Team:NYMU-Taipei/Project/Time Regulation/Reloxilator
From 2008.igem.org
Blackrabbit (Talk | contribs) (→Experimental Results) |
Blackrabbit (Talk | contribs) (→Conclusion) |
||
| Line 387: | Line 387: | ||
| rowspan="2" | '''Initial concentration''' of: || HtlB (higher) || rowspan="2" | (longer) startup time | | rowspan="2" | '''Initial concentration''' of: || HtlB (higher) || rowspan="2" | (longer) startup time | ||
|- | |- | ||
| - | | CII (higher) | + | | CII (higher) |
|- | |- | ||
| rowspan="2" | '''Natural degradation rate''' of || HtlB (longer) || rowspan="2" | (longer) time period | | rowspan="2" | '''Natural degradation rate''' of || HtlB (longer) || rowspan="2" | (longer) time period | ||
| Line 400: | Line 400: | ||
Is a promoter that can be activated by HtlB? because then our calculations would be even more accurate. | Is a promoter that can be activated by HtlB? because then our calculations would be even more accurate. | ||
| - | |||
| - | |||
== Experimental Results == | == Experimental Results == | ||
Revision as of 23:53, 29 October 2008
| Home | Project Overview: | pH Sensor | Attachment | Time Regulation | Waste Removal | Experiments and Parts | About Us |
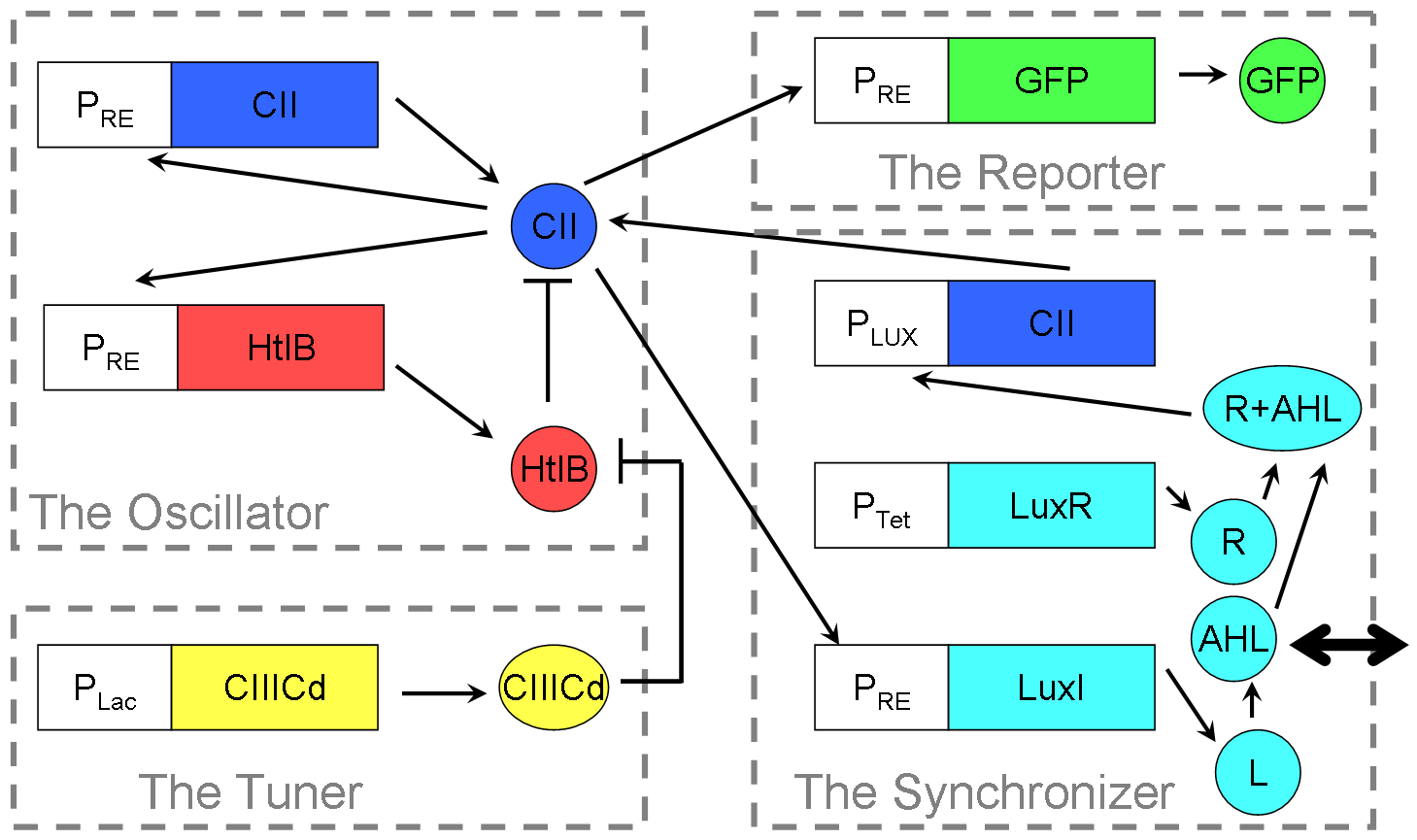
Our Reloxilator (Relaxation Oscillator): A four-part system consisting of an oscillator, a synchronizer, a tuner, and a reporter.
Contents |
Brief Description
The oscillator is a two-component [http://en.wikipedia.org/wiki/Relaxation_Oscillator relaxation oscillator]. The first component consists of the PRE promoter and the gene CII in a positive feedback loop, and both originating from from the λ phage. The second component uses the HtlB gene from E. coli attached to the same promoter as the first component. The HtlB gene produces HtlB proteins that degrade CII proteins.
The synchronizer allows near-full synchronization between different cells after about two time periods of oscillation1. The main parts of the synchronizer is made up of the PLux promoter, and the LuxI and LuxR genes, all coming from the Vibrio Fischeri organism. PRE and the CII gene are the same as the parts used in the oscillator, tying the oscillator and synchronizer together.
The Tuner is a way to control the time period of the system by inhibiting the rate of degradation HtlB has on CII in the oscillator. The protein used is the CIIICd, a DNA synthesised version of the CIIIC protein from the study done by Halder et al2.
The Reporter is the promoter PRE (the same part used in the oscillator) attached to a green flurorescent protein gene. It is used in various tests to check many of the components in the system.
System Description
This section houses the description of our system and the we use.
The Oscillator
A two-component oscillator consisting of two genes: one that induces both genes, while the second gene represses the first one. This oscillation is much like the oscillation of a [http://en.wikipedia.org/wiki/Relaxation_Oscillator capacitor charging and discharging].
Components
Related Biobricks
- [http://partsregistry.org/Part:BBa_R0053 BBa_R0053] works but comes from a different phage (P22). Therefore using it means no proof of oscillation with HtlB [citation required]. Just realised it doesn't mean we can't try it.
- [http://partsregistry.org/Part:BBa_I7108 BBa_I7108] uses BBa_R0053 and produces little GFP output. But that may be good for us?
- [http://partsregistry.org/Part:BBa_R1053 BBa_R1053] is a standardised version of it.
- [http://partsregistry.org/Part:BBa_C0053 BBa_C0053], the related coding region that activates BBa_R0053.
Problems
- [http://www.ncbi.nlm.nih.gov/entrez/viewer.fcgi?db=protein&val=16131068 HtlB] already exists in the strain of E. coli we're going to use (Escherichia coli K12 strain MG1655 (tax-id:511145).
- Go ahead with the experiment, and then find out whether the amount that already exists would significantly impact our results or not by checking with a prediction.
- Due to HtlB already existing in E. coli, if the initial concentration of CII isn't enough, would the oscillator never start?
References
- McMillen, D. et al (2002) "[http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=117365 Synchronizing genetic relaxation oscillators by intercell signalling]", Proc Natl Acad Sci USA, Vol. 22, No. 3, pp. 679-684.
- Where the oscillator idea came from.
The Synchronizer
The synchronizer integrates the use of cell quorum sensing to introduce synchronization of intercell oscillations.
Components
Component positions on Vibrio Fischeri
We performed a bunch of sequence alignments between the sequences from the Ecocyc database and the Biobrick parts to determine position of those components on Vibrio Fischeri.
Orange colour means transcription/translation in reverse direction.
norm means normalised (norm=x-1050153+1)
| Component | length | Start | End | norm start | norm end | Source |
|---|---|---|---|---|---|---|
| LuxI | 573 | 1050153 | 1050725 | 1 | 573 | [http://biocyc.org/VFIS312309/NEW-IMAGE?type=GENE&object=VFA0924 Biocyc] |
| BBa_C0161 | 585 | 1050150 | 1050734 | -2 | 582 | [http://partsregistry.org/Part:BBa_C0161 BBa_c0161] and sequence alignment[2] (87.3% consequence.) |
| BBa_R0062 (pLUX-right) | 55 | 1050749 | 1050803 | 597 | 651 | [http://partsregistry.org/Part:BBa_R0062:Design BBa_R0062] and sequence alignment[2] (72.7% consequence). |
| 55->57 | 1050784 | 1050840 | 632 | 688 | sequence alignment[1] (56% consensus and added 2bp). Looks wrong. | |
| BBa_R0061 | 30->28 | 1050781 | 1050808 | 627 | 656 | [http://partsregistry.org/wiki/index.php/Part:BBa_R0061 BBa_R0061] and sequence alignment[1] (75% consensus, but they deleted the first 2bp). |
| 30->43 | 1050775 | 1050817 | 623 | 665 | sequence alignment[2] (76.5% consensus. This looks wrong.) | |
| BBa_R0063 (pLUX-left) | 151->150 | 1050784 | 1050933 | 632 | 781 | [http://partsregistry.org/Part:BBa_R0063:Design BBa_R0063] and sequence alignment[1] (64.5% consensus and removed 1bp). |
| BBa_c0062 | 756->753 | 1050941 | 1051693 | 789 | 1541 | [http://partsregistry.org/Part:BBa_C0062 BBa_c0062] and sequence alignment[1] (78.9% consensus and removed 3bp) |
| LuxR | 753 | 1050941 | 1051693 | 789 | 1541 | [http://biocyc.org/VFIS312309/NEW-IMAGE?type=GENE-IN-MAP&object=VFA0925 Biocyc] |
[1]: Sequence Alignment with [http://biocyc.org/VFIS312309/sequence-delimited?chromosome=VF-CHROM-ES114-II&object=VFA0924&left=1050153&right=1051693 Biocyc Vibrio Fischeri (1050153-1051693)] (end of LuxI to end of LuxR).
[2]: Sequence Alignment with the reverse compliment of [1].
Related Biobricks
- [http://partsregistry.org/Part:BBa_R0062 BBa_R0062] or [http://partsregistry.org/Part:BBa_R1062 BBa_R1062] in conjunction with or instead of new construct (pRE).
- [http://partsregistry.org/Part:BBa_C0062 BBa_C0062] in conjunction with or instead of LuxR.
- [http://partsregistry.org/Part:BBa_C0061 BBa_C0061] or [http://partsregistry.org/Part:BBa_C0161 BBa_C0161] in conjunction with or instead of LuxI.
Other interesting parts that may come in use (all working):
- [http://partsregistry.org/Part:BBa_F2621 BBa_F2621]
- [http://partsregistry.org/Part:BBa_F2622 BBa_F2622]
- [http://partsregistry.org/Part:BBa_T9002 BBa_T9002]
Other interesting parts we may or may not use.
- [http://partsregistry.org/Part:BBa_J37019 BBa_J37019] titled AHL induced LuxR generator.
- [http://partsregistry.org/Part:BBa_J37032 BBa_J37032] if we want to test if [http://partsregistry.org/Part:BBa_R0062 BBa_R0062] works.
- [http://partsregistry.org/Part:BBa_J37033 BBa_J37033] looks pretty useless.
- [http://partsregistry.org/Part:BBa_I731012 BBa_I731012] looks interesting, but we're probably not going to do anything about it.
Problems
- Aligning [http://partsregistry.org/Part:BBa_R0062 BBa_R0062] and [http://partsregistry.org/Part:BBa_R1062 BBa_R1062] gives 92.2% consensus.
- Aligning [http://partsregistry.org/Part:BBa_R0063 pLUX-left] and [http://partsregistry.org/Part:BBa_R0062 pLUX-right] produced a 64% consensus.
- Why are so many nucleotides different?
- Why do the two promoters overlap so much (exactly according to the sequence alignment), yet the [http://partsregistry.org/Part:BBa_R0062:Design design] doesn't say so?
- Are they using a difference Vibrio Fischeri strain for both?
- A sequence [http://www.ncbi.nlm.nih.gov/entrez/viewer.fcgi?tool=portal&db=protein&term=&query_key=1&dopt=gp&dispmax=20&page=1&qty=1&WebEnv=0NZu2wco5pgt4AFlKGBZSBuHz1xTMr4TRAfhLUXMT0Mg3JomrvvDqnSCvcYGDYXw1omCOSTlDgeb-UU%40264C254084F93770_0012SID&WebEnvRq=1 similar to LuxR] exists in the E. coli strain we're using. Don't know if it'll affect anything.
References
- McMillen, D. et al (2002) "[http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=117365 Synchronizing genetic relaxation oscillators by intercell signalling]", Proc Natl Acad Sci USA, Vol. 22, No. 3, pp. 679-684.
- Same paper as the main paper from the oscillator section.
The Tuner
A way to control the oscillator's time period.
Components
Problems
- The CIIIC protein does not appear in any other papers (nor does it appear on pdb for that matter, but then again CIII doesn't either). So we have to believe that one paper.
- By synthesizing the CIIIC protein from DNA, we have to add the start codon ATG, resulting in an extra Methionine at the start. This may affect our experiments.
References
- Hadler, S. et al (2007) [http://www.ncbi.nlm.nih.gov/pubmed/17890311 Probing the Anitprotease Activity of lCIII, an Inhibitor of the Escherichia coli Metalloprotease HflB (FtsH)]", Journal of Bacteriology, Vol. 189, No. 22, pp. 8130-8138.
- The only paper about the experimental results of CIIIC. Also has a lot of experimental values (such as ratios) that can be used.
- Herman, C. et al (1997) "[http://jb.asm.org/cgi/content/abstract/179/2/358 The HflB Protease of Escherichia coli Degrades Its Inhibitor λcIII]", Journal of Bacteriology, Vol. 179, No. 2, pp. 358-363.
- Also has a lot of experimental values that could be used.
Sequences
This section lists the sequences of promoters and genes.
Promoters
The list of promoters used.
pRE
- few hundred bp upstream from the CI gene.
- 40 to 60bp long.
- Appropriate Literature
- [http://www.ncbi.nlm.nih.gov/pubmed/14872063?ordinalpos=3&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_RVDocSum Interactions among CII protein, RNA polymerase adn λ pRE promoter: contacts between RNA polymerase and the -35 region of pRE are identical in the presence and absence of CII protein]
- [http://www.ncbi.nlm.nih.gov/pubmed/2953648?ordinalpos=13&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_RVDocSum Mutations that improve the pRE promoter of coliphage lambda]
- [http://www.ncbi.nlm.nih.gov/pubmed/2953649?ordinalpos=12&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_RVDocSum Cross-specificities between cII-like proteins and pRE-like promoters of lambdoid bacteriophages]
complement(38391->38340) on lambda phage.
AGAGCCTCGTTGCGTTTGTTTGCACGAACCATATGTAAGTATTTCCTT|AGAT (52bp)
+---- transcription start site ---->
so what we want:
AGAGCCTCGTTGCGTTTGTTTGCACGAACCATATGTAAGTATTTCCTT (48bp)
pRE Overlaps with CII (but in the other direction)
ATGGTTCGTGCAAACAAACGCAACGAGGCTCTACGAATCGAGAGTGCGTTG--> CII cont'
<--ATCTAAGGAAATACTTACATATGGTTCGTGCAAACAAACGCAACGAGGCTCT (<--pRE)
For reference, [http://www.ncbi.nlm.nih.gov/sites/entrez?db=gene&cmd=Retrieve&dopt=full_report&list_uids=3827059 CI]: 37940-37227
Primers: AGAGCCTCGTTGCGTTT (len:17, tm:55deg, GC%:53) AAGGAAATACTTACATATGGTTCGTG (len:26, tm:55deg, GC%:35) Primer used regions: AGAGCCTCGTTGCGTTTGTTTGCACGAACCATATGTAAGTATTTCCTT
There is only 5bp not included in the primer. Feels kinda weird.
pLUX
- [http://partsregistry.org/wiki/index.php?title=Part:BBa_R0062 Part:BBa_R0062-luxR & HSL regulated lux pR]
acctgtaggatcgtacaggtttacgcaagaaaatggtttgttatagtcgaataaa(55 bp)
pLac
[http://partsregistry.org/Part:BBa_R0010 BBa_R0010] (200bp)
Genes
The list of genes used.
CII
- A λ phage gene.
- [http://www.ncbi.nlm.nih.gov/entrez/viewer.fcgi?val=NP_040630.1 NCBI protein]: 97aa
- [http://www.ncbi.nlm.nih.gov/entrez/viewer.fcgi?WebEnv=0_IrKu54rUXySHgWXB0EaSRAhFNRuWQc8t_9Pvcu9deRGYI7MtHyfyf2B79C0DiodB6zisjQl5K8vOD%40264C254084F93770_0012SID&db=nucleotide&qty=1&c_start=1&list_uids=NC_001416.1&uids=&dopt=fasta&dispmax=5&sendto=&fmt_mask=0&from=38360&to=38653&extrafeatpresent=1&ef_CDD=8&ef_MGC=16&ef_HPRD=32&ef_STS=64&ef_tRNA=128&ef_microRNA=256&ef_Exon=512 NCBI nucleotide]: 294bp
>gi|9626243:38360-38653 Enterobacteria phage lambda, complete genome ATGGTTCGTGCAAACAAACGCAACGAGGCTCTACGAATCGAGAGTGCGTTGCTTAACAAAATCGCAATGC TTGGAACTGAGAAGACAGCGGAAGCTGTGGGCGTTGATAAGTCGCAGATCAGCAGGTGGAAGAGGGACTG GATTCCAAAGTTCTCAATGCTGCTTGCTGTTCTTGAATGGGGGGTCGTTGACGACGACATGGCTCGATTG GCGCGACAAGTTGCTGCGATTCTCACCAATAAAAAACGCCCGGCGGCAACCGAGCGTTCTGAACAAATCC AGATGGAGTTCTGA
Primers: ATGGTTCGTGCAAACAAACG (56deg) tcagaactccatctggatttgt
CIIIC
- CIII (A λ phage gene) after residues 1-13 and 42-54 are cleaved (so only residues 14-41) are left.
- Inhibites the degradation rate HtlB has on CII by binding to HtlB.
- HtlB does not degrade CIIIC (unlike CIII).
- [http://www.ncbi.nlm.nih.gov/entrez/viewer.fcgi?db=protein&id=9626284 NCBI (CIII) protein]:54aa
- [http://www.ncbi.nlm.nih.gov/entrez/viewer.fcgi?WebEnv=0cMBABYtiOTu-KvSSXngm5VVsgrz7RFiY24GdDY0xzrnBMiRgRDlrJjhu6Ybt2Q7bn2aiqpN4WI4qWl%40264C254084F93770_0012SID&db=nucleotide&qty=1&c_start=1&list_uids=NC_001416.1&uids=&dopt=fasta&dispmax=5&sendto=&from=33299&to=33463&strand=on NCBI (CIII) gene]:165bp
CIII gene:
>gi|9626243:c33463-33299 Enterobacteria phage lambda, complete genome ATGCAATATGCCATTGCAGGGTGGCCTGTTGCTGGCTGCCCTTCCGAATCTTTACTTGAACGAATCACCC GTAAATTACGTGACGGATGGAAACGCCTTATCGACATACTTAATCAGCCAGGAGTCCCAAAGAATGGATC AAACACTTATGGCTATCCAGACTAA
Start with 165bp (54 residues + stop codon) then cleave off residues 2-13 (39bp) and 42-54 (39bp), leaving residues 1,14-41,55 (90bp)
CIIIC (with added ATG at the start):
ATGCAATATGCCATTGCAGGGTGGCCTGTTGCTGGCTGCCCTTCCGAATCTTTACTTGAACGAATCACCC GTAAATTACGTGACGGATGGAAACGCCTTATCGACATACTTAATCAGCCAGGAGTCCCAAAGAATGGATC AAACACTTATGGCTATCCAGACTAA
Changes to:
ATGCCTTCCGAATCTTTACTTGAACGAATCACCC GTAAATTACGTGACGGATGGAAACGCCTTATCGACATACTTAATCAGCCAGGA TAA
The resulting primers are:
ATGCCTTCCGAATCTTTACTTG ttatcctggctgattaagtatgtcg (56deg)
HtlB
- An E. coli gene.
- HtlB degrades CII.
- [http://biocyc.org/ECOLI/NEW-IMAGE?type=GENE&object=EG11506 Ecocyc]: 1935bp.
- No extra EcoRI,PstI,SpeI,XbaI in gene below.
>EG11506 hflB (complement(3324957..3323023)) E. coli atgGCGAAAA ACCTAATACT CTGGCTGGTC ATTGCCGTTG TGCTGATGTC AGTATTCCAG AGCTTTGGGC CCAGCGAGTC TAATGGCCGT AAGGTGGATT ACTCTACCTT CCTACAAGAG GTCAATAACG ACCAGGTTCG TGAAGCGCGT ATCAACGGAC GTGAAATCAA CGTTACCAAG AAAGATAGTA ACCGTTATAC CACTTACATT CCGGTTCAGG ATCCGAAATT ACTGGATAAC CTGTTGACCA AGAACGTCAA GGTTGTCGGT GAACCGCCTG AAGAACCAAG CCTGCTGGCT TCTATCTTCA TCTCCTGGTT CCCGATGCTG TTGCTGATTG GTGTCTGGAT CTTCTTCATG CGTCAAATGC AGGGCGGCGG TGGCAAAGGT GCCATGTCGT TTGGTAAGAG CAAAGCGCGC ATGCTGACGG AAGATCAGAT CAAAACGACC TTTGCTGACG TTGCGGGCTG CGACGAAGCA AAAGAAGAAG TTGCTGAACT GGTTGAGTAT CTGCGCGAGC CGAGCCGCTT CCAGAAACTC GGCGGTAAGA TCCCGAAAGG CGTCTTGATG GTCGGTCCTC CGGGTACCGG TAAAACGCTG CTGGCGAAAG CGATTGCAGG CGAAGCGAAA GTTCCGTTCT TTACTATCTC CGGTTCTGAC TTCGTAGAAA TGTTCGTCGG TGTGGGTGCA TCCCGTGTTC GTGACATGTT CGAACAGGCG AAGAAAGCGG CACCGTGCAT CATCTTTATC GATGAAATCG ACGCCGTAGG CCGCCAGCGT GGCGCTGGTC TGGGCGGTGG TCACGATGAA CGTGAACAGA CTCTGAACCA GATGCTGGTT GAGATGGATG GCTTCGAAGG TAACGAAGGT ATCATCGTTA TCGCCGCGAC TAACCGTCCG GACGTTCTCG ACCCGGCCCT GCTGCGTCCT GGCCGTTTCG ACCGTCAGGT TGTGGTCGGC TTGCCAGATG TTCGCGGTCG TGAGCAGATC CTGAAAGTTC ACATGCGTCG CGTACCATTG GCACCCGATA TCGACGCGGC AATCATTGCC CGTGGTACTC CTGGTTTCTC CGGTGCTGAC CTGGCGAACC TGGTGAACGA AGCGGCACTG TTCGCTGCTC GTGGCAACAA ACGCGTTGTG TCGATGGTTG AGTTCGAGAA AGCGAAAGAC AAAATCATGA TGGGTGCGGA ACGTCGCTCC ATGGTGATGA CGGAAGCGCA GAAAGAATCG ACGGCTTACC ACGAAGCGGG TCATGCGATT ATCGGTCGCC TGGTGCCGGA ACACGATCCG GTGCACAAAG TGACGATTAT CCCACGCGGT CGTGCGCTGG GTGTGACTTT CTTCTTGCCT GAGGGCGACG CAATCAGCGC CAGCCGTCAG AAACTGGAAA GCCAGATTTC TACGCTGTAC GGTGGTCGTC TGGCAGAAGA GATCATCTAC GGGCCGGAAC ATGTATCTAC CGGTGCGTCC AACGATATTA AAGTTGCGAC CAACCTGGCA CGTAACATGG TGACTCAGTG GGGCTTCTCT GAGAAATTGG GTCCACTGCT GTACGCGGAA GAAGAAGGTG AAGTGTTCCT CGGCCGTAGC GTAGCGAAAG CGAAACATAT GTCCGATGAA ACTGCACGTA TCATCGACCA GGAAGTGAAA GCACTGATTG AGCGTAACTA TAATCGTGCG CGTCAGCTTC TGACCGACAA TATGGATATT CTGCATGCGA TGAAAGATGC TCTCATGAAA TATGAGACTA TCGACGCACC GCAGATTGAT GACCTGATGG CACGTCGCGA TGTACGTCCG CCAGCGGGCT GGGAAGAACC AGGCGCTTCT AACAATTCTG GCGACAATGG TAGTCCAAAG GCTCCTCGTC CGGTTGATGA ACCGCGTACG CCGAACCCGG GTAACACCAT GTCAGAGCAG TTAGGCGACA AGTAA
HtlB primers: atgGCGAAAAACCTAATACTCTG ttacttgtcgcctaactgct
LuxI
- A Vibrio Fischeri gene.
- [http://biocyc.org/VFIS312309/NEW-IMAGE?type=GENE&object=VFA0924 Biocyc]: 573bp
- Biobrick available
- [http://partsregistry.org/wiki/index.php?title=Part:BBa_J37034 BBa_J37034]: LuxI + GFP. We could use this if we can extract the RBS+LuxI part.
>VFA0924 VFA0924 (complement(1050725..1050153)) V. fischeri chromosome es114 ii ATGataaaaa aatcggactt tttgggcatt ccatcagagg agtatagagg tattcttagt cttcgttatc aggtatttaa acgaagactg gagtgggact tggtaagtga ggataatctt gaatcagatg aatatgataa ctcaaatgca gaatatattt atgcttgtga tgatgcggaa gaggtaaatg gctgttggcg tttgttacct acaacgggtg attacatgtt aaaaactgtt tttcctgaat tgctcggaga tcaagtagcc ccaagagatc caaatatagt cgaattaagc cgttttgctg tgggaaaaaa tagctcaaaa ataaataact ctgctagtga aataacaatg aaattgtttc aagctatata taaacacgca gttagtcaag gtattacaga atatgtaaca gtaacatcaa tagcaataga gcgatttctg aaacgtatta aagttccttg tcatcgcatt ggtgataagg agattcattt attaggtaat actagatctg ttgtattgtc tatgcctatt aatgatcagt ttagaaaagc tgtatcaaat taa
LuxI primers: ATGataaaaaaatcggactttttggg ttaatttgatacagcttttctaaactgatc (length=30 too long?)
LuxR
- A Vibrio Fischeri gene.
- [http://www.ncbi.nlm.nih.gov/sites/entrez?Db=gene&Cmd=ShowDetailView&TermToSearch=3280300&ordinalpos=1&itool=EntrezSystem2.PEntrez.Gene.Gene_ResultsPanel.Gene_RVDocSum NCBI]: 753bp.
- Biobrick available
- [http://partsregistry.org/wiki/index.php?title=Part:BBa_J37019 BBa_J37019]: Promoter + LuxR. 862bp
>VFA0925 VFA0925 1050941..1051693 V. fischeri chromosome es114 ii ATGaacatta aaaatataaa tgctaatgag aagataattg ataaaattaa aacttgtaat aataataaag atattaatca atgtttatct gaaatagcaa agataataca ttgtgaatat tacctattcg ctattatcta tcctcactca ataattaaac ctgatgtttc aattatagat aattaccctg aaaaatggcg taaatattat gatgatgccg gactactaga atatgaccct gtagtcgatt actctaagtc ccatcattca ccaattaatt ggaacgtatt cgaaaaaaaa acaataaaaa aagagtctcc gaatgtaata aaagaagcac aggaatcggg actcattact ggatttagct ttccaattca tactgcaagt aatggttttg gaatgctcag ttttgctcat tcagataaag atatttatac tgacagttta tttttacacg ctagtacaaa tgtaccatta atgcttcctt ctttagtcga taattatcaa aaaataaata cgacacgtaa aaagtcagat tctattttaa caaaaagaga aaaagaatgc ttagcgtggg cgagtgaagg aaaaagtaca tgggatattt caaaaatact tggctgcagt gagcgtactg tcacttttca tttaaccaat actcaaatga aactcaatac aactaaccgc tgccaaagta tttctaaagc aattttaact ggcgccatta attgtccata ccttaaaaat taa
LuxR primers: ATGaacattaaaaatataaatgctaatgagaaga (length=34 too long?) ttaatttttaaggtatggacaattaatggc (length=30 too long?)
Testing
How we plan to test our parts when we've finished constructed them.
Promoter Testing
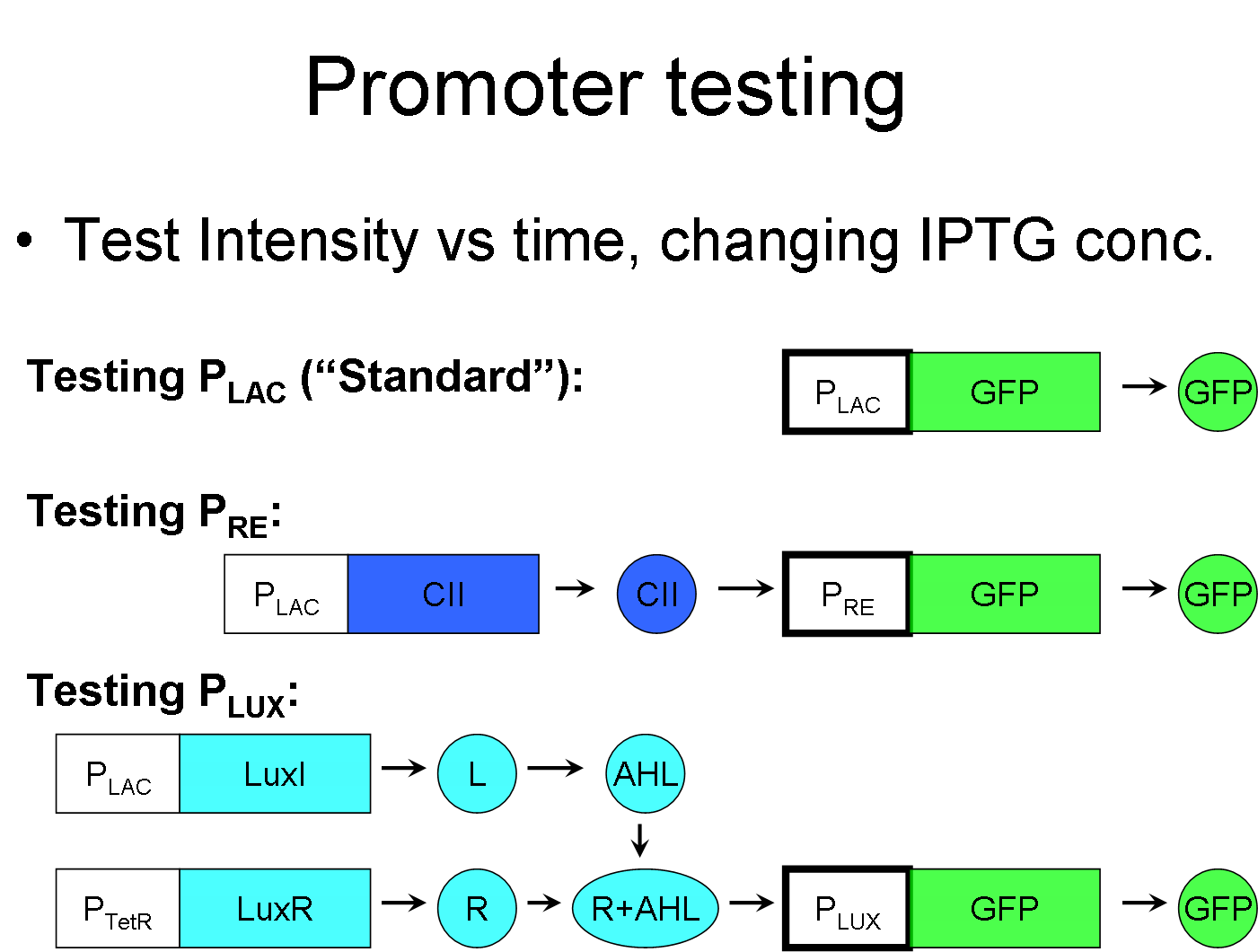
We are attempting to establish a "inducible promoter standard," much like the repressible promoter standard, where instead we define the IPTG inducible "standard" pLac promoter as having a value of 1, while all other inducible promoters will be measured in multiples of this standard. Then the value of IPTG can also be varied to test the ratios between the promoters as to whether the ratio is linear, sinusoidal, polynomial, etc.
We can test pRE promoter via this standard by connecting the |CII gene behind pLac, and connecting a reporter behind pRE.
This standard can also be used to test promoters that require a complex to be induced, such as pLux. pLux requires the R+AHL complex to be induced. To test the the strength of pLux, first let all parts of the complex save one to be always existing, while the last one would be controlled by pLac. In this example, LuxR is always expressed and the amount of AHL is determined by pLac.
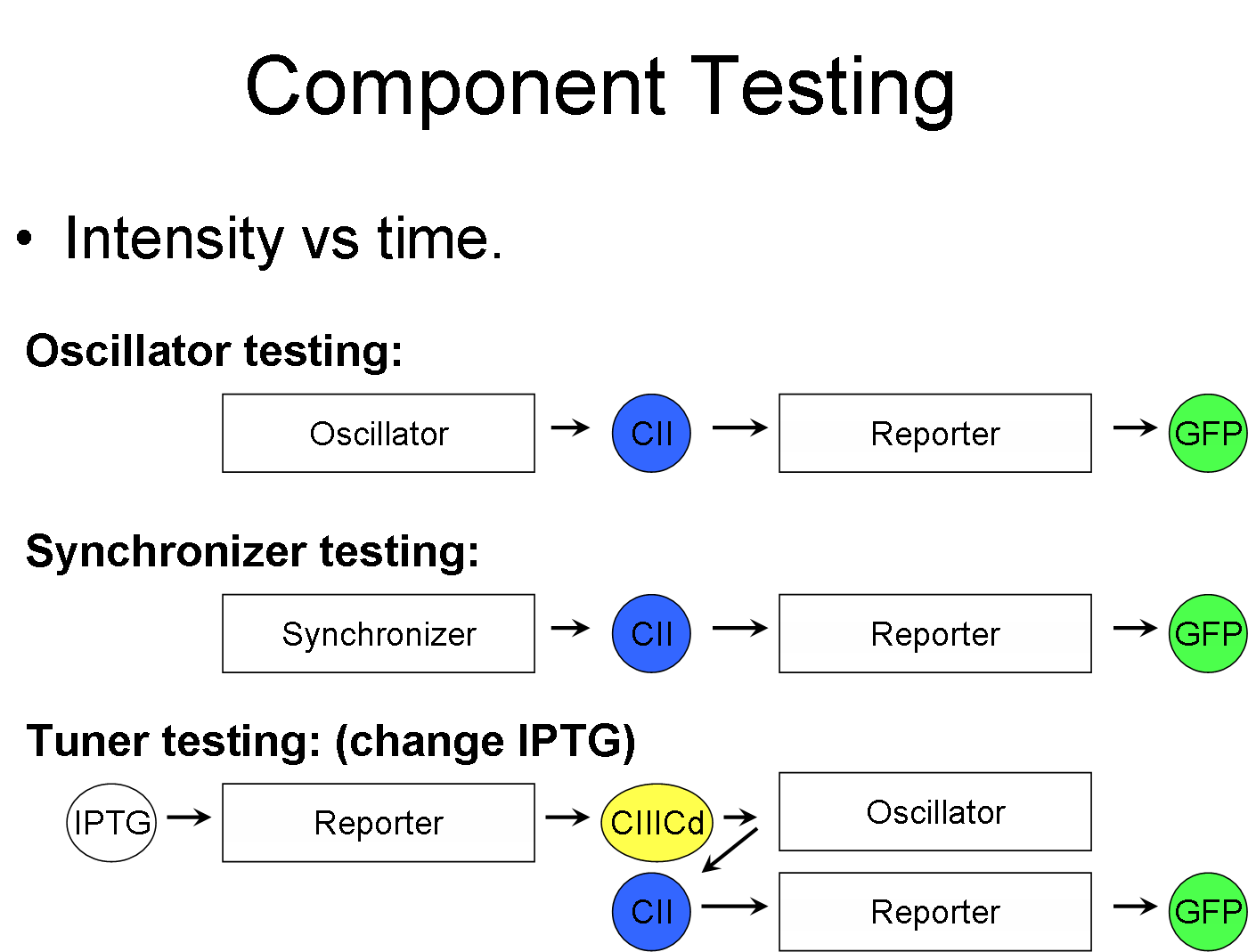
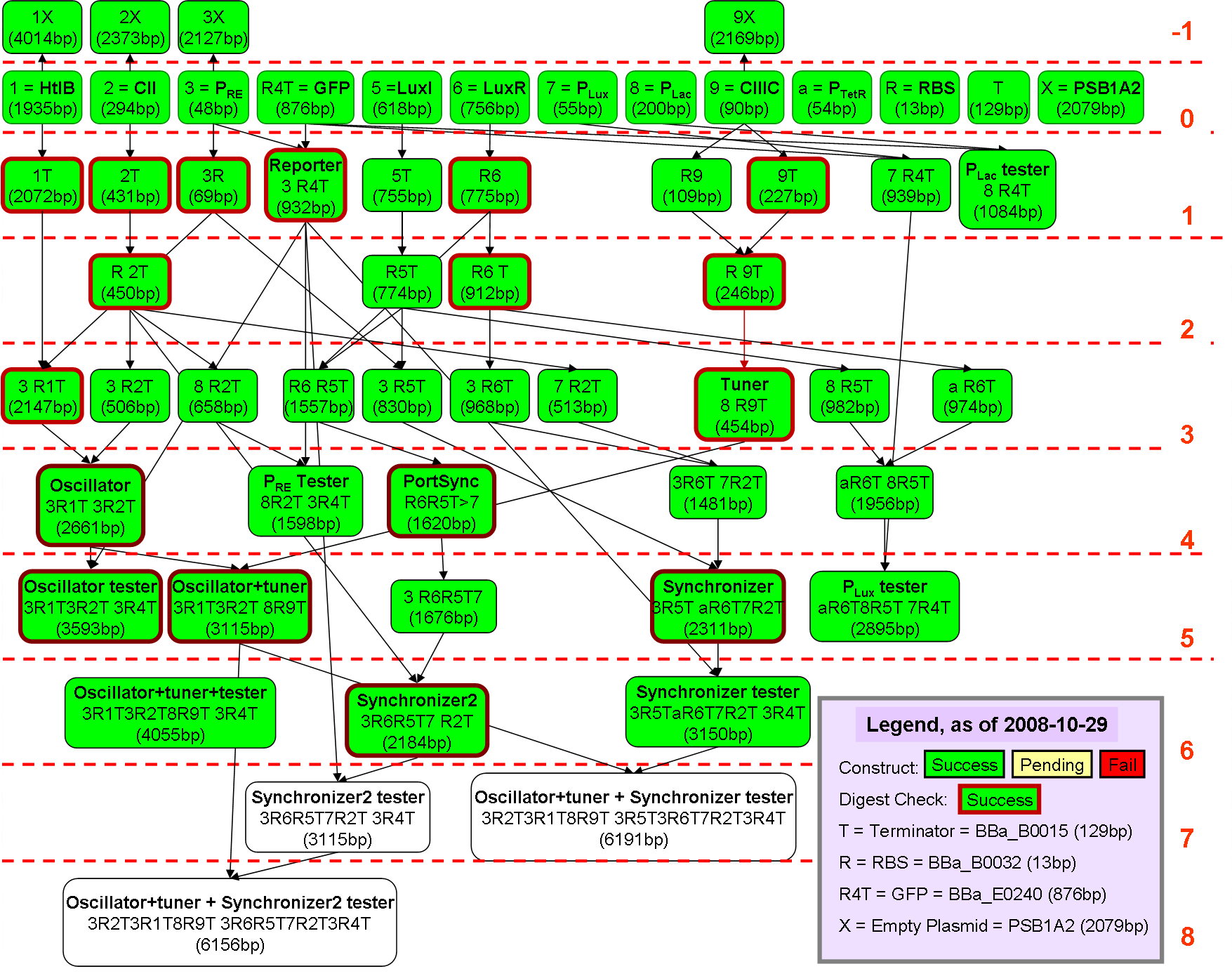
Component Testing
This is the part where we test to see if our component works. The diagram above shows the pathway between the various inputs, and the output (GFP).
Modelling
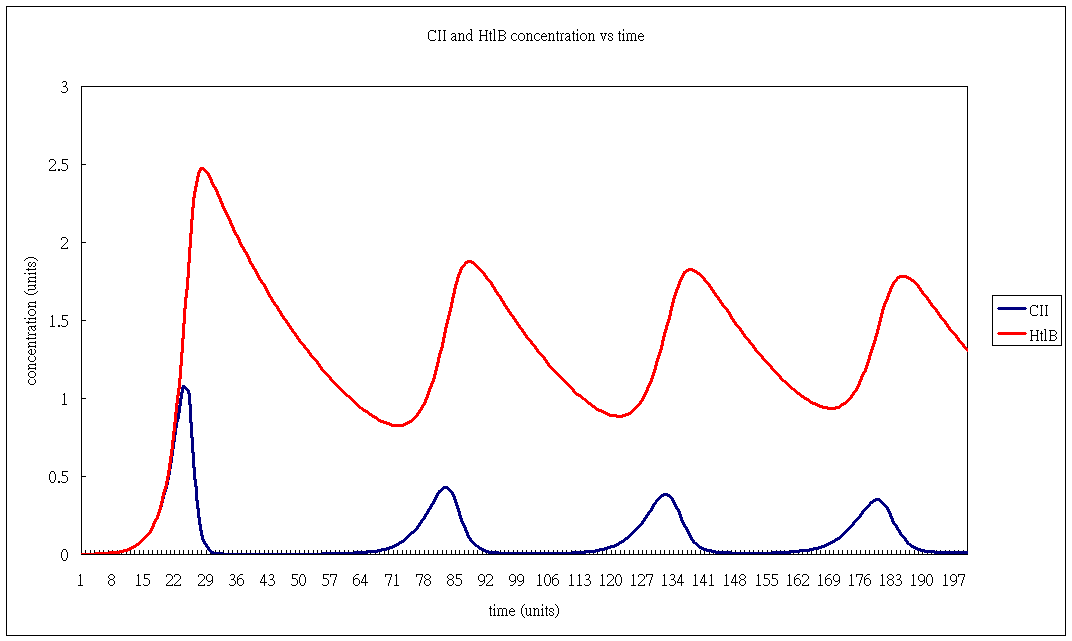
Oscillator Modelling 1
- HtlB concentration
- CII concentration.
- 20sec intervals.
Variables Used
- pRE Stength: 0.4 (1 CII protein produces 0.4 CII and HtlB proteins).
- H-t1/2: 8mins (random value I picked, its probably not the real value.)
- Half life of HtlB.
- Changing this value just makes the oscllation period longer.
- C0: 8mins.
- CII t1/2 with no HtlB present. Based on [http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=94496 Shotland2000])
- C1: 1min.
- Also based on the same paper.
- CII t1/2 with an equal amount of HtlB is what I assume because they don't say.
- Δt: 20sec
- Time segment used for the calculation (can't be too close to the t1/2's).
Formulas used
The HtlB/CII ratio used to calculate the t1/2 at a specific time point:
R = [HtlB] / [CII] * [CII] = [HtlB] ----- (1)
The last *[CII] has something to do with the [HtlB][CII] --> [CII] equation.
Calculation of CII's t1/2 based on the ratio above:
C-t1/2 = C0 x (C1/C0)R ----- (2)
The calculation of amount of substrates remaining w.r.t the half lives:
[HtlB] = [HtlB] x 0.5(Δt / H-t1/2) ----- (3) [CII] = [CII] x 0.5(Δt / C-t1/2) ----- (4)
Conclusion
Fiddling around a bit yields:
| Variable | Affects | |
|---|---|---|
| Initial concentration of: | HtlB (higher) | (longer) startup time |
| CII (higher) | ||
| Natural degradation rate of | HtlB (longer) | (longer) time period |
| CII (longer) | ||
| pRE strength (higher) | (shorter) time period | |
| HtlB to CII degradation rate (higher) | (lower) global maximum concentration (not counting startup) | |
Is a promoter that can be activated by HtlB? because then our calculations would be even more accurate.
Experimental Results
References
- McMillen, D. et al (2002) "[http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=117365 Synchronizing genetic relaxation oscillators by intercell signalling]", Proc Natl Acad Sci USA, Vol. 22, No. 3, pp. 679-684.
- Hadler, S. et al (2007) [http://www.ncbi.nlm.nih.gov/pubmed/17890311 Probing the Anitprotease Activity of λCIII, an Inhibitor of the Escherichia coli Metalloprotease HflB (FtsH)]", Journal of Bacteriology, Vol. 189, No. 22, pp. 8130-8138.
 "
"