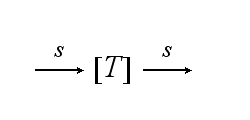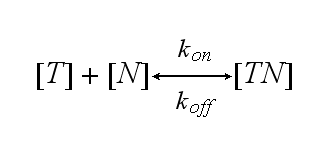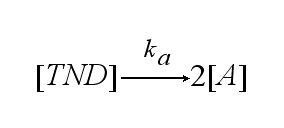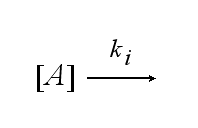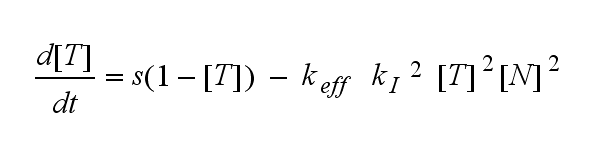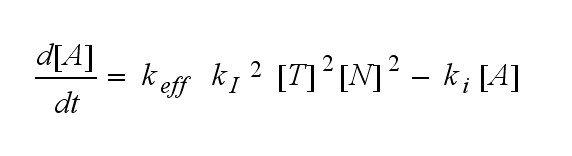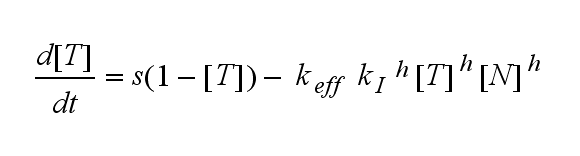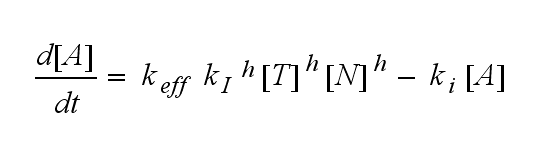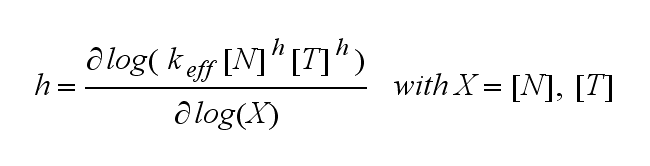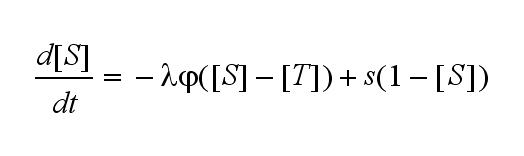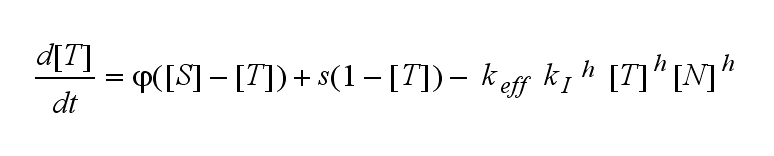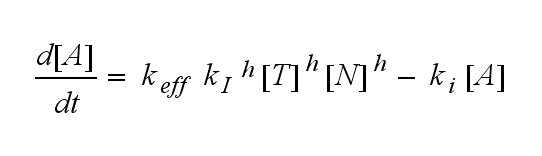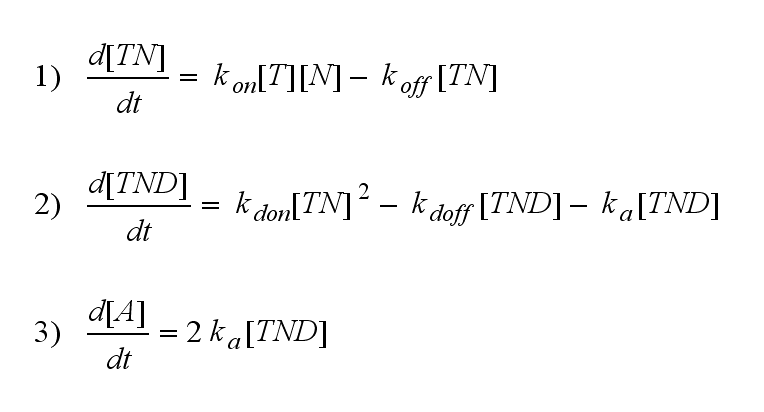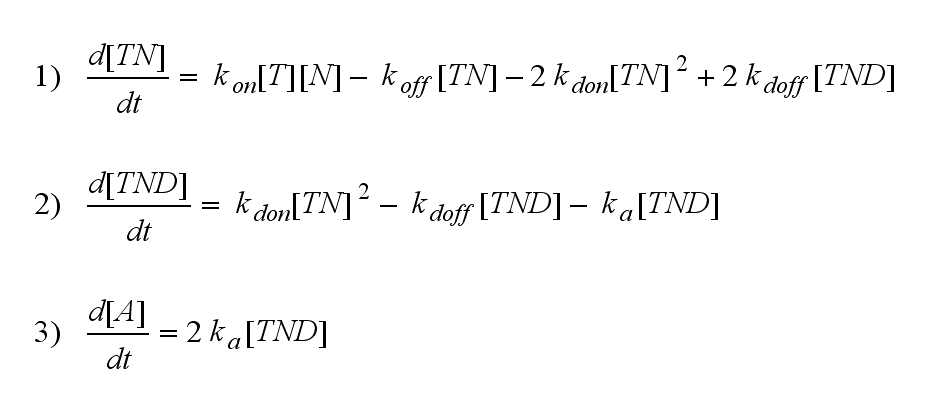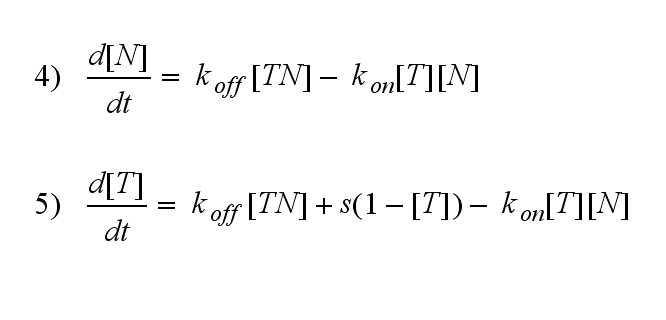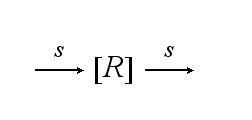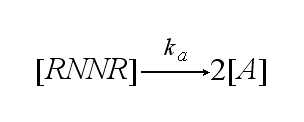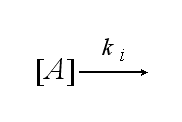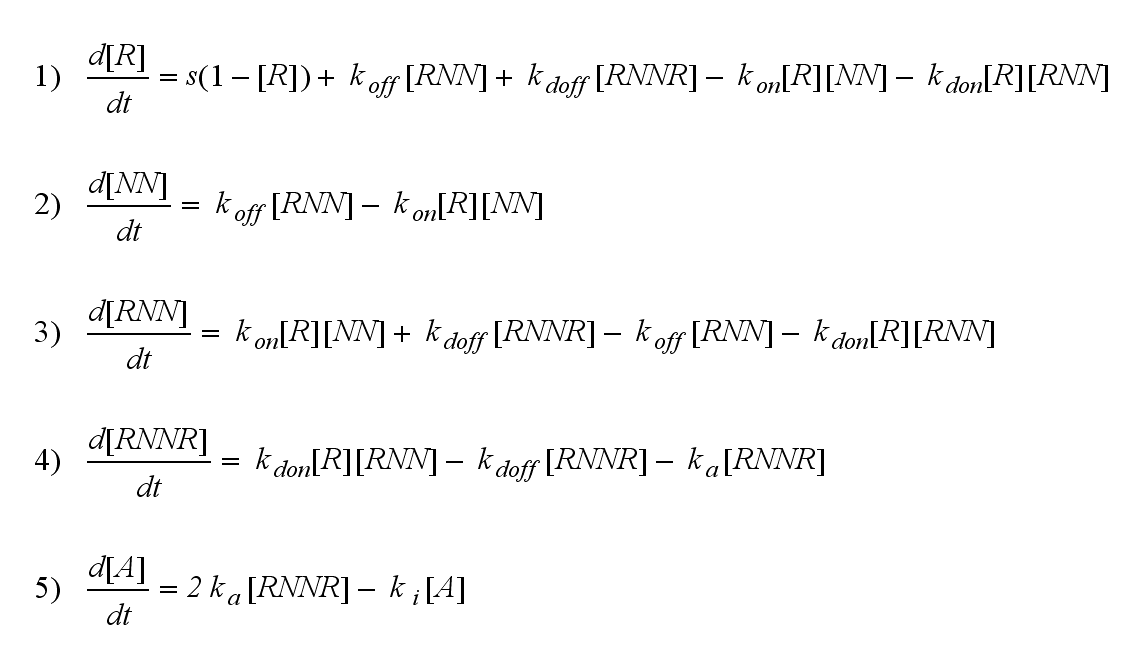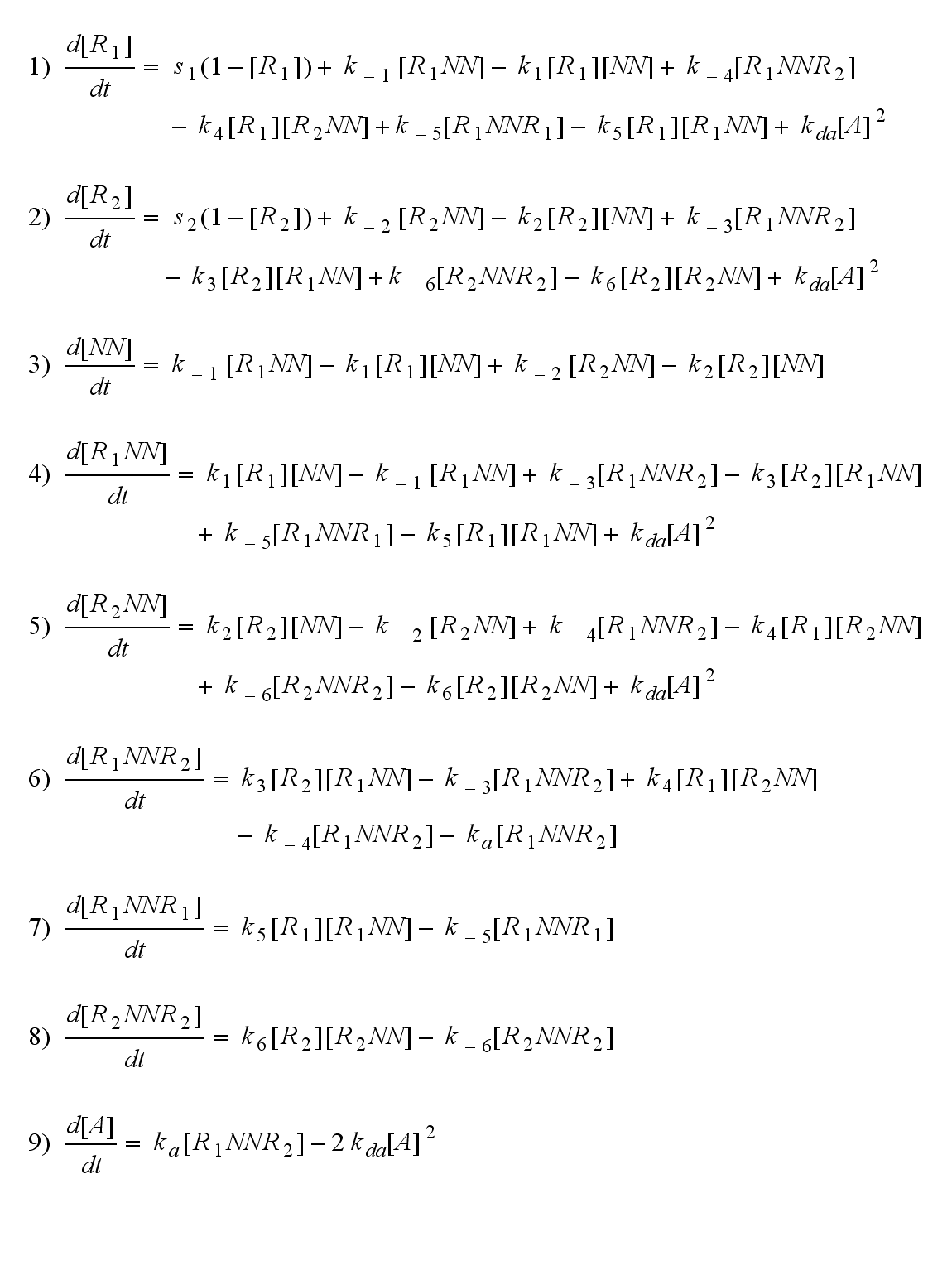Team:Freiburg/Modeling
From 2008.igem.org
| Line 501: | Line 501: | ||
<td> | <td> | ||
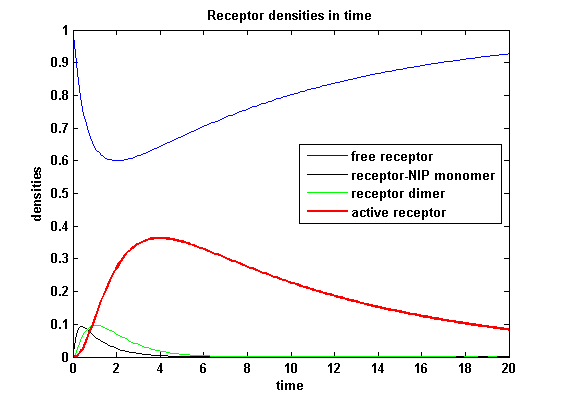
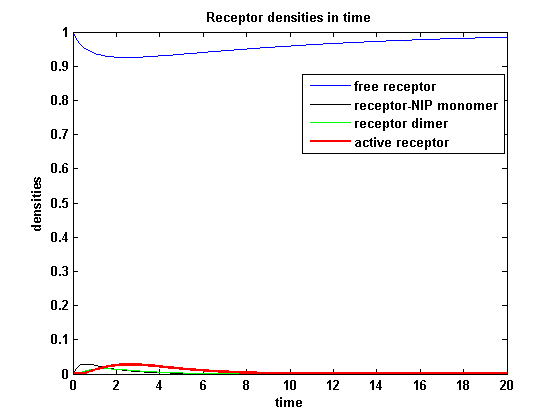
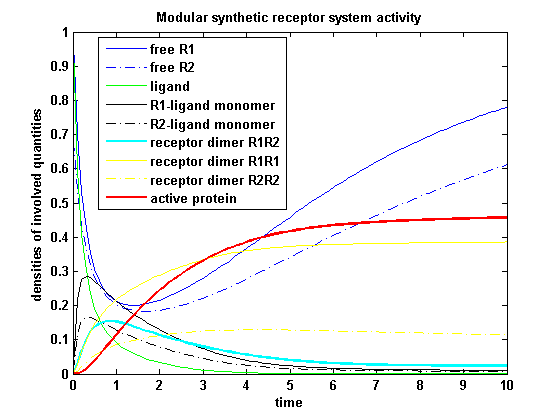
[[image:Freiburg2008_MSRS_2.png|thumb|left|400px|'''Figure 19:''' Involved receptor densities in time]] | [[image:Freiburg2008_MSRS_2.png|thumb|left|400px|'''Figure 19:''' Involved receptor densities in time]] | ||
| - | All densities which are connected to R<sub>2</sub> are shown dashed except the dimer R<sub>1</sub>R<sub>2</sub>, which becomes the active protein. As stated in the parameters, R<sub>1</sub> is a better binding and dimerizing receptor than R<sub>2</sub> and is | + | All densities which are connected to R<sub>2</sub> are shown dashed except the dimer R<sub>1</sub>R<sub>2</sub>, which becomes the active protein. As stated in the parameters, R<sub>1</sub> is a better binding and dimerizing receptor than R<sub>2</sub> and is available in a higher concentration in the cell in the regarded time course, so all densities (free receptor density, monomeric density, dimeric density) connected to R<sub>2</sub> are higher then the according densities of R<sub>1</sub> |
</td> | </td> | ||
<td>chosen parameters:<br> | <td>chosen parameters:<br> | ||
Revision as of 15:35, 28 October 2008
|
Modeling |
_modeling
IntroductionThe dimerization of the extracellular receptor domains is a important necessity for the functionality of our modular receptor system. Presenting the system a stimulus in the form of spatial arranged ligands, the extracellular domains dimerize, thus the corresponding intracellular parts such as the split lactamase halves or split fluorescent proteins complement to measureable output. To analyse the theoretical functionality due to dimerization, first two receptor dimerization models (one T cell receptor model and one general receptor model) are introduced and discussed and then a proper model for the Modular Synthetic Receptor System is constructed.
T cell receptor dimerization model I
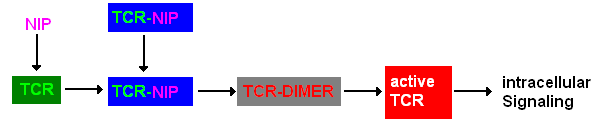
Extracellular signalingA simplified pathway shows the extracellular sequence of TCR activation. After the NIP binding two complexes come together and form a dimer which then leads to activaton of the TCR and further intracellular signaling and T cell activation.
Reaction kinetics
One NIP molecule (N) binds to a TCR (T) with the reaction rate kon or a TCR-NIP complex (TN) dissociates with the reaction rate koff :
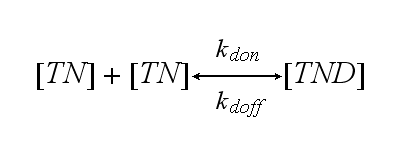
Two TCR-NIP (TN) complexes dimerize to a TCR-NIP dimer (TND) with rate kdon ; the dissociation of a TCR-NIP dimer runs with rate kdoff :
In order to get active TCRs, the TCR-NIP dimer (TND) has to switch into two active TCRs (A) with rate ka :
After activation, the TCR is internalised with rate ki and does not take part anymore in the extracellular signaling :
ODEs derived from the kinetics (Details)In the following equation T represents the free TCR in the T cell membrane where keff is a combination of kdon, kdoff and ka. kI is kon/koff : TCR activity for a set of parametersThe two ODEs above of this first basic model for a set of parameters are solved numerically. They reveal the timecourse of the TCR activity aswell as the one of the unbound TCR.
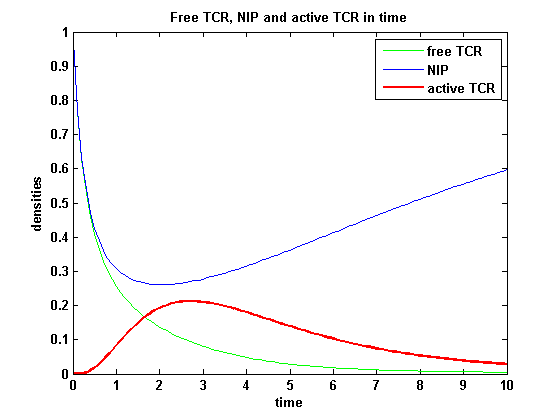
The activity has a maximum a short time after NIP presentation and decreases as the active TCRs are internalized and as the free membrane TCR amount is decreasing. It is remarkable that the response never decreases to complete zero, even for very high internalization rates and in a long timecourse.
Extensions: Ultrasensivity and biphasic kineticsUltrasensitivityThe kinetics of the TCR activation can be generalised by substituting the second order kinetic of the ligand N and the receptor T by a parameter h, which then represents the kinetical order of the system. Biphasic kinetics and parameter analysisNot all TCRs on a T cell membrane can be recruited to NIP binding as a cell´s membrane contains several transmembrane proteins whose size can avoid a TCR-NIP formation when they surround a TCR and make the approaching of the NIP to the binding side of the TCR impossible. Considerating this spatial barriers leads to the idea of a TCR which can switch between two states, one binding state and one non-binding state. Hence a introduction of two different pools of TCRs into the model is appropriate. If a TCR is not available for a NIP molecule, thus it is in the non-binding state,it belongs to the so called spare pool, whereby TCRs belonging to the so called interface pool are in the binding state and can be accessed by the NIP molecule. Moreover the spare pool is in dynamical exchange with the interface pool, so non-binding TCR can become binding TCRs. This exchange is regulated through the parameter λ, a ratio between the spare and the interface pool and φ, the exchange rate constant. S is the spare pool TCR density and T the TCR density of the interface pool. A represents the active TCR density. So the full model I equations are: TCR activity dependent on exchange rate φ and ratio λ :
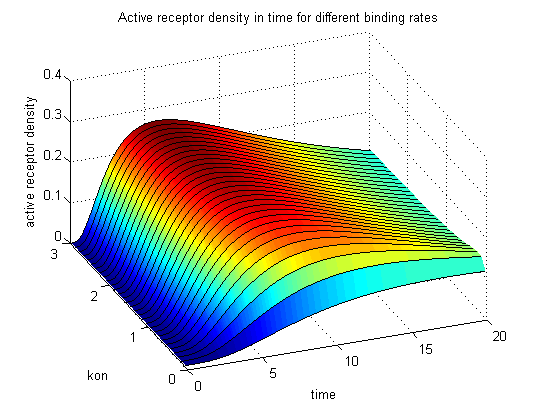
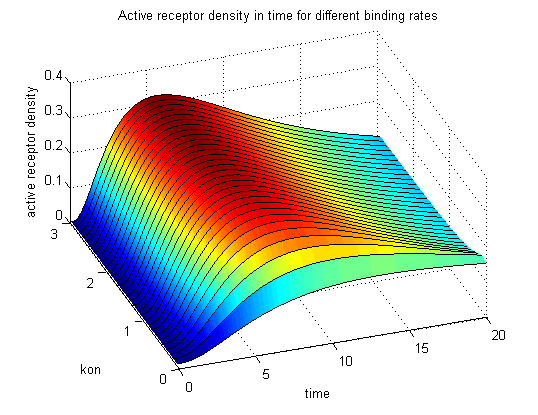
The x-axis represents the timecourse of the activity, the y-axis represents both parameters φ ( = y) and λ ( = 2 - y). So each black line in the plot is a timecourse of the TCR activity for a different φ and λ. The z-axis is the response intensity. With increasing exchange between the interface and spare pool, more TCRs switch to the binding state, hence more TCRs can bind NIP. As a consequence the active TCR density is higher then for a low exchange. Correctional termsRegarding the reaction kinetics and considering the ODEs as a model for TCR dimerization (Bachmann, 1999) led to the realization of an error in the mentioned publication. The ODEs evolved from the reaction kinetics are not :
but :
Furthermore can be derived for the NIP density and the free TCR:
After NIP addition the TCR activity rises to a maximum and decreases as active receptors are internalized and the NIP amount is used up. The free receptor density recovers due to permanent expression of new receptors which are build into the membrane.
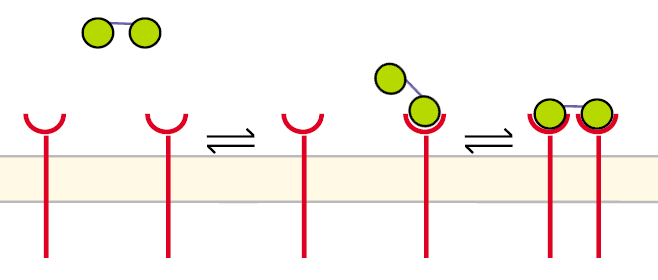
Receptor dimerization model IIA receptor dimerization can proceed in a different way aswell, especially when two NIP molecules are linked to a structure like our DNA-Origami: Before the first step the receptors are unbound and diffuse in the cell´s membrane. The DNA-Origami binds to a receptor in the way that only one NIP molecule is connecting to one receptor. In the second step the second NIP molecule binds to the second receptor and both receptors dimerize.
Reaction kineticsThe continuous production of the cell´s receptor R with rate s is:
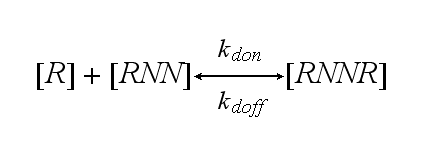
One receptor R binds to one of two NIP molecules NN of the DNA-Origami with rate kon or a receptor-NIP complex RNN dissociates with koff:
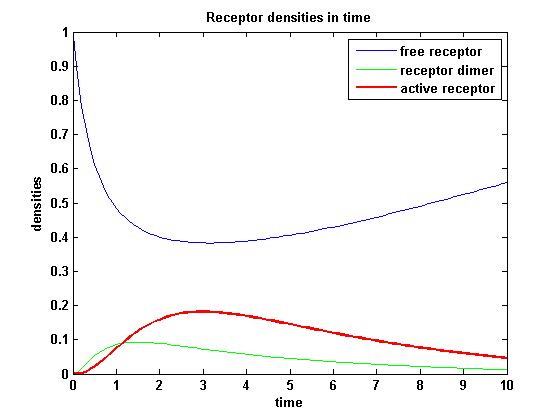
ODE derivationThe ODEs can be derived from the reaction kinetics again and are: Extracellular receptor activityFor a set of parameters the ODEs are solved and reveal amongst the other quantities the receptor densities in the time course:
The receptor activity rises after NIP addition, has a maximum and decreases due to the internalization of the active receptor. The dimer density increases fast after addition of NIP and decreases as dimers fade into active receptors.
Discussion of model I and IIFor the comparison, the corrected model I is taken and model II. The receptor NIP binding mechanism differs in the two models. Model I assumes the production of two independent receptor-NIP complexes which can dimerize after formation. In model II, dimerization happens after one of two receptors connects to the bivalent ligand consisting of two NIP molecules. Setting several conditions unravels the consequences of this differences for the active receptor, the dimer and the receptor recovery. Different binding rates, dimerization rates and NIP amounts are used in the solutions of the model I ODEs and model II ODEs. receptor densities dependent on binding rate konThe two model ODEs are solved for the mentioned parameters below whereby the binding rate kon was variated.
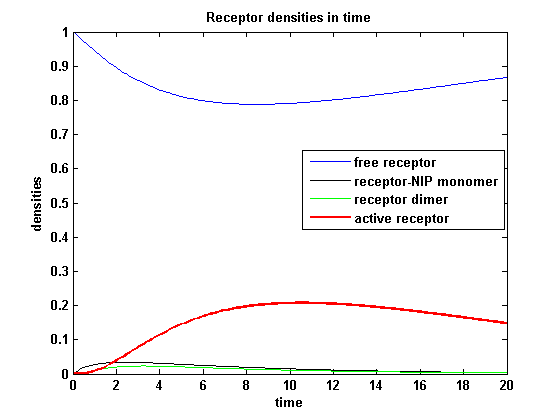
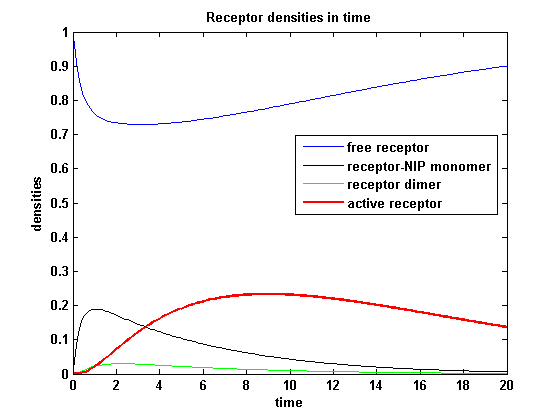
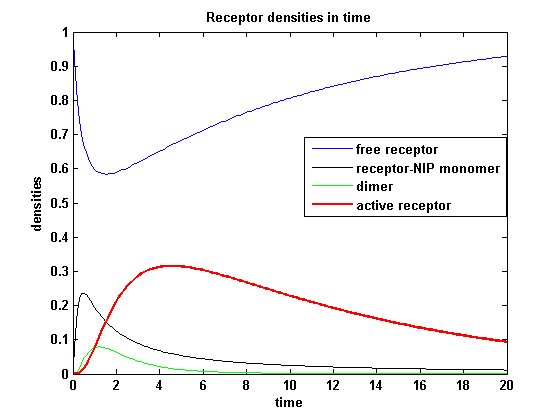
So the course in time for the free receptor, the monomeric receptor-NIP complex, the dimer and the active receptor is plotted:
kon = 0.2 :
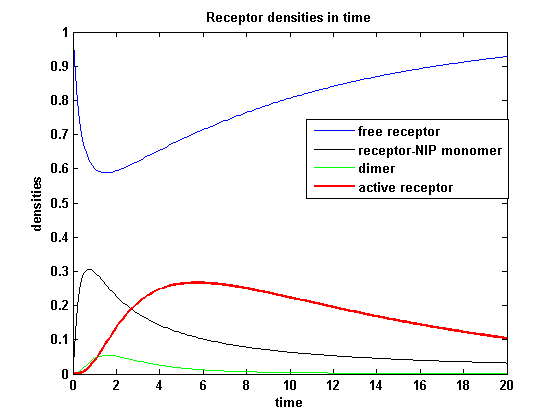
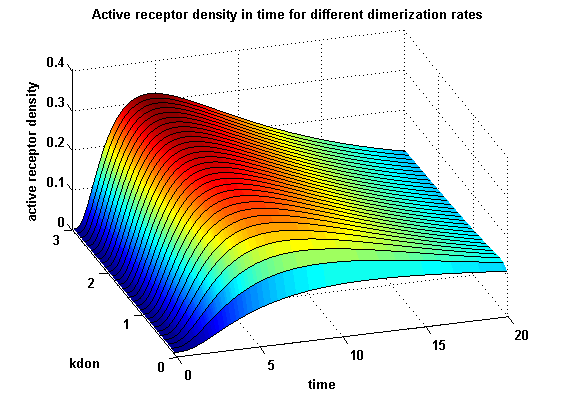
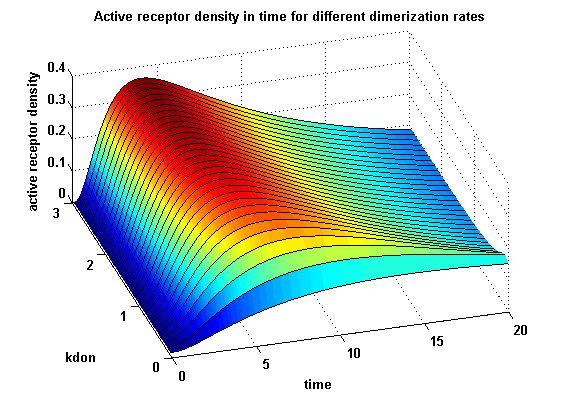
receptor densities dependent on dimerization rate kdonApplying different kdon leads to the following result:
kdon = 0.3 :
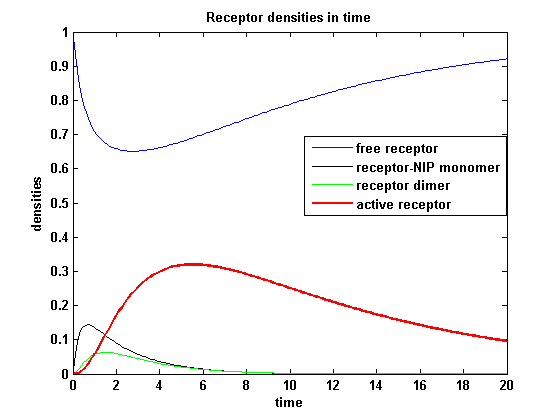
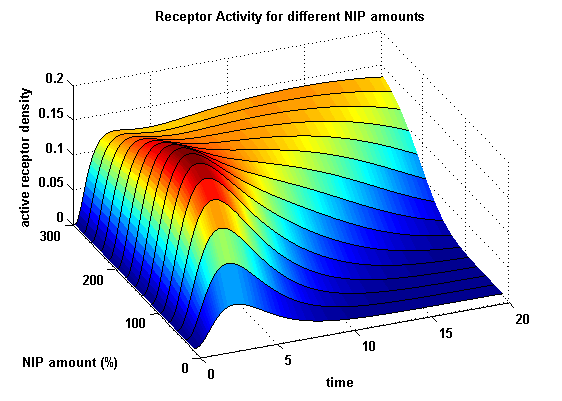
receptor densities dependent on NIP AmountThe use of different amounts of ligand in the form of NIP is maybe the most important influence on the receptor activation. As the ligand is the stimulus in order to activate our system, it´s amount will determine strongly the activity of the receptor. So different amounts of NIP are used and a higher internalization rate ki is used aswell.
NIP = 0.1 :
Modular Synthetic Receptor System modelThe dimerization model II is used to form the base of the model for our Modular Synthetic Receptor System. It has two receptors R1 and R2. The bivalent ligand are two NIP molecules NN which are linked to DNA-Origami. Aswell two linked lipocalin molecules can serve as the bivalent ligand. Each receptor consists of a extracellular detector domain, which is connected to an intracellular split protein half by a short transmembrane protein. Only when receptor R1 and R2 dimerize, the intracellular split protein halves complement to become the complete functional protein A in the form of a β-lactamase or a fluorescent protein. A dimerization of R1 and R1 or R2 and R2 can happen to R1R1 or R2R2, but does not lead to an active intracellular protein and measurable output. Of course all receptor dimers (R1R2, R1R1, R2R2) dissociate into monomers much easier than an receptor dimer which is active in the form of A, as the connected split halves proteins halves attract each other.
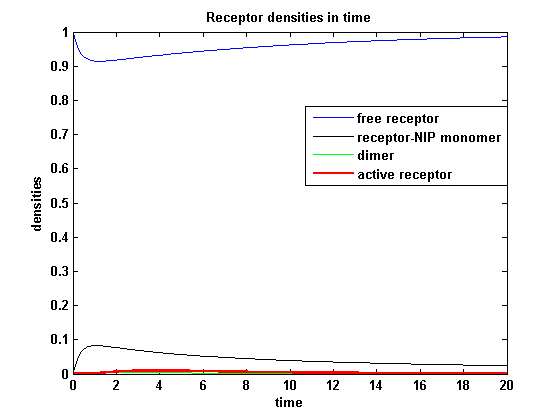
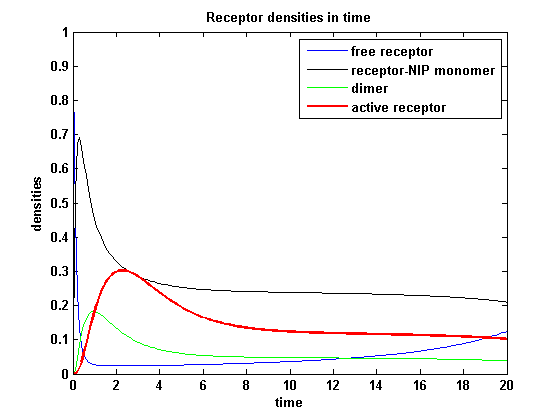
Reaction kineticsODE derivationThe ODEs can be derived from the reaction kinetics again and are: Involved quantities and split protein activityThe ODEs are solved for a set of parameters and reveal the time courses of the quantities:
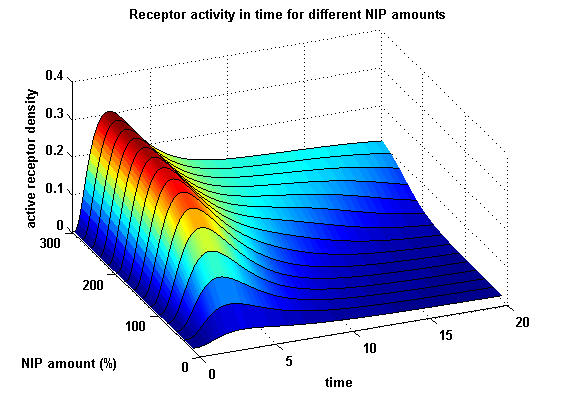
Active split protein density dependent on input ligand amountIn order to find the activity behaviour dependent on the input, the same parameters were used as above except the initial ligand parameter, which was variated.
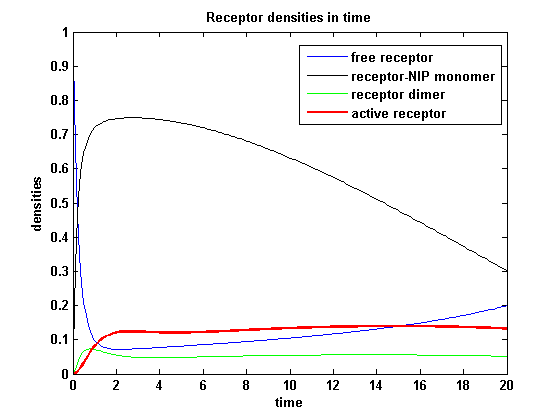
Activity with two kinetical different receptorsAs two different receptors are used, they can differ in the kinetics. Assuming that receptor R1 is binding the ligand better, so its binding rate is higher and its dissociating rate is lower than of receptor R2. Choosing not only a higher turnover rate s1 but also a higher dimerization and a lower dimer dissociating rate for receptor R1 and a lower initial free receptor density of R2, gives the following theoretical result:
Matlab file listLiterature |
 "
"