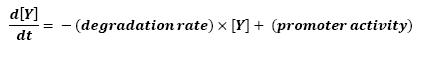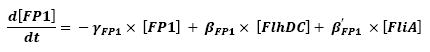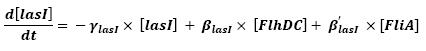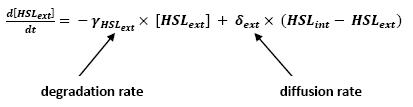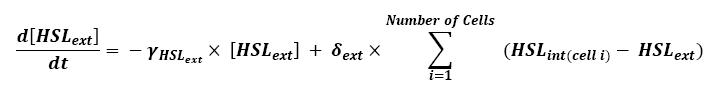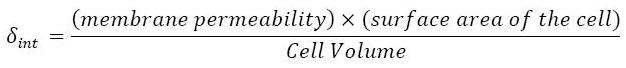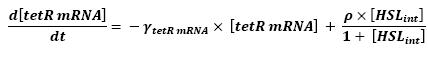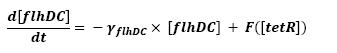Team:Paris/Modeling/linear approach
From 2008.igem.org
(→An Oscillatory Biological Model) |
(ssdwerhj) |
||
| (10 intermediate revisions not shown) | |||
| Line 29: | Line 29: | ||
** Influence of flhDC and fliA over flhB - lasI | ** Influence of flhDC and fliA over flhB - lasI | ||
** Influence of lasI over HSL<sub>int</sub> | ** Influence of lasI over HSL<sub>int</sub> | ||
| - | ** Relation between HSL<sub>int</sub> and HSL<sub>ext</ | + | ** Relation between HSL<sub>int</sub> and HSL<sub>ext</sub> |
** Influence of HSL<sub>int</sub> over tetR mRNA | ** Influence of HSL<sub>int</sub> over tetR mRNA | ||
** Influence of tetR mRNA over tet R | ** Influence of tetR mRNA over tet R | ||
| Line 36: | Line 36: | ||
===Establishing the model=== | ===Establishing the model=== | ||
* <b>Steps</b> | * <b>Steps</b> | ||
| - | ** flhDC | + | ** Influence of flhDC over fliA |
| - | ** flhDC | + | ** Influence of flhDC and fliA over fliL - Fluorescent Protein 1 (FP1) |
| - | ** flhDC | + | ** Influence of flhDC and fliA over flgA - Fluorescent Protein 2 (FP2) |
| - | ** flhDC | + | ** Influence of flhDC and fliA over flhB - Fluorescent Protein 3 (FP3) |
| - | ** flhDC | + | ** Influence of flhDC and fliA over flhB - lasI |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
<br> | <br> | ||
| Line 72: | Line 67: | ||
** As presented in the first equation below, there is a retroaction from fliA over fliA. In a first approach, we decided not to take into account this interaction, that is setting β'<sub>FliA</sub> to zero. In a second approach, we will consider this retroaction, and find a way to describe it mathematically. | ** As presented in the first equation below, there is a retroaction from fliA over fliA. In a first approach, we decided not to take into account this interaction, that is setting β'<sub>FliA</sub> to zero. In a second approach, we will consider this retroaction, and find a way to describe it mathematically. | ||
** [[Team:Paris/Modeling/linear_approach#Discussion|Another step]] consists in choosing the proper units, so as to have a homogenous set of parameters. | ** [[Team:Paris/Modeling/linear_approach#Discussion|Another step]] consists in choosing the proper units, so as to have a homogenous set of parameters. | ||
| - | ** Finally, it is important to understand at this stage the choice we made when we decided to present this model. Our goal was to have a rough but somehow realistic (i.e. grounded on studies available on the literature) model with which we could start to work, for example to have an idea of time scales. This is the reason why we ended with this | + | ** Finally, it is important to understand at this stage the choice we made when we decided to present this model. Our goal was to have a rough but somehow realistic (i.e. grounded on studies available on the literature) model with which we could start to work, for example to have an idea of time scales. This is the reason why we ended with this linear appproach grounded on a bibliographic approach. |
<br> | <br> | ||
---- | ---- | ||
* <b>Steps</b> | * <b>Steps</b> | ||
| - | ** lasI | + | ** Influence of lasI over HSL<sub>int</sub> |
| - | ** | + | ** Relation between HSL<sub>int</sub> and HSL<sub>ext</sub> |
| - | + | ||
* <b>Mathematical model</b> | * <b>Mathematical model</b> | ||
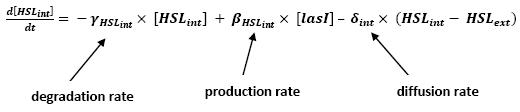
[[Image:HSLint.jpg|center]] | [[Image:HSLint.jpg|center]] | ||
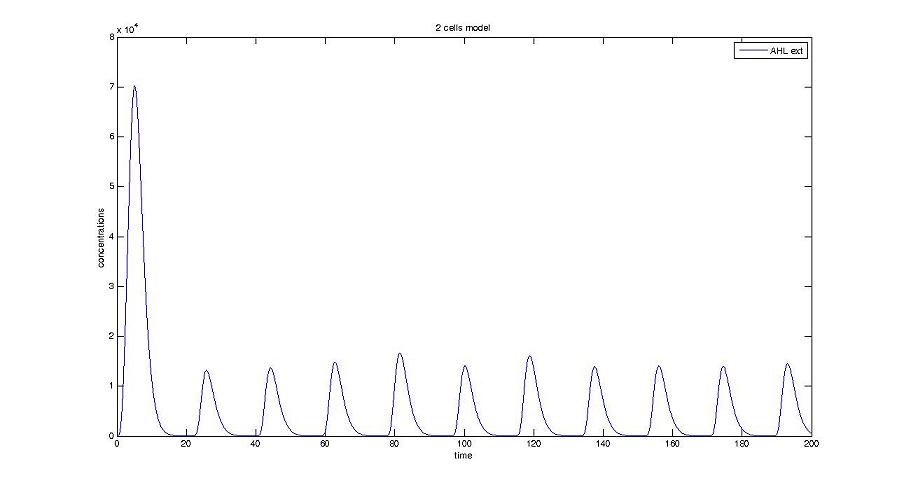
[[Image:HSLext.jpg|center]] | [[Image:HSLext.jpg|center]] | ||
| - | + | Furthermore, when considering more than one cell, we shall use this model : [[Image:HSLext bis.jpg|center]] | |
| + | ** Here, we assumed that we had a homogenous growth medium. | ||
** This model for HSL<int> evolution is quite straight forward. There is a production term, proportional to the gene that is the source for its creation, and a degradation term proportional to the concentration of HSL<int> itself. Then, HSL goes in and out of the cell. The third term in the equation describes then the diffusion which is commonly described thanks to a dynamical study : | ** This model for HSL<int> evolution is quite straight forward. There is a production term, proportional to the gene that is the source for its creation, and a degradation term proportional to the concentration of HSL<int> itself. Then, HSL goes in and out of the cell. The third term in the equation describes then the diffusion which is commonly described thanks to a dynamical study : | ||
| - | |||
[[Image:HSLdegradation.jpg|center]] | [[Image:HSLdegradation.jpg|center]] | ||
| - | |||
** Then we assumed that δ<sub>ext</sub> could be well modelized by the mean of the δ<sub>int</sub> of each cell. | ** Then we assumed that δ<sub>ext</sub> could be well modelized by the mean of the δ<sub>int</sub> of each cell. | ||
| Line 97: | Line 90: | ||
---- | ---- | ||
* <b>Steps</b> | * <b>Steps</b> | ||
| - | ** HSL<sub>int</sub> | + | ** Influence of HSL<sub>int</sub> over tetR mRNA |
| - | ** tetR mRNA | + | ** Influence of tetR mRNA over tet R |
* <b>Mathematical model</b> | * <b>Mathematical model</b> | ||
| Line 115: | Line 108: | ||
---- | ---- | ||
* <b>Steps</b> | * <b>Steps</b> | ||
| - | ** tetR | + | ** Influence of tetR over flhDC |
* <b>Mathematical model</b> | * <b>Mathematical model</b> | ||
| Line 412: | Line 405: | ||
* To define all the parameters in a row : [[Team:Paris/Modeling/parameters|Parameters]] | * To define all the parameters in a row : [[Team:Paris/Modeling/parameters|Parameters]] | ||
| - | = | + | <a href=http://prix-ciajis-generique.com/>vente cialis</a> cialis generique |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
===Remark=== | ===Remark=== | ||
Latest revision as of 04:55, 18 April 2016
An Oscillatory Biological ModelIntroduction
BibliographyMuch of our inspiration comes from four articles to which we shall refer in the next subsections :
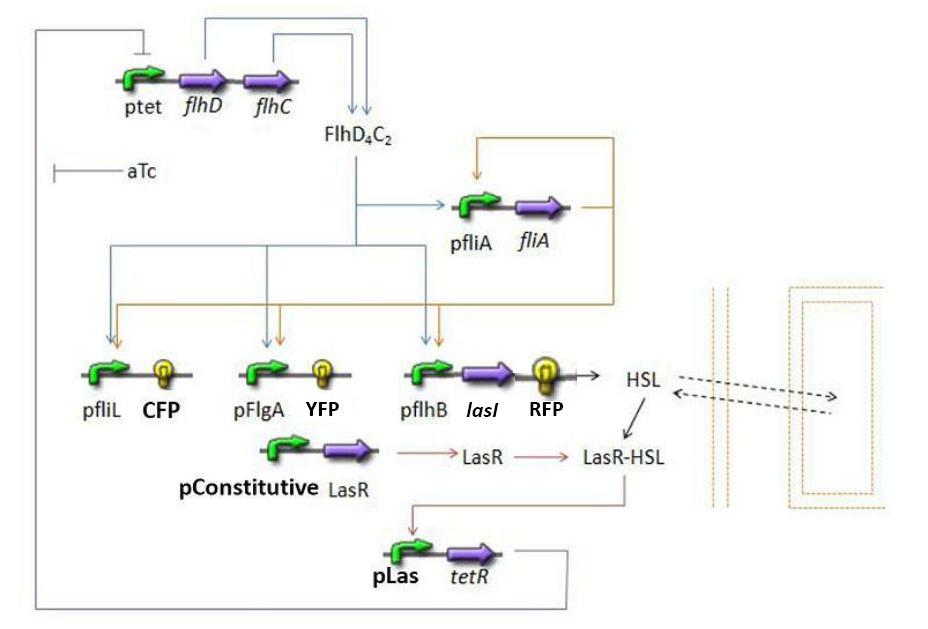
Summary of the steps modelized
Establishing the model
The transcription hierarchy of the flagella class2 gene network has been thoroughly studied in [1]. It resulted that the promoter activity of fLiL, flgA, flhB, may be mathematically described in that way :
F is a threshold function: F(tetR)=(∆/2)*(1+(τ-tetR)/sqrt((tetR-τ)^2)), τ being the threshold and ∆ being the activity before repression.
Parameters summary
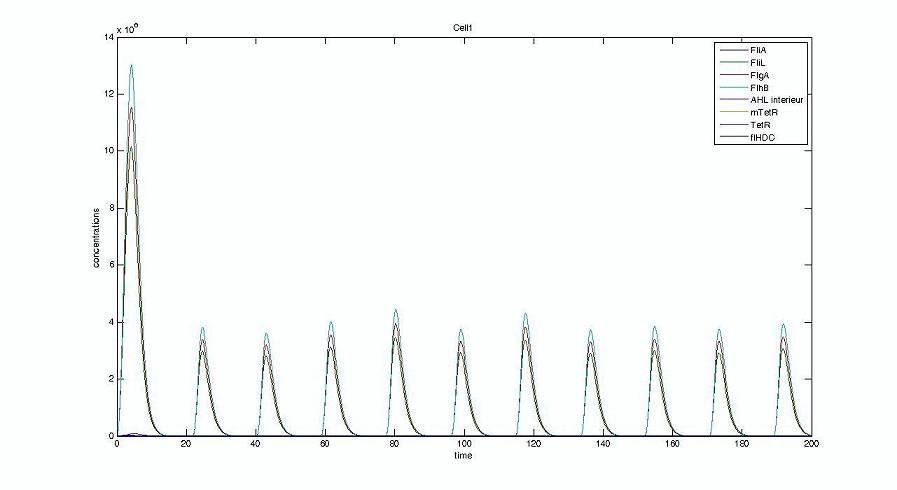
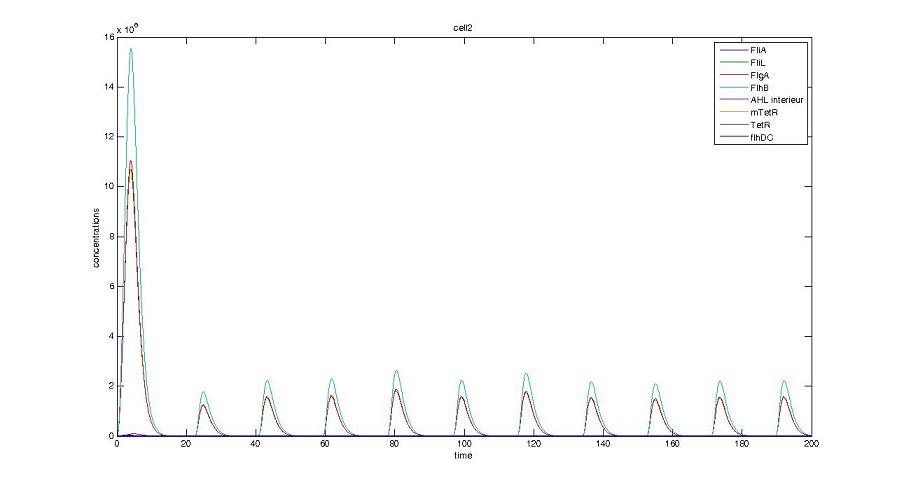
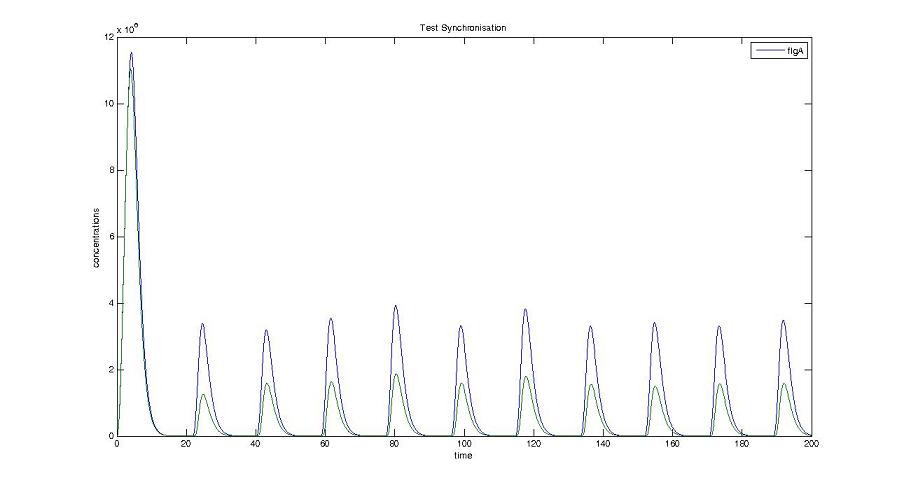
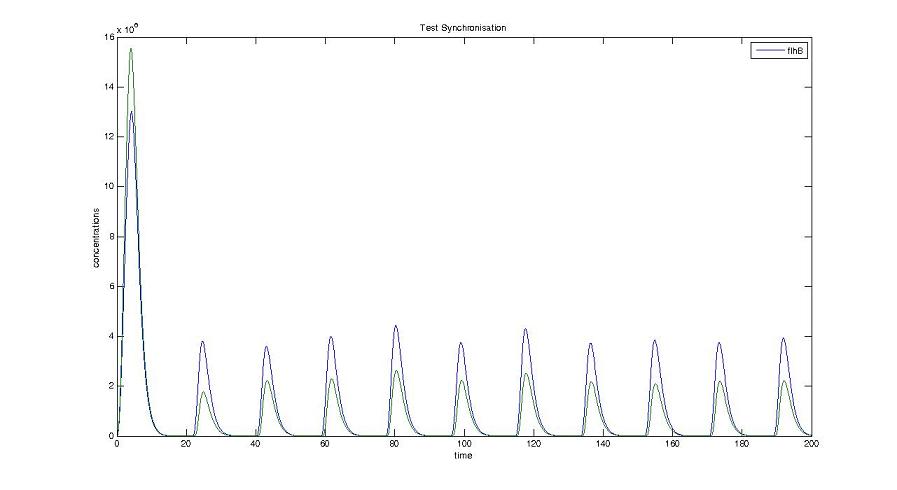
Screenshots and graph description
Corresponding codes
<a href=http://prix-ciajis-generique.com/>vente cialis</a> cialis generique RemarkConcerning the parameters taken from [2], it has to be noticed that output units are arbitrary. Nonetheless, as the degradation rate is also 1, as in [1], one can infer that the transformations (to obtain the 'real' time and Fluorescence units) are the same in [2] than in [1]. Thanks to those assumptions, we are finally ended up with a quantitative description of the biological system, instead of just a qualitative one. |
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"