Team:University of Lethbridge/Project
From 2008.igem.org
The “Bacuum" Cleaner – an intelligent self-propelling keener cleaner
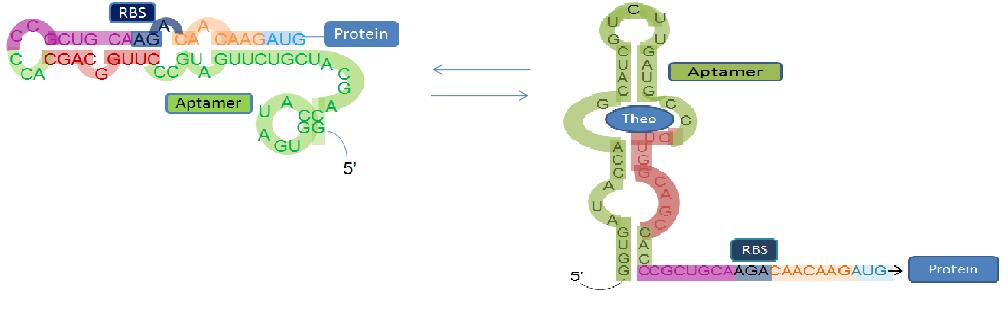
Our goal is to create a modified Escherichia coli bacterium capable of seeking out and degrading toxic aromatic pollutants created during the oil refinery and mining processes. Our “Bacuum" cleaner will respond to this destructive compound through interaction with a programmed riboswitch (an mRNA capable of binding a target molecule with its 5’ UTR which causes a conformational change, thereby regulating any downstream protein translation). By using riboswitches that switch at varying concentrations of target ligand, we can alter the signal which is induced. At low concentrations, we intend to have our riboswitch express the motility protein cheZ in E. coli, thus directing the bacterium towards higher concentrations of our target molecule (i.e. positive chemotaxis). Once it reaches a threshold concentration, a catabolic pathway capable of degrading our target pollutant will be activated.
We plan to use [http://en.wikipedia.org/wiki/Systematic_Evolution_of_Ligands_by_Exponential_Enrichment SELEX] to reprogram the theophylline riboswitch initially characterized by our team last year, aiming to have it bind one of many toxic aromatic compounds. This not only builds on our past work with riboswitches but also demonstrates an alternative function for riboswitch-controlled chemotaxis, changing from a simple detection to active “search and destroy” role. Fluorescent protein expression will be used to demonstrate and characterize the functionality of the riboswitch.
Our project also has major environmental implications, especially in Alberta, where the oil industry is the driving economic sector. The tailings ponds used to house discarded mining refuse from the oil refineries pose a major environmental dilemma. The problem arises with being unable to collect and degrade the compounds which make up these toxic soups. These not only toxic but often corrosive water beds not only affect the local environment but can also cause severe repercussions to other ecosystems as much of the wildlife relies on getting their food and water from the now disrupted bionetwork. These tailings ponds are often laced with heavy metals and dangerous chemicals, including these toxic aromatic hydrocarbons which can be extremely difficult to breakdown. Despite the efforts of the corporations responsible, some of these chemicals can leech into and contaminate the surrounding ground water and soil. It is our intention that our “Bacuum” will not only remove these compounds from such an aqueous system, but will also eliminate them. This goes one step further than the ordinary less-intelligent bag-dependent vacuum cleaners. Ultimately we aim to have our bacteria convert the target aromatic into a useable compound. One example would be the biosynthesis of fatty acids as an alternative fuel source, creating a potential “Bioreactor” which is capable of creating a more environmentally friendly fuel source.
Project Details
Chemotaxis - the "search" from "search and destroy"
Our subproject is to ensure our 'bacuum cleaner' will search out and move towards our potential target. Using a riboswitch which responds to this target, a gene involved in motility in Escherichia coli (CheZ) will be switched to an 'on' state. CheZ controls bacterial movement by dephosphorylating CheY, altering the bacteria from random tumbling to directed movement. By doing so, we hope to observe the bacteria moving towards the target molecule in a positive chemotaxis manner.
Because we wish to eventually have our bacteria degrade this activating compound, we need to have a multi-level response system established. Our bacteria should respond to low levels of the target by activating our CheZ riboswitch (through a strong binding aptamer) and to high levels by activating the catabolic pathway responsible for its degradation. By doing so, we will have created our 'search and destroy' system.
Our subgroup will attempt to establish a working motility assay to prove control of chemotaxis is possible with an already characterized theophylline-binding riboswitch and later with our new target molecule. We will also use a colour read-out system of two fluorescent proteins to demonstrate the differential binding of the ligand at varying concentrations.
Riboswitch Characterization
This subproject was created to quantify and characterize the binding affinity of the theophylline riboswitch to theophylline as well as other similar aromatic ligands. We will use Green Fluorescent Protein (GFP) sequenced behind the riboswitch to help in this endeavour. When the target ligand is bound to the riboswitch, the production of GFP will be induced, producing green colonies. When the ligand is not bound, GFP will not be produced. We intend to try to characterize the riboswitch with and without the iGEM scar to determine if this produces any effect on the relative effeciency of the riboswitch on the protein.
We will use variable concentrations of Theophylline, Caffeine and 3-methylxanthine to help with the characterization of the initial theophylline riboswitch. We will also test other aromatic compounds to determine wether or not, the theophylline riboswitch can be utilized to bind them as well.
Once we have successfully produced mutant riboswitches through SELEX, will we then perform these same characterization experiments to determine their binding affinities.
SELEX
IRES
Internal ribosomal entry sites (IRESs) are elements found in certain eukaryotic mRNAs that facilitate translation initiation by noncanonical, end independent interactions known as internal ribosomal entry. We are going to incorporate the IRES from the rpsA gene, encoding ribosomal protein S1, of E. coli into our Bacuum cleaner as a means of quantifying translational efficiency. In order to do this we will compare translation rates for GFP to other canonical RBS. This will allow us to standardize the number of ribosomal initiations per second (RiPS).
Results
 "
"

 ]
]
 ]
]
 ]
]
 ]
]
 ]
]
 ]
]