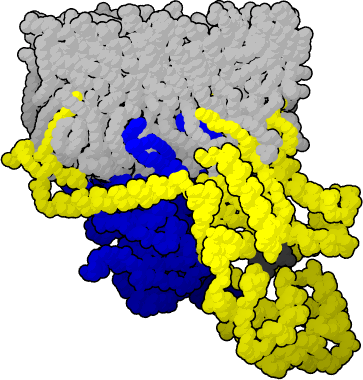
Team:Illinois/Antibody GPCR Fusion
From 2008.igem.org
(→Specific Plans, Supplies, and Protocols) |
|||
| Line 62: | Line 62: | ||
g. a replication origin for producing the plasmid in bacteria. | g. a replication origin for producing the plasmid in bacteria. | ||
| + | [http://www.pubmedcentral.nih.gov/pagerender.fcgi?artid=365362&pageindex=1#page] <--- This paper shows us how to fuse the FUS 1 gene. | ||
*Use site specific directed evolution to increase the effectiveness of the new GPCR. | *Use site specific directed evolution to increase the effectiveness of the new GPCR. | ||
**[http://www.escience.ws/b572/L4/L4.htm]Methods of site-directed mutagenesis, including non-PCR techniques | **[http://www.escience.ws/b572/L4/L4.htm]Methods of site-directed mutagenesis, including non-PCR techniques | ||
Revision as of 21:03, 15 June 2008
| Home | The Team | Notebook | Research Articles | Protocols | The Project | Pictures | Parts Submitted to the Registry |
|---|
Contents |
Announcements & Meetings
- Friday 13th Research Fest
- Sunday 15th Research Fest II
Core Team Members
Dave Luedtke, Bobak Hadidi, Kiruthika Selvadurai, Namita Bakshi, Toni Espina, Sarah Grajdura
- Add yourself!
Project Abstract
G protein-coupled receptors, or GPCRs, are transmembrane receptors that sense extracellular objects on the scale of small molecules to large proteins. "The importance of GPCR systems is illustrated by the fact that 30% of all clinically prescribed drugs function as GPCR agonists or antagonists" [2]. Activation of the GPCR by ligand binding begins a signal transduction pathway that ultimately results in the transcriptional activation or repression of one or more genes. The signal is transduced with G-proteins, which are signalling proteins that associate with GTP and GDP, as well as kinase cascades. Yeast cells are known to utilize two GPCRs signal transduction pathways, one to detect the presence of glucose and the other to initiate mating. We hope to engineer the well-characterized mating pathway to produce a colorimetric change in the cell upon detecting a novel molecule-- a surface protien of some water-borne pathogen, or possibly a toxin secreted by such a water-borne pathogen. One issue that may have to be dealt with is the cell wall of the yeast: it may prevent the target protein from drawing near enough to the receptor to activate it.
Specific Plans, Supplies, and Protocols
In theory, this is how we will progress:
- Create a fusion protein that links an antibody against cholera toxin to the Ste2 GPCR of S. cereviviae, the pheremone response GPCR.
- 1)Find the sequences of the GPCR [http://www.bio.davidson.edu/Courses/genomics/2002/Statler/STE2.htm] and the antibody. Did someone already find the sequence of the antibody? [http://www.hytest.fi/product/cholera-toxin-antibody.html] <--- We can buy the antibody to the beta subunit here.
- 2)Select site of fusion
- Cys187 in EL2 and Cys110 in TM3: highly conserved cysteines found in most GPCRs
- "extracellular loop 2 has extensive contacts with the other extracellular domains and also plunges down into the transmembrane bundle, contacting the retinal ligand." If we add the antibody here, will it interfere with the proper folding of the surface protein? [1]
- "the first extracellular loop is important in ligand-mediated activation of the receptor, whereas the second loop functions as a negative regulator of receptor activation and serves to stabilize the inactive state of the receptor" [2]
- Possible sites for fusion of antibody: Tyr101 through Gln135 in EL1 domain (because they did not have major effects on binding and signal transduction when they were mutated to Cysteine)
- Also, we want to add the antibody at a point where the unbound state (of the GPCR) is in contact with the solvent: residues 101 and 106
- "mutation of residues Leu102, Asn105, Ser108, Tyr111, and Thr114 to cysteine resulted in receptors that could effectively bind pheromone, but they were partially (Leu102 and Gln135) or severely (Asn105, Ser108, and Tyr111) compromised with respect to signal transduction." [3]
- 3)Have the gene sequenced--This will be a lengthy gene. We might be better off somehow obtaining a gene containing the natural GPCR (perhaps from a research group) and inserting the gene of the antibody.
- Professor Chris Rao explained to us an online tool (it was some company) for determining what restriction enzymes and where in the gene they could act to aid us in this.
- One yeast Ste2 gene was sequenced to be ~1.5kb, including restriction sites for EcoRI and BamHI [9].
- Express the antibody/GPCR fusion protein in yeast that lack the wild type receptor.
- Found [http://www.lgcpromochem-atcc.com/common/catalog/yeastGeneticStock/yeastGeneticStockIndex.cfm 6 strains] (type "ygk ste3|ste2" without quotes into the text search) of yeast of interest from the Yeast Genetic Stock Center, 2 of mating type a, 2 of mating type alpha, and 2 that are diploid (both). Three of each type have their Ste2 gene knocked out; the other 3 likewise have their Ste3 gene knocked out. Each costs about $150, in addition the media for their growth conditions is listed, WOW!
- MATa expresses Ste2 (the alpha-factor receptor) while MATalpha expresses Ste3 (the a-factor receptor) [1]. We should thus either order MATa without Ste2 or MATalpha without Ste3.
- Which mating type would better suite our purposes?
- The gene will most likely be on a plasmid. We may have to integrate the gene into the yeast's chromosome, however.
- The size of the gene may be substantial, if it turns out to be too large for a plasmid, integration into the yeast chromosome may be the way to go (hopefully not).
- [9] cites that a form of the Ste2 gene has been carried by a vector into yeast, size should not be a problem unless the antibody module is large.
- The size of the gene may be substantial, if it turns out to be too large for a plasmid, integration into the yeast chromosome may be the way to go (hopefully not).
- Found [http://www.lgcpromochem-atcc.com/common/catalog/yeastGeneticStock/yeastGeneticStockIndex.cfm 6 strains] (type "ygk ste3|ste2" without quotes into the text search) of yeast of interest from the Yeast Genetic Stock Center, 2 of mating type a, 2 of mating type alpha, and 2 that are diploid (both). Three of each type have their Ste2 gene knocked out; the other 3 likewise have their Ste3 gene knocked out. Each costs about $150, in addition the media for their growth conditions is listed, WOW!
- Measure the activation of the GPCR by the toxin (and by the natural pheremone) using a reporter gene.
- Several studies have used the FUS1 promoter in conjunction with HIS3 selection. The FUS1 promoter seems to be a good choice for our purposes.
- Here's a recombinant plasmid we will need to contruct:
[http://www.freepatentsonline.com/5063154.html] a. the FUSI promoter isolated from Saccharomyces cerevisiae;
b. DNA encoding approximately the first 254 amino acids of said FUSI gene;
c. DNA encoding a protein or a polypeptide of interest (Could be His3), the DNA positioned so as to be under the control of said FUSI promoter;
d. DNA encoding a selectable marker in yeast;
e. a two micron autonomously replicating sequence;
f. DNA encoding a selectable marker in bacteria; and
g. a replication origin for producing the plasmid in bacteria. [http://www.pubmedcentral.nih.gov/pagerender.fcgi?artid=365362&pageindex=1#page] <--- This paper shows us how to fuse the FUS 1 gene.
- Use site specific directed evolution to increase the effectiveness of the new GPCR.
- [http://www.escience.ws/b572/L4/L4.htm]Methods of site-directed mutagenesis, including non-PCR techniques
- [http://web.archive.org/web/20010222093603/http://www.labmarket.co.kr/methodpcr5.htm]Protocol for site-directed mutagenesis using PCR
- Altered-States Method[http://www.escience.ws/b572/L4/L4.htm]:
- 1. Start with a plasmid carrying a defective selectable marker (e.g. Amp)
- 2. Link the mutation you are making elsewhere in the plasmid to a correction of the defective selectable marker.
- 3. Select for correction of the selectable marker, and you are likely to also find plasmids with your specific mutation introduced as well
- QuikChange Method[http://www.bio.davidson.edu/courses/Molbio/MolStudents/spring99/sarah/method.html]
- Gene of interest in plasmid, but it is modified (bp's added, removed, substituted), DNA is denatures and primers anneal to both strands, there are "staggered nicks" where mutation is, Dpn I added (digests methylated DNA), parent DNA is methylated but new mutated DNA is not, result is that only new mutant DNA (see below, brown region w/ green x's) is transformed in bacteria cell
- http://www.bio.davidson.edu/courses/Molbio/MolStudents/spring99/sarah/quikchg4.gif
Alternatively, we might begin looking into completely replacing the yeast receptor with a mammalian or novel receptor with an affinity for our target protein. If we go down this route, we will also have to replace the G-alpha subunit with a chimeric one that has affinity for both the new receptor and the yeast G-betagamma complex. These have already been synthesized/cloned/will be fairly easy to procure, and may or may nor be receptor specific. This method is the major procedure for researching GPCRs today, typically used in pharmaceutical research -- knock out the pheromone GPCR while leaving the endogenous signal pathway/reporter intact, then introduce the uncharacterized GPCR of interest (with its G-aplha). In our case, we would not be studying an unknown GPCR to find what its target ligand or purpose is; instead we would simply already have one that we had/would evolve using mutagenesis to have an affinity for the toxin.
To Research
- Cell Wall Issue
- "The potency of some ligands might be reduced in yeast compared with mammals [for receptors taken from mammalian cells]. Using Gpa1–Ga chimeras probably explains part of this discrepancy [original yeast G-alpha subunits have low affinity for the new receptors and the mammalian G-alpha has low affinity for yeast G-betagamma, we will bypass this problem altogether by fusing the antibody to the original yeast receptor], but another factor might be the ability of ligands to penetrate the yeast cell wall. Potencies for small ligands are similar in yeast and mammals, those for intermediate ligands are about two orders of magnitude less in yeast, and large ligands such as chemokines (polypeptides with 70–80 residues) are often unable to activate receptors when applied exogenously to yeast. The fact that smaller versions of these ligands can be more efficient agonists suggests that accessibility is an issue."[1]
- Antibody Sequence
- Sources of yeast strains deficient in specific genes
- Cholera Toxin or possible target protein
G Proteins and their Receptors
G proteins are comprised of 3 subunits, collectively known as a heterotrimeric or large G protein. The individual pieces are Gα, Gβ and Gγ. When the appropriate molecule is bound to the receptor, the GDP bound to the Gα unit becomes GTP. More conformational changes are induced causing Gβγ and Gα to dissociate. Depending on the signal cascade, either complex, Gα-GTP or Gβγ may now induce the appropriate linked signal pathway.
As opposed to other signal receptors, the ligands that bind GPCRs typically bind within the transmembrane domain. When the receptor is bound by its ligand, it shifts shape, activating the G protein (by allowing the exchang of GDP for GTP). Also, G protein complexes may or may not be coupled with the receptors, instead, if they happen across a receptor with its activating ligand bound, the G protein will be activated (GDP -> GTP) and induce its signal cascade. (wikipedia).
Literature Research and References
[1]Functional analysis of heterologous GPCR signaling pathways in yeast
[2]http://www.nature.com/embor/journal/v2/n7/full/embor385.html
Ste2 information:http://www.bio.davidson.edu/Courses/genomics/2002/Statler/STE2.htm
[9] [http://www.pubmedcentral.nih.gov/picrender.fcgi?artid=280318&blobtype=pdf Ste2 Protein]
2 GPCR pathways in yeast: http://www.nature.com/embor/journal/v2/n7/images/embor385-f1.gif
 "
"