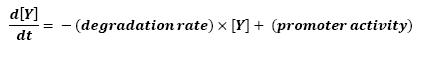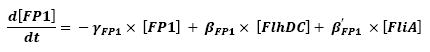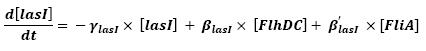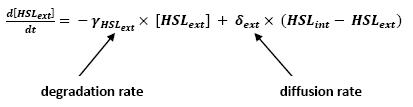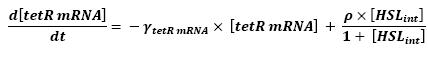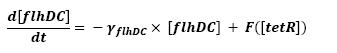Team:Paris/Modeling
From 2008.igem.org
(→An Oscillatory Biological Model) |
(→An Oscillatory Biological Model) |
||
| Line 38: | Line 38: | ||
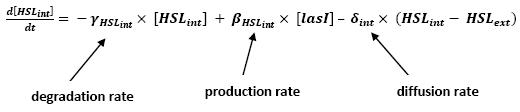
<br>The goal here is to present the differential equations we used for our system modelization. At each step, we shall describe why we chose this precise model, its drawbacks and possible improvments, the parameters involved and enventually a biologically coherent value. | <br>The goal here is to present the differential equations we used for our system modelization. At each step, we shall describe why we chose this precise model, its drawbacks and possible improvments, the parameters involved and enventually a biologically coherent value. | ||
| - | <br>The key problem with a | + | <br>The key problem with a dynamic system consists in the fact that adding a new equation gives a more detailed idea of the overall process, but one looses precision by doing so since new parameters appear. |
<br>In that respect (in all cases but one) we chose not to model the mRNA steps, that is translation and transcription. We then assumed that we could act as if a protein would directly beget another one, without loosing too much precision. | <br>In that respect (in all cases but one) we chose not to model the mRNA steps, that is translation and transcription. We then assumed that we could act as if a protein would directly beget another one, without loosing too much precision. | ||
<br>This first approach only refers to a single cell. We shall examine later on what happens if we put more cells together. | <br>This first approach only refers to a single cell. We shall examine later on what happens if we put more cells together. | ||
===Bibliography=== | ===Bibliography=== | ||
| - | + | Musch of our inspiration comes from four articles to which we shall refer in the next subsections : | |
| + | <br> | ||
<br>[1] Shiraz Kalir, Uri Alon. Using quantitative blueprint to reprogram the dynamics of the flagella network. Cell, June 11, 2004, Vol.117, 713-720. | <br>[1] Shiraz Kalir, Uri Alon. Using quantitative blueprint to reprogram the dynamics of the flagella network. Cell, June 11, 2004, Vol.117, 713-720. | ||
<br>[2]Jordi Garcia-Ojalvo, Michael B. Elowitz, Steven H. Strogratz. Modeling a synthetic multicellular clock : repressilators coupled by quorum sensing. PNAS, July 27, 204, Vol. 101, no. 30. | <br>[2]Jordi Garcia-Ojalvo, Michael B. Elowitz, Steven H. Strogratz. Modeling a synthetic multicellular clock : repressilators coupled by quorum sensing. PNAS, July 27, 204, Vol. 101, no. 30. | ||
Revision as of 14:10, 6 August 2008
|
 "
"