Team:University of Washington/Project
From 2008.igem.org
(→Introduction: Controlled Trans-Kingdom Genetic Transfer) |
(→Introduction: Controlled Trans-Kingdom Genetic Transfer) |
||
| Line 44: | Line 44: | ||
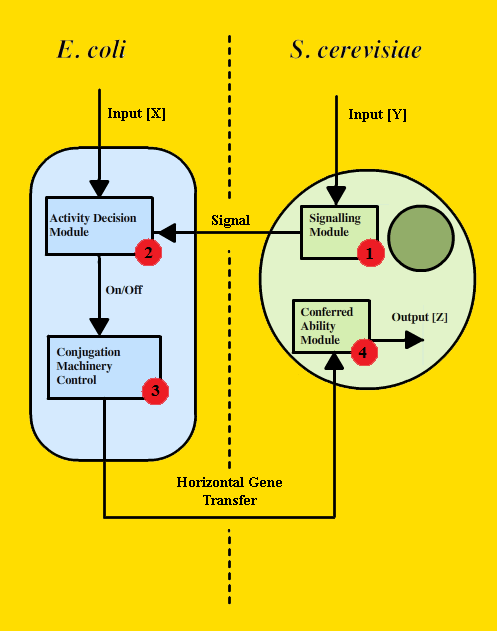
The modular design for trans-kingdom genetic transfer could be adapted for use with various organisms under different circumstances. However, our particular design is intended for a scenario where ''E. coli'' and ''S. cerevisiae'' are in close proximity, the yeast cannot naturally digest lactose, lactose is the primary food source, and the ''E. coli'' possess a transferrable plasmid containing genes for lactose digestion. Because expending resources on lactose machinery in the presence of the more readily usable glucose would be disadvantageous, ''E. coli'' will only produce the LuxR protein in the presence of lactose, and the absence of glucose. The yeast will be engineered to produce the quorum sensing molecule, AHL. When the AHL diffuses to the LuxR proteins contained in ''E. coli'', the production of conjugation machinery will be activated, and genetic material will be transferred to yeast. Pragmatically, this schema for trans-kingdom genetic transfer could be used to modify and engineer organisms which are normally difficult to access or operate on, and also as a means of delivering functional DNA for gene therapy. Additionally, this design may serve as an entry point for more complex, multi-organism systems. | The modular design for trans-kingdom genetic transfer could be adapted for use with various organisms under different circumstances. However, our particular design is intended for a scenario where ''E. coli'' and ''S. cerevisiae'' are in close proximity, the yeast cannot naturally digest lactose, lactose is the primary food source, and the ''E. coli'' possess a transferrable plasmid containing genes for lactose digestion. Because expending resources on lactose machinery in the presence of the more readily usable glucose would be disadvantageous, ''E. coli'' will only produce the LuxR protein in the presence of lactose, and the absence of glucose. The yeast will be engineered to produce the quorum sensing molecule, AHL. When the AHL diffuses to the LuxR proteins contained in ''E. coli'', the production of conjugation machinery will be activated, and genetic material will be transferred to yeast. Pragmatically, this schema for trans-kingdom genetic transfer could be used to modify and engineer organisms which are normally difficult to access or operate on, and also as a means of delivering functional DNA for gene therapy. Additionally, this design may serve as an entry point for more complex, multi-organism systems. | ||
| + | |||
| + | {| style="background | ||
| + | | | ||
[[Image:uwbigpicture.jpg|400px]] | [[Image:uwbigpicture.jpg|400px]] | ||
Revision as of 07:14, 28 October 2008

| Home | The Team | The Project | Modeling | Notebook | Protocols | Parts Submitted to the Registry | Measurement Kit | SeToB | Safety |
|---|
Introduction: Controlled Trans-Kingdom Genetic Transfer
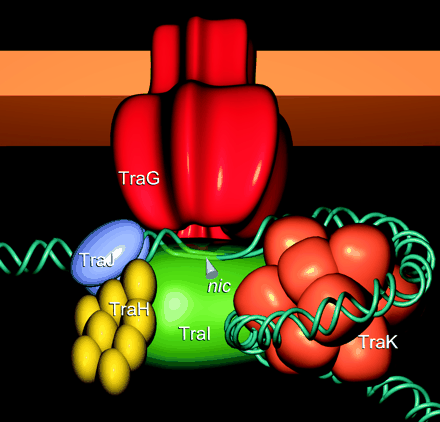
Certain bacteria exhibit a means of horizontal gene transfer called conjugation that contrasts with traditional heritable genetics. Specifically, the mechanism of bacterial conjugation does not rely on the transfer of genetic information from parent to progeny. Instead, a unique type of plasmid induces the replication and transmission of itself or other plasmids, called shuttle plasmids between bacteria via direct cell-to-cell contact. The genes on the conjugative plasmid encode for a suite of enzymes and structural proteins, including a sex pilus nanotube that connects the bacteria and accommodates the transmission of DNA. In this manner, bacteria can accept and express genes separate from their chromosomal DNA. This mode of genetic transfer is fascinating on many levels, including its possible role in evolution and phylogenetic discrepancies, it has the potential to be utilized as an invaluable mechanism for genetic alteration.
In addition to horizontal gene transfer between bacteria, bacteria can also transfer genetic material across kingdoms to the eukaryote S. cerevisiae, more commonly known as baker's yeast, an organism incredibly distant evolutionarily from any bacterial species [1]. Unlike sexual reproduction, which requires great genetic similarity between mating partners, it would appear that conjugation provides a means for transferring genetic material across vast evolutionary expanses. Our goal is to engineer E. coli to conjugate across phylogenetic kingdoms under the presence and absence of specific environmental signals. However, in order to prevent the genetic circuitry from being “evolved out,” the plasmid will confer some adaptive function to the yeast. Eventually, the yeast will be engineered to produce some beneficial product for E. coli, and the two species will live symbiotically, exchanging DNA and cellular products necessary for survival. In this way, there will be a mutualistic interaction between the yeast and E. coli, and it is hoped that the functional integrity of the genetic circuitry will remain intact.
The modular design for trans-kingdom genetic transfer could be adapted for use with various organisms under different circumstances. However, our particular design is intended for a scenario where E. coli and S. cerevisiae are in close proximity, the yeast cannot naturally digest lactose, lactose is the primary food source, and the E. coli possess a transferrable plasmid containing genes for lactose digestion. Because expending resources on lactose machinery in the presence of the more readily usable glucose would be disadvantageous, E. coli will only produce the LuxR protein in the presence of lactose, and the absence of glucose. The yeast will be engineered to produce the quorum sensing molecule, AHL. When the AHL diffuses to the LuxR proteins contained in E. coli, the production of conjugation machinery will be activated, and genetic material will be transferred to yeast. Pragmatically, this schema for trans-kingdom genetic transfer could be used to modify and engineer organisms which are normally difficult to access or operate on, and also as a means of delivering functional DNA for gene therapy. Additionally, this design may serve as an entry point for more complex, multi-organism systems.
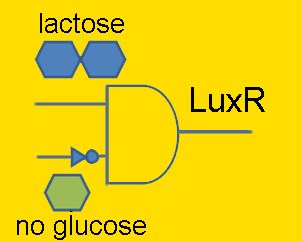
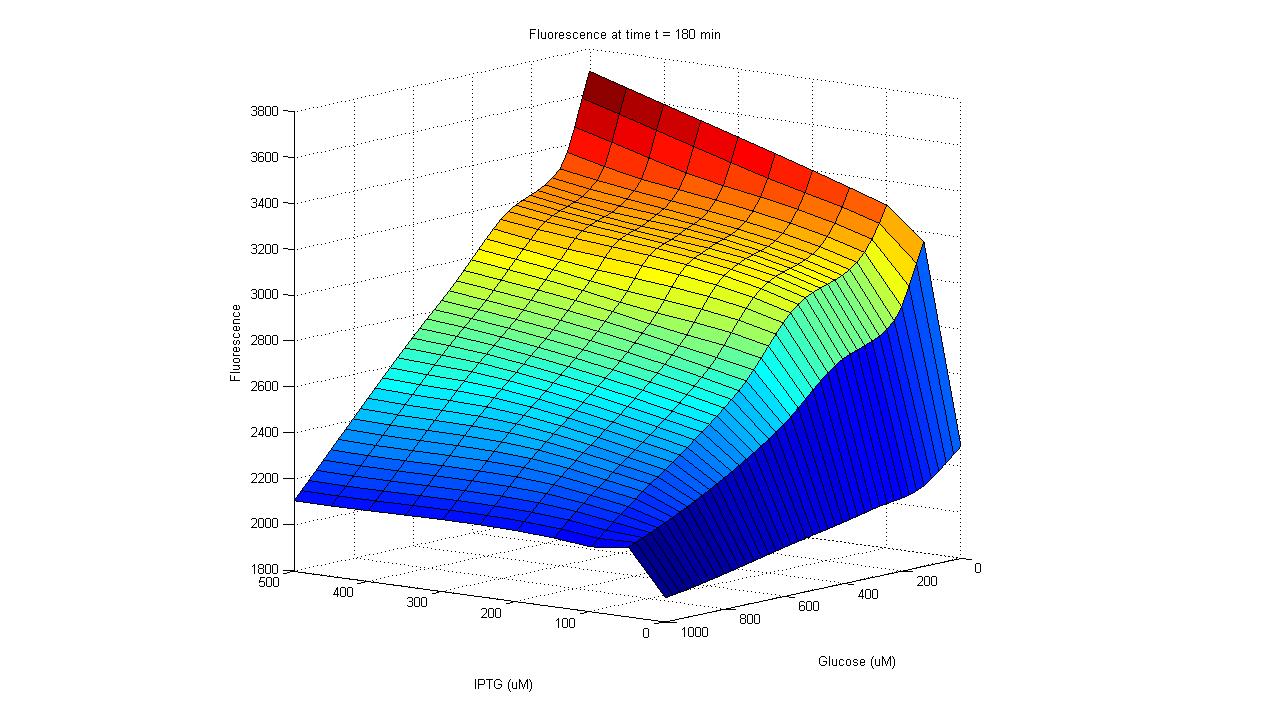
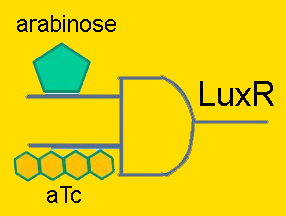
Individual ProjectsSub#1:Activity Decision Module - LuxR production under pLac/pAraC+TetR in bacteriaRegulators: pLacLactose is a disaccharide composed of glucose and galactose, and in order for an organism to metabolize lactose, it first must be broken-up into these constituent monomers. β-galactosidase is an enzyme capable of doing just that, and is naturally produced by E. coli. However, E. coli cleverly regulate their production of this enzyme to times only when it’s required. Being that lactose is a less efficient fuel than simple glucose, E. coli only want to metabolize lactose when glucose is absent. This regulation is carried out at the transcriptional level by putting the β-galactosidase gene (and 2 other genes) under the control of a promoter (the lac promoter) which is inhibited by LacI and induced by CAP. LacI is a molecule that is constitutively produced by E. coli; it can either bind to the promoter or bind to lactose, meaning that LacI only inhibits the promoter in the absence of lactose. CAP is another molecule naturally present in E. coli which is only free to bind to the lac promoter in the absence of glucose. Given that gene repression is stronger than activation, the production of β-galactosidase is only carried out when required. In order to have LuxR produced only when glucose is absent and lactose is present, the LuxR protein generator BioBrick (part BBa_I0462) was placed under the control of the lac promoter (part BBa_R0010). The lac promoter is a commonly used transcriptional regulator, as it enables genes to be turned on/off with addition/subtraction of lactose (or IPTG) from the growth medium. On the other hand, the use of glucose as a means to regulate the activity of the promoter is not common. Thus, promoter part BBa_R0010 had to be tested to verify that it indeed behaves as the natural lac promoter does (i.e. IPTG can only successfully activate the promoter in the absence of glucose). To carry out such a test, part BBa_R0010 was placed in front of the GFP generator BioBrick (part BBa_E0240). An assay was then carried out that characterized the promoter activity in varying concentrations of IPTG and glucose: 25 wells of a 96-well plate were filled with the mid-log culture of a LacI expressing E. coli strain (DH5a+LacIq) grown in M9 minimal media. IPTG and glucose was then added to the wells in different amounts. The GFP fluorescence of each well was then measured over a six hour period. As the plot shows below, GFP has the highest expression in the well with the highest concentration of IPTG and no glucose, and the lowest expression in the well with the highest glucose concentration and no IPTG. Additionally, in wells with both glucose and IPTG, the repression of the glucose proved stronger than the activation of the IPTG. Thus the promoter part behaves just as required. After parts BBa_R0010 and BBa_I0462 were ligated together, they were placed inside the low/medium copy, kan resistant plasmid pSB3K3. This new composite part was given name BBa_K109702. Activity of pLac in Varying Concentrations of IPTG and Glucose A GFP generator gene (part BBa_E0240) was placed under the control of the Lac promoter (part BBa_R0010). Fluorescence was then measured under varying concentrations of IPTG and glucose to characterize the promoter's activity. Regulators: pAraC+TetR
Sub#2:Signaling Module - Biosynthesis of the AHL distress signal (in Yeast)Ultimately, our goal is to place bacterial conjugation under the control of an external distress signal from a metabolically-stressed yeast cell. Thus, it is necessary to engineer S. cerevisiae to produce the AHL signaling molecule. Our proposed method to accomplish this is to clone the AI synthase gene (BBa_C0161) into a nonstandard yeast expression plasmid, which should theoretically result in the conversion of amino acid precursors into the AHL molecule. To test whether the yeast transformants are indeed capable of AHL signaling, they can be tested in co-culture with the AHL receiver cells (BBa_T9002). Although AHL production will initially be under control of a constitutive promoter, eventually we may place it under control of a glucose-repressible promoter such as pJEN1 [3], so that the yeast will produce AHL proportionally in response to glucose starvation. Sub#3: Conjugation Machinery Control - Replacement of Conjugative Promoters with an Inducible Promoter
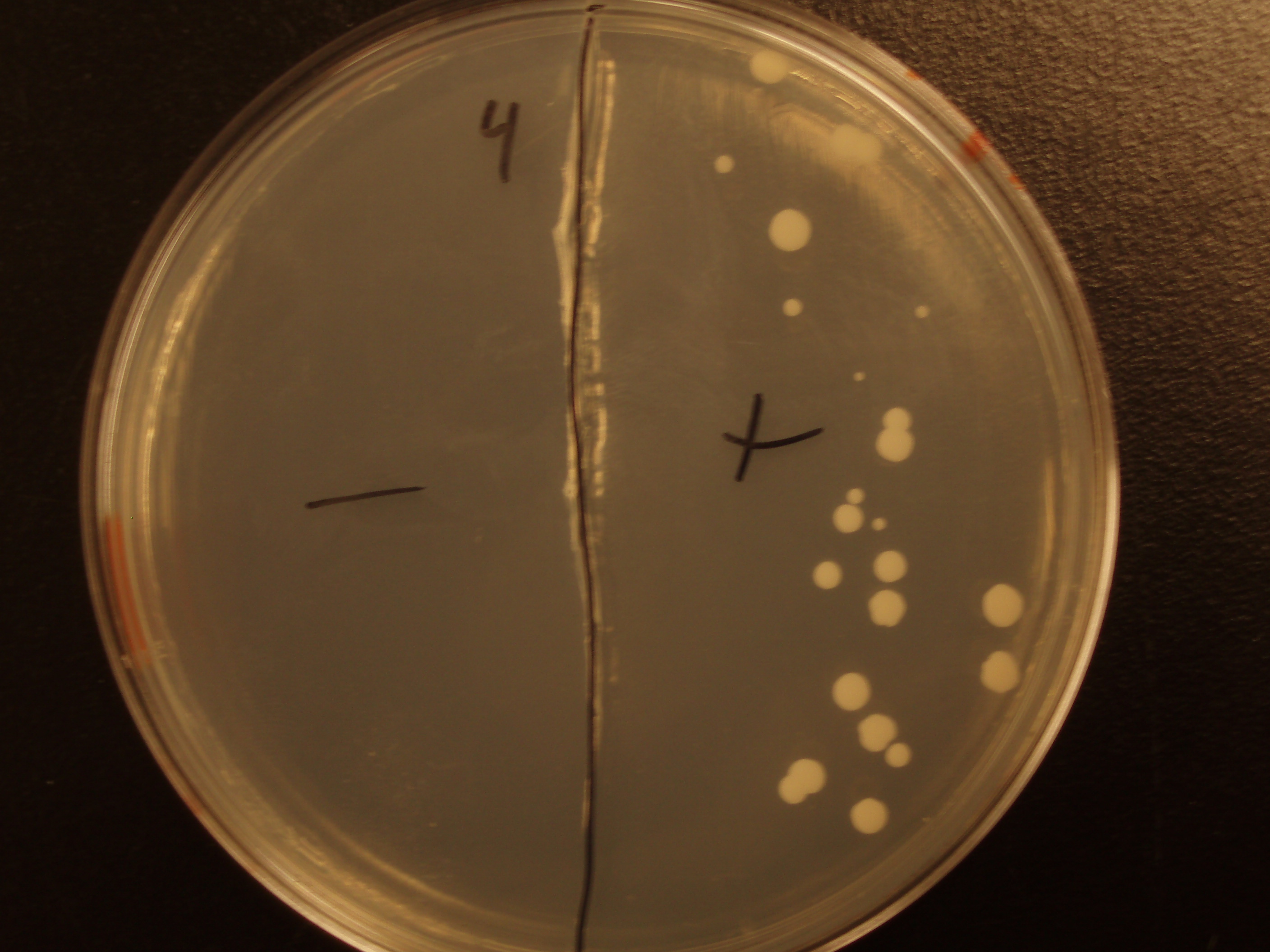
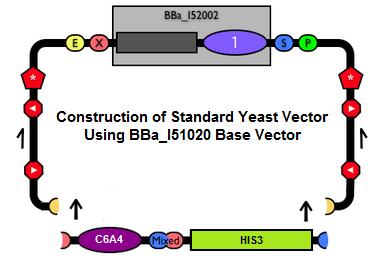
Sub#4: Conferred Ability Module- Yeast Shuttle Vector
[1] Sprague, George F., and Jack A. Heinemann. “Bacterial conjugative plasmids mobilize DNA transfer between bacteria and yeast.” Nature. Institute of Molecular Biology. University of Oregon. 340. 20 Jul 1989. [2] Robert Sidney Cox, III, Michael G Surette, and Michael B Elowitz. "Programming gene expression with combinatorial promoters." Nature. Institute of Molecular Biology. 145. 13 November 2007 [3] Chambers P, Issaka A, Palecek SP. "Saccharomyces cerevisiae JEN1 promoter activity is inversely related to concentration of repressing sugar." Appl Environ Microbiol. 2004 Jan;70(1):8-17. [4] Zatyka M, Jagura-Burdzy G, Thomas CM. "Transcriptional and translational control of the genes for the mating pair formation apparatus of promiscuous IncP plasmids." J Bacteriol. 1997 Dec;179(23):7201-9. [5] Cashmore, Annette M., Steven Bates, and Brian M. Wilkins. “IncP Plasmids Are Unusually Effective in Mediating Conjugation of Escherichia coli and Saccharomyces cerevisiae: Involvement of the Tra2 Mating System.” Journal of Bacteriology. Dec 1998. pp 6538-6543.
|
 "
"