Team:Hawaii/Notebook/2008-08- 1
From 2008.igem.org
| Projects | Events | Resources | ||
|---|---|---|---|---|
| Sponsors | Experiments | Milestones | Protocols | |
| Notebook (t) | Meetings (t) |
Things we did today
Wetlab work
Restriction Digest
- Grace
- Sequential digest of B0015 with XbaI and EcoRI
Checked transformants from yesterday's ligations
- Grace
Transformation of DB3.1 with nir and gfp constructs
| Constructs | Colony forming units |
|---|---|
| nir+rbs (1:6) | 5 |
| nir+rbs (1:3) | 6 |
| GFP+tt | 0 |
| GFPf+tt | 5 |
| BB-pRL1383a (1:2) | 19 + 2 clusters |
| BB-pRL1383a (1:4) | 5 |
PCR
- Grace & Krystle
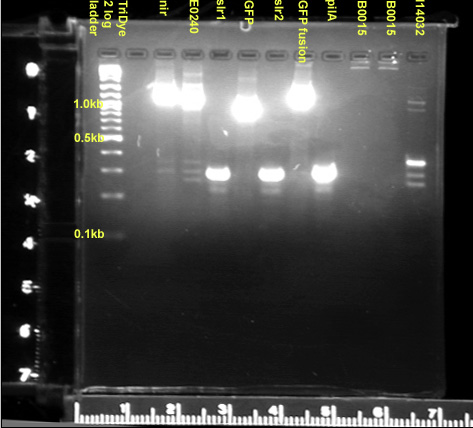
- Colony PCR'd all transformants from yesterday's ligations and slr1, slr2, nir, pilA, GFPf
- Annealed at 58C, elongated for 90 sec.
- PCR of E0240 and I14032 from filter paper
- Annealed at 58C, elongated for 90 sec.
- Ran on 2.5% gel
- Contaminant at 250bp
- pRL1383a = 5 bands (~1.2kb, ~0.9kb, ~0.35kb, ~0.3kb, ~0.25kb); desired band = 958bp
- nir+rbs = 2 bands (~0.275kb, ~0.25kb); desired band = 353bp
- nir was not inserted
- E/X sites not compatible; ligase ligated blunt ends created by RE digest leftovers?
- GFPf+tt = 2 bands (~0.32kb, ~0.25kb); desired band = 1081bp
- GFPf not inserted
- Extracted E0240, slr1, slr2, GFP, pilA, I14032 from gel (correct sizes)
- GFPf = ~1150bp (too big, what's going on? we're consistently getting GFPf this big)
- nir = ~1100bp (too big)
- Since sequencing returned correct sequences for GFPf and nir, regrow E. coli from colony used for sequencing and see if desired/expected results can be obtained or if results are the same
Inoculated LB+amp100
- Grace
- GFP
- nir
Made LB media
- Krystle
Discussion
Quote of the Day
Logically, it makes sense, but we're not always very logical people so it's still a weird numbering system - KS, GK in reference to the BioBrick part numbers
[http://manoa.hawaii.edu/  ][http://manoa.hawaii.edu/ovcrge/
][http://manoa.hawaii.edu/ovcrge/  ][http://www.ctahr.hawaii.edu
][http://www.ctahr.hawaii.edu  ]
]
 "
"