Team:EPF-Lausanne/Project
From 2008.igem.org
| Home | The Team | The Project | Parts | 2-step PCR | Microfluidics | Modeling | Notebook |
|---|
Contents |
Genetic network generating spatial patterns through cell-cell communication and controlled information processing
Abstract
Biological systems are unique in their ability to combine information and energy to generate complex entities. Genetically encoded networks drive many of these patterning processes. Furthermore, developmental studies have highlighted the importance of gradient formation and cell-cell communication for the generation of cellular patterns in the early stages of life. It has been shown that simple networks can form both static and dynamic patterns. Nonetheless, a system whose pattern formation is dependent on combinations of multiple signals has yet to be demonstrated. Here we address this question by designing a network, involving two different quorum-sensing based signaling mechanisms. Upon introduction in E.coli, the system can sense the relative amounts of two input molecules. Using a pre-define set of rules which was selected on its ability to generate spatial patterns, the cell can then express its final state by emitting red or green fluorescence and transmit its state to its neighbors.
General Introduction and Context
Quorum sensing has been a wide-spread interest of synthetic biology in recent years. The ability to have populations of cell communicate in such a way that they either relay information or create specific patterns in culture is one which has been used in several studies (cite...). Some have relied on two different cell populations (predator-prey ref. for example) to engineer in vitro behaviors previously non existing in the bacteria.
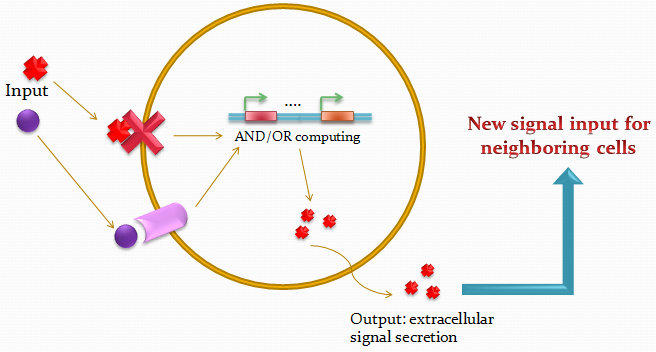
The idea of the project is to build a system which uses two different quorum sensing sets of molecules that make the cell able to integrate three different levels of signal, and emit a different response, by producing a given fluorescent protein and a quorum sensing molecule.
Our project uses a single population of cells, which all contain the same genetic information. Thus, the different behavior of the sub-populations will only be dictated by neighboring sub-populations. We also take advantage of the high-level engineering competences found in our Institute (EPFL) by using microfluidic devices in order to culture the cells in an ordered and controllable manner. These devices are custom-made for the project and consist in PDMS chips in which we design an array of miniature chambers to culture cells, linked to each other with a network of ducts. This enables us to culture sub-populations of our cells in contact with each other in an environment of medium, and to make these communicate with each other in a controlled manner for signal propagations.
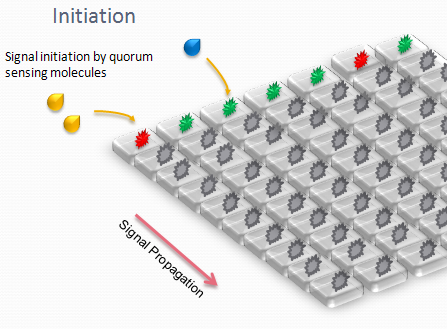
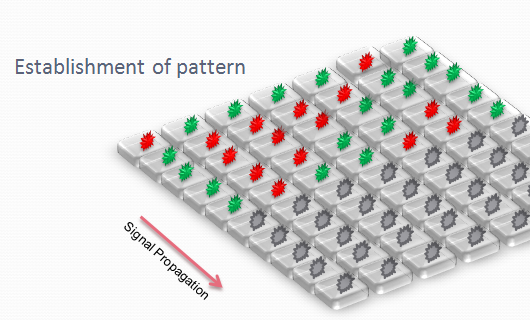
The aim is to initiate a signal on one side of the channel and observe the propagation along the plate.
Our microfluidic devices are also used in a side-project in order to characterize transcription factors (the MITOMI devices).
Biomedical application
We may be able to apply this idea to automated system of drug producing for personal medicine. In such a system, you just put a sample of blood in the first column of a microfluidics chip, then engineered cells in the wells detect some solutes, antibodies, and so on, and produce a signal given what they detect. Then the signal is integrated along the chip and when it reaches the last column, the engineered cell put in the wells are able to produce drugs responding to diseases detected along the chip.
As you would certainly suppose, we will not be able to engineer all the cell types to aim this goal in one summer, but we want through our project to make a few first steps to make such a system feasable in the future.
Initiation of the project
Our goal was to produce a system as simple as possible giving patterns as complex as possible. We therefore decided to use only two quorum sensing systems - and two fluorescent proteins - and to go through different network geometry and rule sets. We tried two different networks, and looked for all the rule sets possible for each network. We mean by "rule" the law determining the output response of the system to a given input.
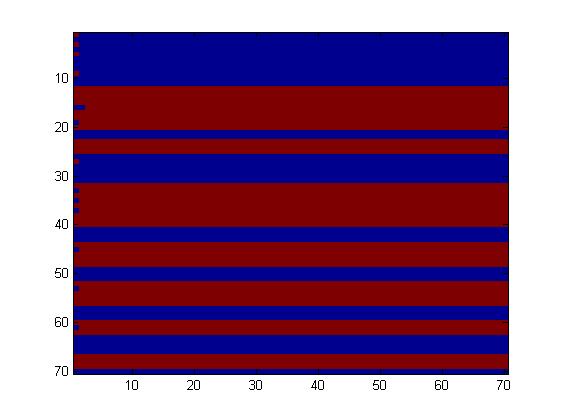
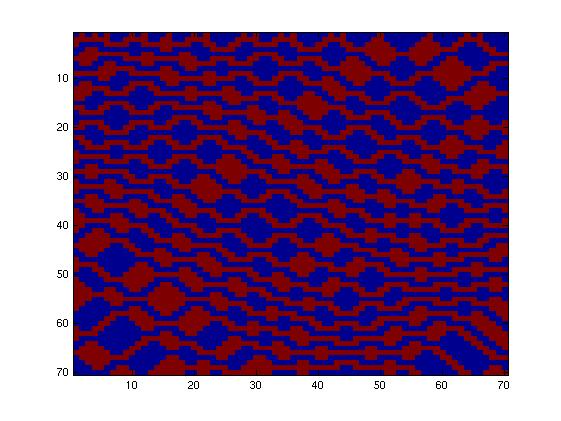
The first one, with really simple connections, gave too simple patterns, like the "barcode" one.
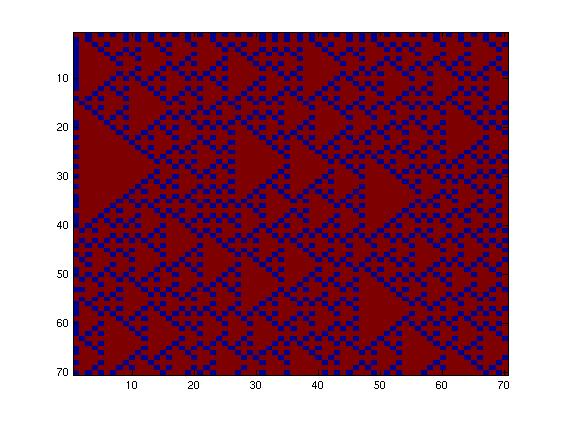
The second network, with more tricky connections, gave more interesting patterns like the "triangles" one.
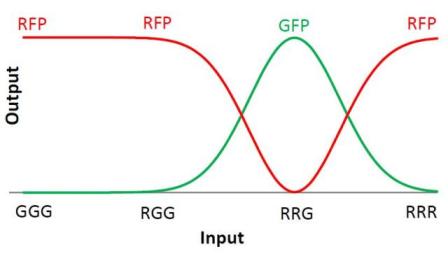
We choose to keep this pattern, and to try to implement its rule set, given in the following table. You can see the desired expression level of each fluorescent protein on the graph below.
Project Details
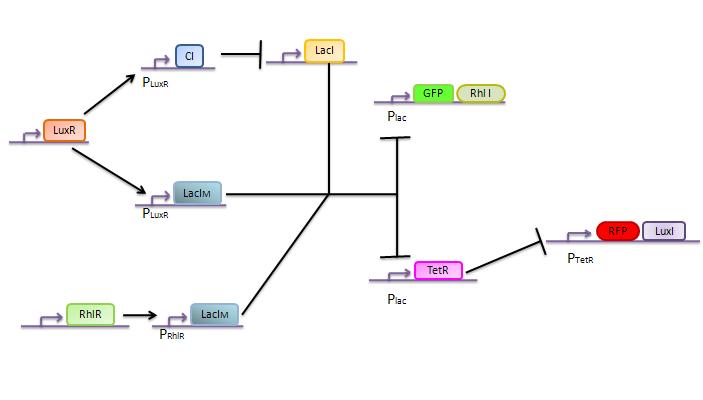
The genetic circuit which will be implemented in the cells is the following :
This genetic circuit must give this following pattern :
Our system is the combination between a new plasmid and a pre-existing system ("A synthetic multicellular system for programmed pattern formation", from Ron Weiss et al., Nature, vol.434, April 2005) which detect different level of the luxI quorum sensing molecule and give a specific response according to the concentration received. As shown on the previous genetic circuit, we will add the RhlI molecule after the GFP, Rhl being our second quorum sensing molecule used. In addition to the two Ron Weiss system plasmids, we have to build an extra plasmid with the second part of the genetic circuit. We will add an extra copy of the LacIm gene so that we can test it idenpendently from the Ron Weiss system.
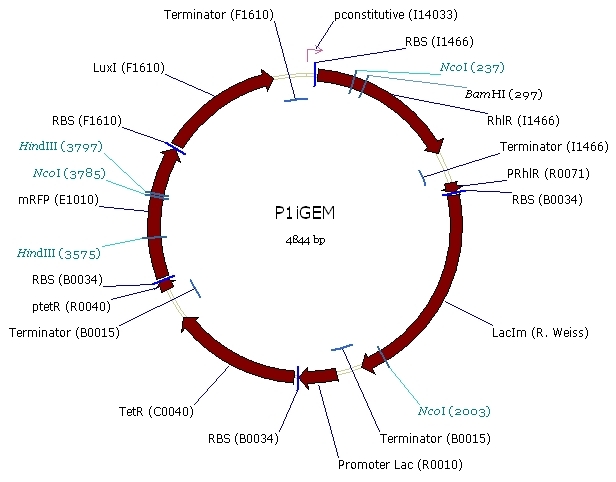
Here is the scheme of the plasmid with the Registry of Standard Parts references for each Biobrick we use.
Experiments
In order to complete our project, we followed mainly three courses of action. The first focuses on the network design and the mathematical modeling; it aims to draw the principles of information processing by individual cells and the cell population, with ODEs used to model the reaction of the system. Details on this part are available in the corresponding page of the Wiki : Modeling. This part was done with the help of professor Felix Naef.
The second part of the project was of course the concrete realisation of the plasmid in the laboratory. In the facilities provided by the Laboratory of Systems Biology and Genetics (http://deplanckelab.epfl.ch/) of Prof. Bart Deplancke. Using Biobricks and material obtained from the Ron Weiss group, we followed a cloning scheme to obtain our genetic circuit in E. coli.
Most of the information on the cloning can be found on the Parts section of this Wiki, with respect to the parts submitted, on this Project page higher up (description of the plasmid), and in the Notebook section for the protocols used in culture, transformation, digestion and ligation.
The third part of the project consisted in the building and using of the microfluidic chips for the two sides of the project in the Laboratory of Biological Network Characterization (http://lbnc.epfl.ch/) of Prof. Sebastian Maerkl at EPFL. First, the MITOMI approach to used these devices in order to characterise transcription factors was implemented in the laboratory. Then, the chips for cell culture and test of the genetic circuit designed and cloned were also constructed. Information on this part of the project can be found on the corresponding page of the Wiki : Microfluidics. Protocols for the fabrication of the devices and the preparation of DNA and transcription factors can also be found on the Notebook page of this Wiki.
Results
We were able to obtain the cloned plasmid with all the components.
Several MITOMI experiments were conducted with the chips.
We really need more detail on all of this, including Keshav's fluorescence measurements. We can link to the significant steps of the cloning in the Notebook part as well.
 "
"