Team:Beijing Normal/Project
From 2008.igem.org

| Home | The Team | The Project | Parts Submitted to the Registry | Modeling | Notebook |
|---|
Contents |
Overall
We are aiming to create some magic intelligent bacteria to track and ‘eat’ pollutants PCBs (Polychlorinated Biphenyl) and dioxins efficiently, based on the methods of synthetic biology.
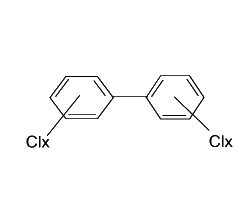
Polychlorinated biphenyls (PCBs) are a family of compounds produced commercially by the direct chlorination of biphenyl using ferric chloride and/or iodine as the ctalyst.The total amount of PCBs produced in the world is estimated 1.2 million tons.Because PCBs have been released into the environment in many countries over decades, these compounds have become serious and global environmental contaminants.PCBs tend to accumulate in biota owing to their lypophilic property.
The biphenyl molecule is made up of two connected rings of six carbon atoms each, and a PCB is any molecule having multiple chlorines attached to the biphenyl nucleus.
Two distinct classes of bacteria have now been identified that biodegrade PCBs by different mechanisms, including aerobic bacteria which live in oxygenated environments and anaerobic bacteria which live in oxygen free environments such as aquatic sediments. The aerobes attack PCBs oxidatively , breaking open the carbon ring and destroying the compounds. Anaerobes, on the other hand, leave the biphenyl rings intact while removing the chlorines.
Polychlorinated dibenzo-p-dioxins (PCDD) and polychlorinated dibenzofurans (PCDF) were introduced into the biosphere on a large scale as by-products from the manufacture of chlorinated phenols and the incineration of wastes. Due to their high toxicity they have been the subject of great public and scientific scrutiny.
The evidence in the literature suggests that PCDD/F compounds are subject to biodegradation in the environment as part of the natural chlorine cycle. Lower chlorinated dioxins can be degraded by aerobic bacteria from the genera of Sphingomonas, Pseudomonas and Burkholderia.However, higher chlorinated dioxins requires anaerobic degradation process.
Organic pollutants such as PCB and dioxins, produced in human beings activities in the last century, are toxic and carcinogenic which are able to promulgate widely and accumulate to a high level of concentration by food chain. Due to their inherent thermal and chemical stability, it is commonly considered as indestructible under normal incineration or burial.
Nonetheless, by endowing some bacteria ability of utilizing such molecules as carbon source, cooperative evolution makes all possible! Enzymes assembled from related degradation pathways into our host strain serve as the function part. We introduce popular components involved in chemotaxis, quorum-sensing to regulatory parts, sense the environment signal, respond to move, accelerate growing and produce related degradation enzymes. After the cleaning work being finished, bacteria will return to the normal state.
Taking the condition of our lab into account, we decide just deal with the aerobic degradation path way. And do some work on increasing the degradation effeciency.
to be continued and refreshed...
Project Details
background information
- Although many environmental pollutants are efficiently degraded by microorganisms, others such as PCBs and dioxins persist and constitute a severe health hazard. In some instances, persistence is a consequence of the inadequate catabolic potential of the available microorganisms. Gene technology, combined with a solid knowledge of catabolic pathways and microbial physiology, enables the experimental evolution of new or improved catabolic activities for such pollutants.
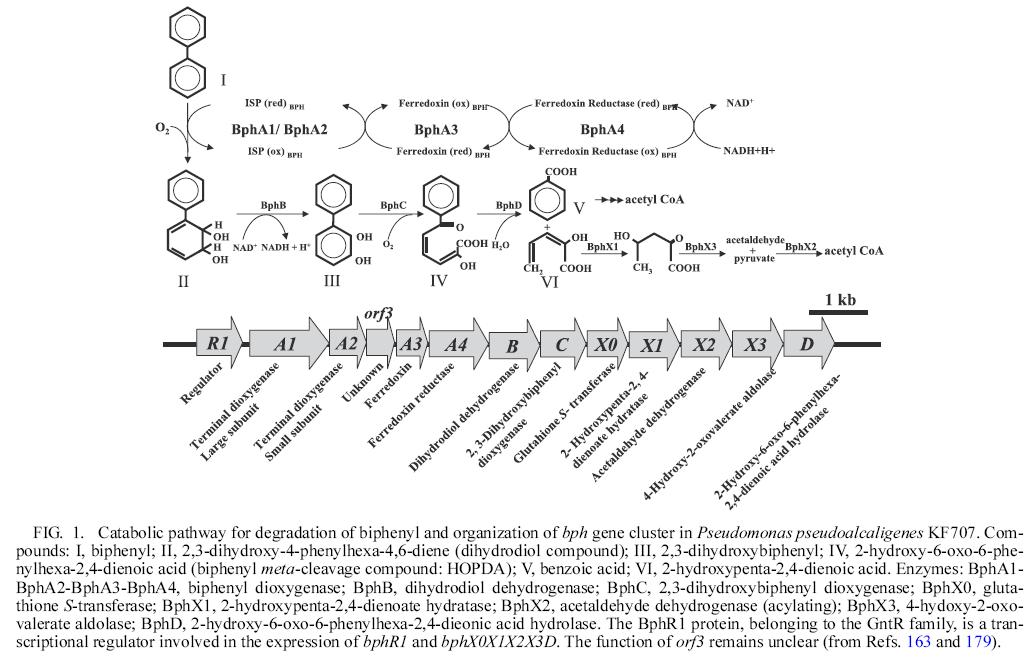
- Catabolic pathway for degradation of biphenyl and organization of bph gene cluster is as follows:
reference: Microbial Degradation of Polychlorinated Biphenyls: Biochemical and Molecular Features.Kensuke Furukawa and Hidehiko Fujihara,Journal of bioscience and bioengineering,Vol. 105, No. 5, 433–449. 2008
permitted to use this figer by professor Kensuke Furukawa
Enzymes responsible for oxidative degradation of PCBs
1 BphA: biphenyl dioxygenase
Biphenyl dioxygenase is a Riesketype three-component enzyme, comprising a terminal dioxygenase that is composed of a large subunit (encoded by bphA1) and a small subunit (encoded by bphA2), a ferredoxin (encoded by bphA3) and a ferredoxin reductase (encodedby bphA4)
It catalyzed the conversion of biphenyl to dihydrodiol compound in step 1.
2 BphB: dihydriol dehydrogenase
In the second step of the upper pathway BphB enzymes catalyzes the conversion of dihidrodiol to dihydroxy compound.
3 BphC: 2,3-dihydroxybiphnyl dioxygenase
The third enzyme in upper pathway is a 2,3-dihyroxybiphnyl dioxygenase involved in the ring meta-cleavage at the 1,2 position.
4 BphD: hydrolase
In the forth step BphD, a hydrolase, hydrolyzes the ring meta-cleavage yellow compound(HOPDA) to chlorobenzoic acid and 2-hydroxypenta-2,4-dienoate.
The Experiments
- 1.Get the parts from biobrick
We largely follow the instructions provided by the webpage, however, more TE buffer is added(10ul). It seems that this will increase the amount of the plasmids dissolved and improve transformation effeciency.
- 2 PCR
- 2.1 The pfu/Taq complex system
Reagent Concentration/Activity 50ul in total taq buffer 10x 5 pfu/taq complex 0.8~1.0 dNTPmix 10mM each 4 Primer 1 10uM 1.5 Primer 2 10um 1.5 Template DNA changeable -- ddH2O --- add to 50ul
- 2.2 The program under pfu/Tag complex system
Progress Program I
Predenaturing 95℃ 5 min Denaturing 95℃ 30sec Annealing (Tm-5) ℃ 30sec Extension 72℃ theoretically 1min/1kb Last extention 72℃ 5min Hold 4℃
- 3 restriction enzyme digestion
select suitable enzymes and buffer. Analyse the system using NEB cutter in case of double digestion. [http://www.neb.com/nebecomm/DoubleDigestCalculator.asp NEB cutter finder]. 37℃ water bath for 2~3h, 4h is preferable
- 4 ligation
The key to a successful ligation according to our experience is avoiding high temperature. After a gel extraction with 32ul elution buffer, a approximately 20ul product is obtained. Then mixed with 20ul Buffer 1(Takara Ligation Kit). Reaction at 16℃ water bath is widely recommended, and time for ligation we use is 2h or more.
- 5 transformation
Add 10ul ligation product into 100ul competence cells(Top10 or others) and keep it in 4℃ refrigeratory for 30min. After a hotshock of 45sec(for chemical competent) or 90sec(for CaCl2 competent), the cells are placed in the 4℃ refrigeratory again for 3-5min. Then add 600~800ul SOC to hotshock cells and incubate in 37℃ shaking table for 1h(160rpm-180rpm). At last the cells are palced on petri dishes with relative antibiotics at appropriate concentration. Then place in 37℃(temperature accords to the properties of different hosts and plasmids),incubate for 12h or more.
Special protocol
To obtain some genes, we often have to use bacteria chromosome or large size plasmid as template in which high GC% content and complex secondary structures are seriously hampering PCR and thus leading to complete failure. To solve this problem, we have developed a special protocol-- hot start method combined with additive(sole or mixed)-- which is most helpful .
- Simplified Hot Start PCR
Hot start method is to prevent primer dimer and low specified product yield. In our experiment, We use common taq polymerase to replace commercial high-priced hot-start polymerase.
After a 5min 95℃ pre-heat step, DNA polymerase is added to each PCR tube before the PCR cycling begin. After that, the regular PCR cycling could begin.
- explaination
When reaction components are mixed at room temperature, reaction set up below the optimal primer annealing temperature, which permits nonspecific primer annealing and extention.Undesired, non-specific primer extetion products formed this way may be amplified in the PCR, resulting in misprimed products and primer ologomers.In hot start PCR, DNA polymerase is withheld from the mixture until the system has reached a temperature that favours specific primer annealing. As a result hot start PCR can greatly improve specificity, sensitivity and yield in a PCR.
- effective additives
The most simple additive in our PCR is DMSO(>=99.9% purity) at a concentration of 5%(V/V). It works well in most 'problematic' PCR, however, when DMSO fails in some case, we have to turn to a magic mixed additive (2.7 M betaine, 6.7 mM DTT, 6.7% DMSO, and 55 ug/ml BSA) for help. This excellent additive has solved several PCR where even 5% DMSO fails.
In certain PCR there is no yield without this additives, for example the ones by which we amplify bphA1A2A3A4 and bphBC.
- explaination
One major factor limiting the output of PCR routines is that a number of DNA sequences are poorly or not amplifiable under standard reaction conditions, either because of their high GC-content or/and their instrict propertoties to form secondary structures. This protocol is intended for GC-rich DNA sequences. This is a concentration dependent combination of betaine, dithiothreitol, and dimethyl suloxide. According to the references the concentration ratio is: a mixture containing 2.7 M betaine, 6.7 mM DTT, 6.7% DMSO, and 55 ug/ml BSA. It is stable at -20℃ for at least 3 months.
- reference:
1.Simplified hot start PCR, David E. Birch et al. Nature 381:445-446 (1996)
2.An alternative hot start technique for PCR in small volumes using beads of wax-embedded reaction components dried in trehalose, S.Kaijalainen et al. Nucleic Acids Research 21:2959-2960 (1993)
3.Effect of DMSO on PCR of Porphyra yezoensis(Rhodophyta)gene, Yukihiro Kitade et al. Journal of Applied Phycology 15:555-557 (2003)
4.An efficient and economic enhancer mix for PCR, Markus Ralser et al. Biochemical and Biophysical Research Communications 347:747–751 (2006)
 "
"