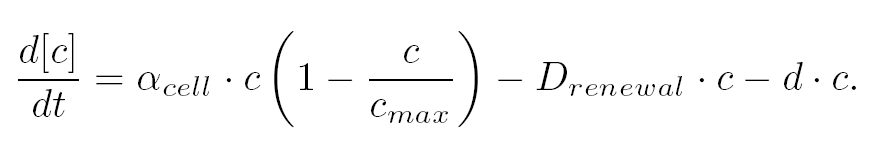Team:Paris/Analysis/Construction2
From 2008.igem.org
(→Kinetics) |
(→Kinetics) |
||
| Line 31: | Line 31: | ||
<TABLE WIDTH=90%> | <TABLE WIDTH=90%> | ||
<TR ALIGN=CENTER VALIGN=MIDDLE> | <TR ALIGN=CENTER VALIGN=MIDDLE> | ||
| - | <TD COLSPAN=2 BGCOLOR="#649CD7"><B>Common Dynamics: | + | <TD></TD> |
| + | <TD COLSPAN=2 BGCOLOR="#649CD7"><B>Common Dynamics: Chemostat</B></TD> | ||
| + | <TD></TD> | ||
</TR> | </TR> | ||
<TR ALIGN=LEFT VALIGN=MIDDLE> | <TR ALIGN=LEFT VALIGN=MIDDLE> | ||
| + | <TD></TD> | ||
<TD COLSPAN=2>The variation of cells' concentration in the chemostat over time can be expressed in terms of a production (positive) term and degradation (negative) terms:</TD> | <TD COLSPAN=2>The variation of cells' concentration in the chemostat over time can be expressed in terms of a production (positive) term and degradation (negative) terms:</TD> | ||
| + | <TD></TD> | ||
| + | </TR> | ||
| + | |||
| + | <TR ALIGN=CENTER VALIGN=MIDDLE> | ||
| + | <TD></TD> | ||
| + | <TD COLSPAN=2>[[Image:final_chemostat.png|center|300px]] </TD> | ||
| + | <TD></TD> | ||
| + | </TR> | ||
| + | |||
| + | <TR ALIGN=LEFT VALIGN=MIDDLE> | ||
| + | <TD></TD> | ||
| + | <TD COLSPAN=2> For the production term, we use a logistic equation to model cell growth, according to standard assumptions [[Team:Paris/Modeling/BOB#Bibliography|[Garcia-Ojalvo]]]. The behaiviour obtained is the following one: at low population density, the concentration of cells in the chemostat (c) increase exponentialy with a growth rate α<sub>cell</sub> and at high population density, the population reaches a maximum concentration, c<sub>max</sub>. | ||
| + | </TD> | ||
| + | <TD></TD> | ||
| + | </TR> | ||
| + | |||
| + | <TR ALIGN=LEFT VALIGN=MIDDLE> | ||
| + | <TD></TD> | ||
| + | <TD COLSPAN=2> For the degradation term, we consider that c decrease proportionaly to both a dilution phenomena cause by the renewal of the medium in the chemostat (D<sub>renewal</sub>) and cell death (d). | ||
| + | </TD> | ||
| + | <TD></TD> | ||
| + | </TR> | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <TR ALIGN=CENTER VALIGN=MIDDLE> | ||
| + | <TD></TD> | ||
| + | <TD COLSPAN=2 BGCOLOR="#649CD7"><B>Common Dynamics: Quorum Sensing </B></TD> | ||
| + | <TD></TD> | ||
</TR> | </TR> | ||
<TR ALIGN=LEFT VALIGN=MIDDLE> | <TR ALIGN=LEFT VALIGN=MIDDLE> | ||
| - | <TD COLSPAN=2> | + | <TD></TD> |
| + | <TD COLSPAN=2> To model the quorum sensing dynamics, we consider: </TD> | ||
| + | <TD></TD> | ||
</TR> | </TR> | ||
<TR ALIGN=LEFT VALIGN=MIDDLE> | <TR ALIGN=LEFT VALIGN=MIDDLE> | ||
| - | <TD COLSPAN=2> | + | <TD></TD> |
| + | <TD COLSPAN=2>cont.. </TD> | ||
| + | <TD></TD> | ||
</TR> | </TR> | ||
<TR ALIGN=CENTER VALIGN=MIDDLE> | <TR ALIGN=CENTER VALIGN=MIDDLE> | ||
| - | <TD BGCOLOR="#D4E2EF"><B>Bimodular | + | <TD COLSPAN=2 BGCOLOR="#D4E2EF"><B>Bimodular System Model</B></TD> |
| - | <TD BGCOLOR="#D4E2EF"><B>Single module | + | <TD COLSPAN=2 BGCOLOR="#D4E2EF"><B>Single module System Model</B></TD> |
</TR> | </TR> | ||
<TR ALIGN=LEFT VALIGN=MIDDLE> | <TR ALIGN=LEFT VALIGN=MIDDLE> | ||
| - | <TD>2</TD> | + | <TD COLSPAN=2>2</TD> |
| - | <TD>3</TD> | + | <TD COLSPAN=2>3</TD> |
</TR> | </TR> | ||
</TABLE> | </TABLE> | ||
Revision as of 18:33, 27 October 2008
|
Model Construction
MotivationWe are interested to understand the factors that account for an oscillating behaviour at population level. We build two models to study systems that are equipated with quorum sensing capabilities but that relay on different designs principles. The proposed models are:
Both the modular and the all-in-one models describe events that happend not only at the cellular level (as in the core system) but also at the population level due to interactions needed bettwen a cell and its environment during quorum sensing. In the following sections, we first describe the population modeling (the common part amoung our two proposed models) to then focus our attention to the characteristics that are exclusive to each of the modeling alternatives.
Population Modeling
Kinetics
Parameters Searchmanly from literature but also from S0 analysis.
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"