Team:BCCS-Bristol/Modeling
From 2008.igem.org
Revision as of 21:54, 12 August 2008 by Tgorochowski (Talk | contribs)
|
Approach
Due to the nature of this project a wide range of modelling techniques are required to help understand the validity and guide experimentation in the wet lab. The reason for this is that in addition to the creation of synthetic genetic circuits to regulate the internal state of a cell, we require the cell to physically interact with the environment in such a way that by bringing a large number of bacteria following to same rules emergent behaviour is exhibited.
Models
 |  |  |
| | | |
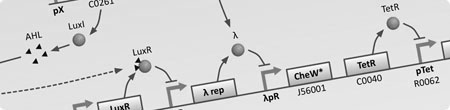
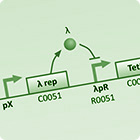
| To understand the dynamics of genetic networks that we design, differental equations are used to model the changes in concentration of mRNA and proteins. | With the project requiring global behaviour to emerge via the physical en masse movement of bacteria, stochastic agent based simulations are used to investigate the viability and limits of different approaches. | With the aim of improving simulation accuracy this model integrates the intercellular GRN dynamics with the extra-cellular interactions of the stochastic agent based simulation. |
Modelling Notebook
- Progress Report - To track our progress see our weekly reports. These let you know what we have been up to, including comments on the reasons behind any important decisions that were made.
- Modelling Parameters - Full listings of all parameters investigated for the modelling part of the project, with values that have been chosen, reasons for their selection and references to the original sources.
Results
 "
"