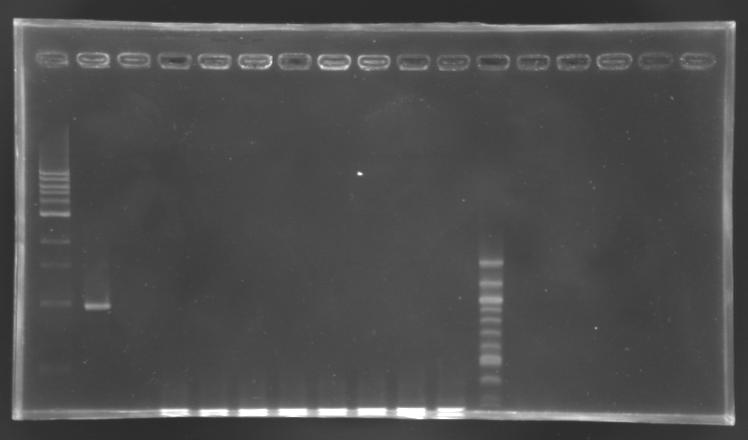
| + | [[Image:KR000256.JPG|thumb|Screening of the cloning of EnvZ* - clones 9 to 16]] | ||
| + | '''PCR''' | ||
| + | elongation time: 2 min 30 | ||
| + | <br> | ||
| + | '''Electrophoresis''' | ||
| + | {|border="1" style="text-align:center" | ||
| + | |'''well n°''' | ||
| + | |1 | ||
| + | |2 | ||
| + | |3 | ||
| + | |4 | ||
| + | |5 | ||
| + | |6 | ||
| + | |7 | ||
| + | |8 | ||
| + | |9 | ||
| + | |10 | ||
| + | |11 | ||
| + | |12 | ||
| + | |- | ||
| + | |'''sample''' | ||
| + | |1 kb DNA ladder | ||
| + | |positive<br>control | ||
| + | |negative<br>control | ||
| + | |L165.9 | ||
| + | |L165.10 | ||
| + | |L165.11 | ||
| + | |L165.12 | ||
| + | |L165.13 | ||
| + | |L165.14 | ||
| + | |L165.15 | ||
| + | |L165.16 | ||
| + | |100 bp DNA ladder | ||
| + | |- | ||
| + | |'''expected size''' | ||
| + | | | ||
| + | | | ||
| + | | | ||
| + | |colspan="8"|1659 bp | ||
| + | | | ||
| + | |- | ||
| + | |style="background: #ff6d73" |'''measured size''' | ||
| + | | | ||
| + | | | ||
| + | | | ||
| + | |colspan="8"|0,3 kb | ||
| + | | | ||
| + | |- | ||
| + | |} | ||
| + | <br> | ||
| + | '''Results''': All the clones analysed were not correct. | ||
| + | |||
| + | |||
| + | =Cloning of OmpR*= | ||
| + | ==Digestion== | ||
| + | ===Determination of the concentration of DNA=== | ||
| + | We used the biophotometer | ||
| + | *5 µL of template DNA or 5 µL of EB buffer for th blank | ||
| + | *55 µL of pure water | ||
| + | {|border="1" style="text-align:center" | ||
| + | |- | ||
| + | |'''Template DNA''' | ||
| + | |'''Concentration of DNA''' | ||
| + | |- | ||
| + | |PCR 147 | ||
| + | |150 µg/mL | ||
| + | |- | ||
| + | |PCR 148 | ||
| + | |101 µg/mL | ||
| + | |- | ||
| + | |MP 101.2 | ||
| + | |353 µg/mL | ||
| + | |} | ||
| + | |||
| + | ===Name of the digestions=== | ||
| + | {|border="1" style="text-align:center" | ||
| + | |- | ||
| + | |'''Name of the digestion''' | ||
| + | |'''Template DNA''' | ||
| + | |'''What's in?''' | ||
| + | |'''Enzymes used''' | ||
| + | |- | ||
| + | |D 188 | ||
| + | |PCR 148 | ||
| + | |OmpR* | ||
| + | |XbaI-PstI | ||
| + | |- | ||
| + | |D 189 | ||
| + | |MP 101.2 | ||
| + | |pSB1A2 | ||
| + | |XbaI-PstI | ||
| + | |} | ||
| + | |||
| + | ===Protocol of digestion=== | ||
| + | |||
| + | * D 188 : 3 µL of PCR 148 | ||
| + | * D 189 : 3 µL of MP 101.2 | ||
| + | * 3µL Buffer (n°2) 10X | ||
| + | * 0.3µL BSA 100X | ||
| + | * 22.7 µL of pure Water | ||
| + | * 1 µL of each enzyme | ||
| + | |||
| + | * Incubate during about 2h30 at 37°C | ||
| + | * 20 minutes at 65°C | ||
| + | |||
| + | ===Cleaning of the digestion products=== | ||
| + | [[Team:Paris/Notebook/Protocols#Purification_.28Kit_Promega.29|Standard protocol.]] | ||
| + | |||
| + | ==Ligation== | ||
| + | ===Determination of the concentration of DNA=== | ||
| + | We used the biophotometer | ||
| + | *10 µL of template DNA or 10 µL of EB buffer for th blank | ||
| + | *50 µL of pure water | ||
| + | |||
| + | {|border="1" style="text-align:center" | ||
| + | |- | ||
| + | |'''Template DNA''' | ||
| + | |'''Concentration of DNA''' | ||
| + | |- | ||
| + | |D 188 | ||
| + | |9 µg/mL | ||
| + | |- | ||
| + | |D 189 | ||
| + | |30 µg/mL | ||
| + | |} | ||
| + | |||
| + | ===Protocol of ligation L171=== | ||
| + | |||
| + | * 2 µL Ligase Buffer 10X | ||
| + | * 1.5 µL D 189 (vector) | ||
| + | * 5 µL D 188 (insert) | ||
| + | * 11.5 µL pure Water (qsp 20 µL) | ||
| + | * 1 µL T4 DNA ligase at 400 000 U/mL concentration | ||
| + | * O/N at 16°C | ||
| + | |||
| + | |||
| + | =Checking mutagenesis FliA= | ||
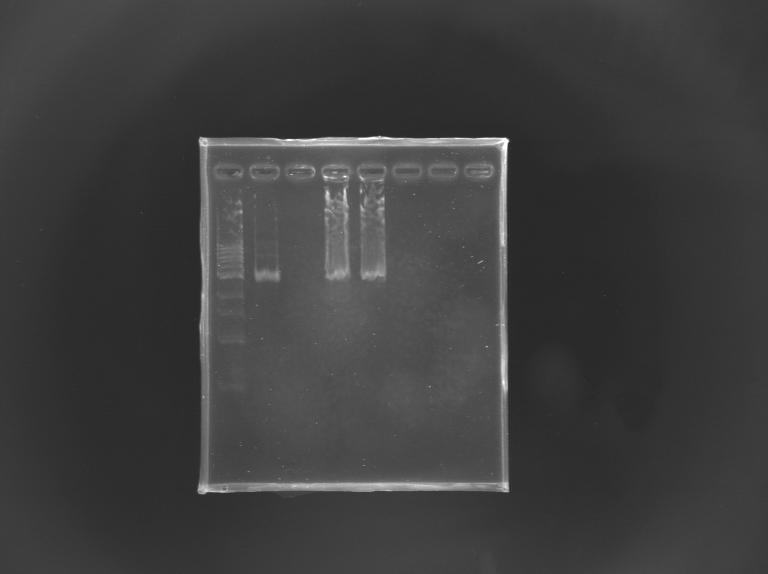
| + | [[Image:Digestion_mut_FliA.jpg|thumb|EcorI/PstI digestion of mutated FliA]] | ||
| + | |||
| + | For this, i digested mutated FliA and non-mutated FliA with EcoRI and PstI and put in migration the digestion products running on gels.<br> | ||
| + | Results : No digestion for the mutated sequence --> successful mission ! | ||
Latest revision as of 19:16, 9 September 2008
Construction of pFlhB - mRFP Tripart (LVA+)Aim : Construction of "pFlhB-RBS-mRFP-dbl ter" (pFlhB-I732078) DigestionMeasurement of the concentration of D187 purifiedProtocol (it's same that for Miniprep) => the experiments of Gel Extraction have failed, so we need to repeat the step of digestion. Digestion
Gel Verification
Cloning of EnvZ* in pSB1A2Transformation results
PCR screening
Electrophoresis
PCR
elongation time: 2 min 30
Results: All the clones analysed were not correct.
Cloning of OmpR*DigestionDetermination of the concentration of DNAWe used the biophotometer
Name of the digestions
Protocol of digestion
Cleaning of the digestion productsLigationDetermination of the concentration of DNAWe used the biophotometer
Protocol of ligation L171
Checking mutagenesis FliAFor this, i digested mutated FliA and non-mutated FliA with EcoRI and PstI and put in migration the digestion products running on gels. Results : No digestion for the mutated sequence --> successful mission ! |
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"