Team:BCCS-Bristol/Modeling-GRN
From 2008.igem.org
Contents |
Gene Regulatory Networks (GRN) Modelling
Introduction
Sender
The sender device consists of two genes (LuxI, GFP) and a single promoter (pCpxR). We are interested in the relationship between the activator protein CpxR and the production of the signalling compound N-acyl-homoserine lactone (AHL). AHL is produced by the LuxI enzyme which uses S-adenosylmethionine and the acylated-acyl carrier protein as substrates for the reaction. The concentration of the activator protein CpxR is specified as a contact profile mimicking the discovery of a particle. A linear piece-wise function was selected, ranging from 0 at t = 0, to a maximal CpxR protein concentration CpMax after 1 hour, taken from [15]. In the following equation time t is measured in minutes,
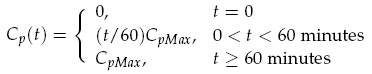
Both LuxI and GFP are controlled by the promoter pCpxR. In reality, because both genes are under the control of a single promoter, transcription would lead to the creation of a single mRNA strand. This mRNA would contain both genes, however, have separate ribosome binding sites (RBS) for each to allow for parallel translation. Due to the negligible effect on dynamics and a lack of information regarding degradation rates of this mRNA sequence, each gene transcription was treated separately. The transcription phase was modelled using a Hill function for the activation of the promoter with a degradation term for existing mRNA. This gives the following for mRNA concentrations of LuxI Im and GFP Gm:
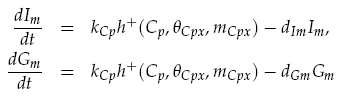
Translation of the mRNAs into proteins of LuxI Ip and GFP Gp was modelled as proportional to the current mRNA concentrations minus a degradation term, yielding the model:
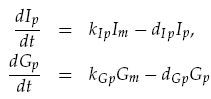
Transport
<math>\alpha\,\!</math> <math>&alpha</math> <math>$ alpha $</math> &alpha
<math>\sqrt{1-e^2}</math>
<math>\sqrt{2}</math>
<math>{}_1^2\!\Omega_3^4</math>
<math>-></math>
We were interested in understanding the relationship between AHL production at a point and the spatial concentration distribution over time. For the transportation of AHL between cells we assumed diffusion was the only process taking place. There is some evidence that AHL may be actively transported across cell membranes [8], however, by assuming only diffusion, a good indication of the worse case scenario could be gained. Assuming the diffusion co-efficient D is the same in all directions we have,
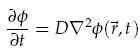
where f(~r, t) is the concentration of the diffusing chemical at location ~r and time t. As evolution of the concentration distribution is the same in all directions, we only considered a 1-dimensional case. The following ‘Forward Time Centred Space’ (FTCS) discrete numerical scheme was used to calculate the time evolution of the system, namely we have:
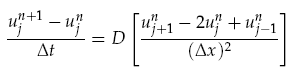
where unj<math>\u^n_j</math> is the chemical concentration at the discrete spatial index j and time index n, Dt is the time step, and Dx is the width of each spatial index. When computing the time evolution, the spatial environment was split into 1mm portions and a time step of 0.01s used.
Reciever
The receiver device consists of two genes (LuxR, mCherry) and two promoters (ptetR, Lux pR). We are interested in how the AHL concentration effects the output of our reporter protein mCherry. The protein LuxR is under the control of the constitutive promoter ptetR. This means that the promoter is permanently at its maximal transcription rate kTp, unless repressed. In our system, repression of the ptetR promoter is assumed to never occur. This allows the equation determining mRNA concentration of LuxR Rm to be simplified. The normal h+ term becomes equal to 1, removing the non-linearity. Degradation of the mRNA must also be considered to give the actual output concentration:
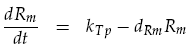
The activation of the Lux pR promoter, controlling mCherry output, is more complicated. To create a working transcription factor for Lux pR, two LuxR–AHL complexes form a dimer. To incorporate this feature we replicated the work in [3], modelling LuxR–AHL concentration CRA as:
Using this concentration for the LuxR–AHL complex as an input to the Hill function and including a standard degradation term, the mCherry mRNA concentration Mm can be modelled by:
Translation was modelled as being proportional to the mRNA concentration, minus a degradation term. Thus for LuxR and mCherry concentrations, Rp and Mp respectively, we have:
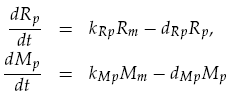
 "
"